Science Forum: Vision, challenges and opportunities for a Plant Cell Atlas
Figures
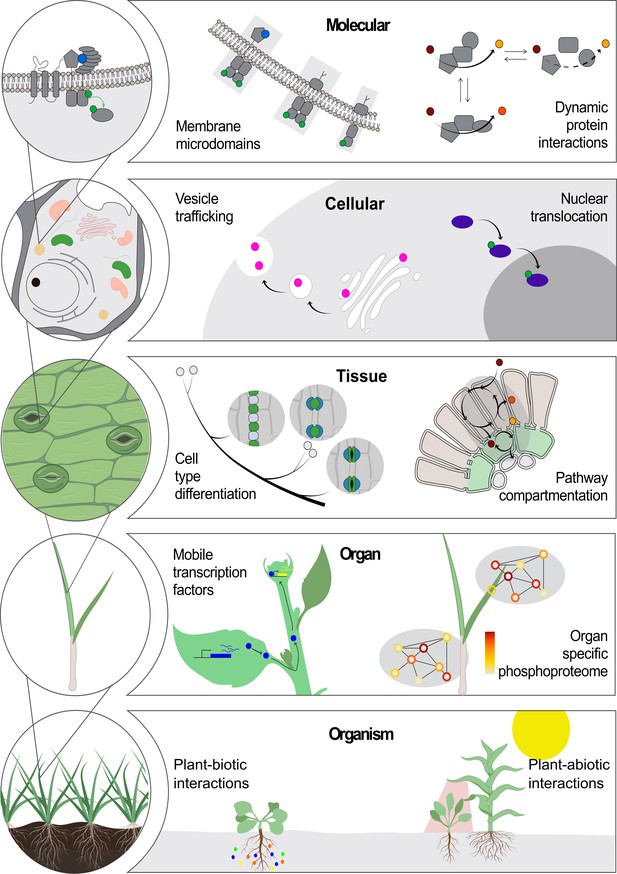
Location-to-function paradigm.
The illustration shows the levels of organization of a plant (left) and a few examples highlight how location determines function at each level (right). At the molecular level, functions of protein complexes can be determined by their localization in membrane microdomains (Jarsch et al., 2014) or by dynamic protein interactions (Obata, 2019). At the cellular level, a protein can be located differentially through transport mechanisms, including vesicle trafficking (Goring and Di Sansebastiano, 2018) and nuclear translocation (Marchive et al., 2013), regulating its function. At the tissue level, cell position can drive its fate into a specialized cell type (Shao and Dong, 2016). Metabolic pathways can operate specifically in specialized cell types across tissues (Marchive et al., 2013; Schlüter and Weber, 2020). At the next level, the existence of non-cell-autonomous transcription factors can transverse intercellular scales across plant organs (Han et al., 2014). Also, an organ-dependent post-translational proteome has been described as a mechanism of protein function regulation (Uhrig et al., 2019). At the organism level, plant interaction with biotic (Harrison et al., 2002) and abiotic factors (Michaud et al., 2017) can occur through a localized cue perception.
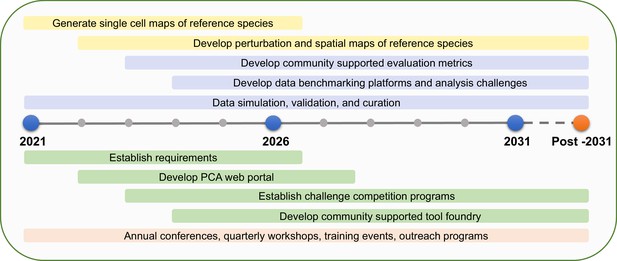
PCA milestones.
PCA milestones for the next 10 years and beyond in data generation (yellow bars), data analysis (purple bars), software development (green bars), and building PCA community (orange bar).
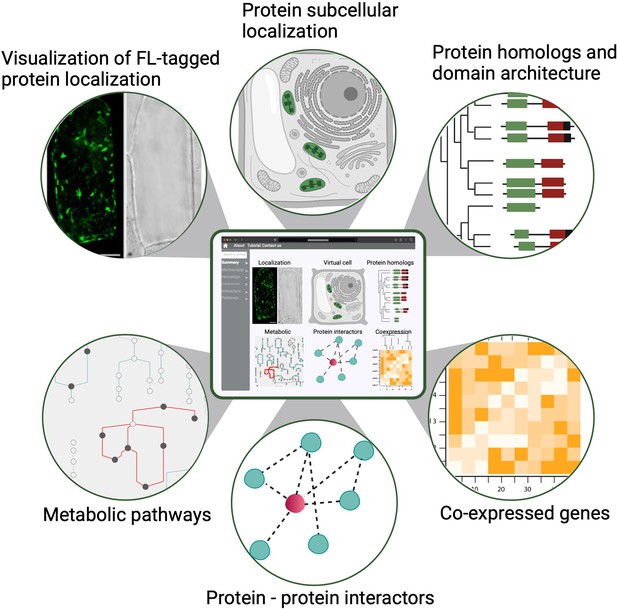
A conceptual diagram of a PCA user interface.
An example of a PCA user interface that integrates various data types from molecular, biochemical, cellular and evolutionary contexts to connect location to function. Shown are example data types that would be seamlessly connected to enable easy navigation and discovery (FL: fluorescence).
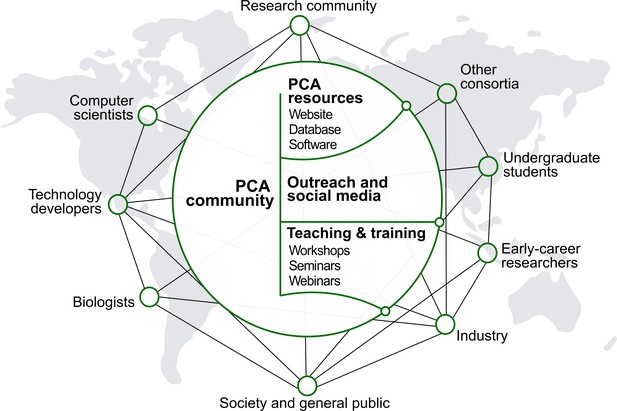
People and culture.
Major stakeholders of the PCA are described. The goal is to establish a broad network of collaboration between developers and users through the creation of an accessible platform, educational tools and outreach activities. The PCA community should strive to be inclusive, diverse and transparent.
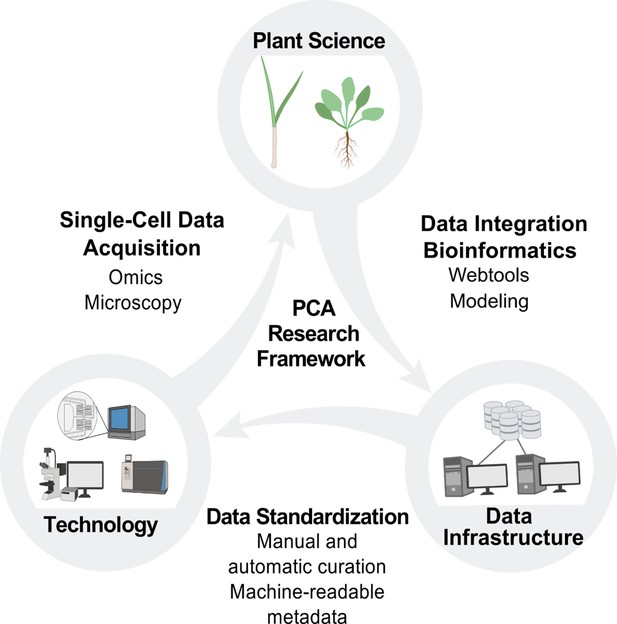
PCA research framework.
Building the PCA will require cooperation and coordination between plant science, technology and data infrastructure. Plant scientists will use new technologies to generate data at the single-cell level, which will be integrated by data scientists into the PCA infrastructure through manual and automated curation. This data integration will enable the development of new technologies and drive new hypotheses and investigations for plant scientists, who will then contribute additional data to the PCA resource.
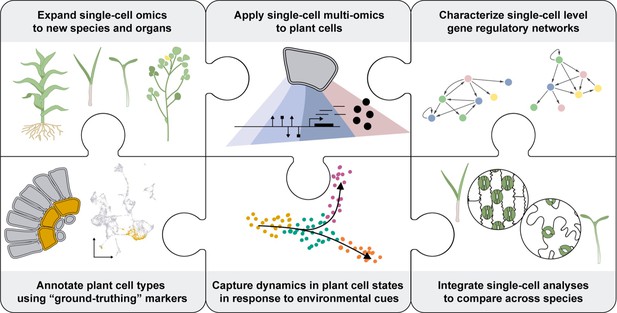
Knowledge gaps to fill for the PCA.
Several limitations inherent to plants (shown as pieces of a jigsaw puzzle) must be overcome to enhance the understanding of plant cells as biological systems. To characterize the unique molecular attributes of each cell type composing a plant, plant biologists will need to develop reliable methods to broadly access the different cell types across various plant species. Also, single-cell multi-omics technologies will be necessary to get a deeper understanding of plant cell processes across all layers of molecular regulation. This data must be gained in the context of functional annotation of the cells. This would require the identification of reliable marker genes for each type of plant cell-. Ultimately, the spatiotemporal distribution of the molecular attributes of each cell will help to understand their dynamic regulation during plant cell development and in response to environmental stresses. Integrating the information collected using bioinformatics tools will enable the characterization of regulatory networks at a single-cell resolution and comparative analyses across plant species at the cellular level. This data will fuel the establishment of new synthetic biology strategies to enhance plant biology.
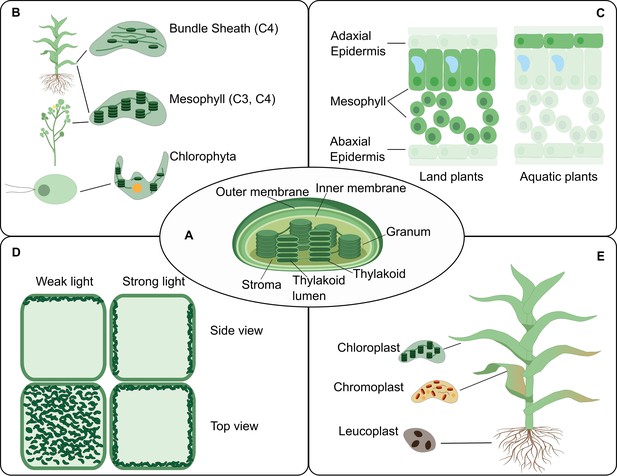
The chloroplast structure and function in connection to location.
The PCA could facilitate an increased understanding of the biology of the chloroplast by providing accessible, integrative and spatially-resolved data for transcripts, proteins and metabolites across cell types (Box 2). (A) A representation of a typical chloroplast structure and its compartments. (B) The chloroplast structural differences in some plant lineage. (C) A leaf cross-section showing different chloroplast location at the tissue level in land and aquatic plants. (D) The chloroplast location can change within the cell upon environmental cues such as light intensity. (E) Various forms of chloroplasts in different organs.





