Chondroitin sulfate proteoglycan 4,6 sulfation regulates sympathetic nerve regeneration after myocardial infarction
Figures
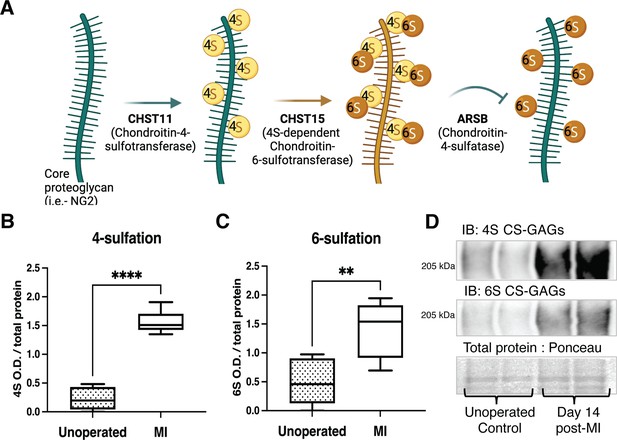
Chondroitin sulfate proteoglycan (CSPG) 4,6-sulfation increases in the cardiac scar after myocardial infarction (MI) caused by ischemia/reperfusion (I/R).
(A) CSPG sulfation patterning schematic with key enzymes. (B) 4-Sulfation (4S CS-GAGs) or (C) 6-sulfation (6S CS-GAGs) quantification assessed by western blot in the healthy myocardium (unoperated) or cardiac scar 14 days after MI (MI). Quantification of n=6 animals per treatment group, mean optical density (OD) ± SD, Student’s t-test (Welch’s test), 4S ****p-value < 0 .0001, 6S **p-value = 0.003. (D) Example blot images for 4S CS-GAGs, 6S CS-GAGs, and total protein from two unoperated and two MI animals.
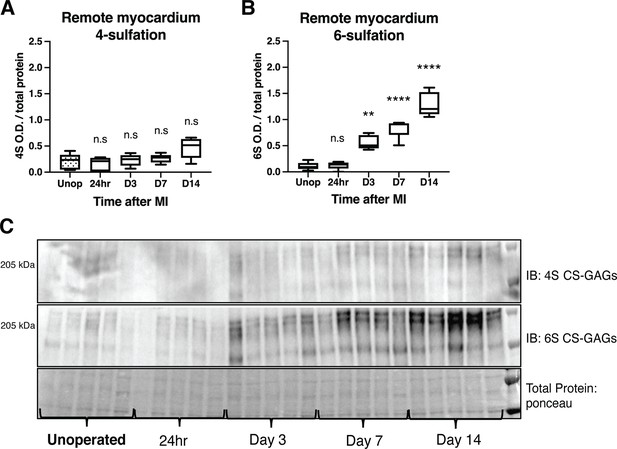
Time course of chondroitin sulfate proteoglycan (CSPG) sulfation in the remote myocardium (non-scar tissue) after myocardial infarction (MI) caused by ischemia/reperfusion (I/R).
Western blot quantification of (A) 4-sulfation (4S CS-GAGs). Statistics: one-way ANOVA (Tukey’s post-test), ns – not significant p-value = 0.989, 0.997, 0.955, 0.053 respectively left to right, comparisons to unoperated tissue, n=5 animals per group. (B) Western blot quantification of 6-sulfation (6S CS-GAGs). Statistics: one-way ANOVA (Tukey’s post-test), ns p-value = 0.999, **p-value = 0.001, ****p-value < 0.0001, comparisons to unoperated tissue, n=5 animals per group. (C) Example western blot images of A and B.
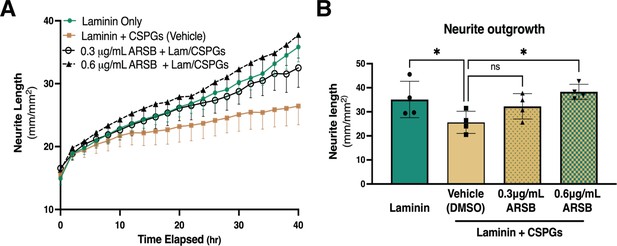
Reducing CS-GAG 4-sulfation promotes sympathetic neurite outgrowth in vitro.
(A) Example of neurite outgrowth experiment with arylsulfatase-B (ARSB). Data are mean neurite length ± SD at 9 locations per well and 3 wells per condition. (B) Removal of 4-sulfation with ARSB restores neurite outgrowth to control levels. Quantification of dissociated sympathetic neurite length at 40 hr post-plating on indicated plate coatings. Data are mean neurite length ± SD; one-way ANOVA (Dunnett’s post-test). All comparisons made to neurite outgrowth on laminin + CSPG; 0.3 μg/mL ARSB ns – not significant p-value = 0.193, laminin only *p=0.039, 0.6 μg/mL ARSB *p-value = 0.042, n=4 experiments.
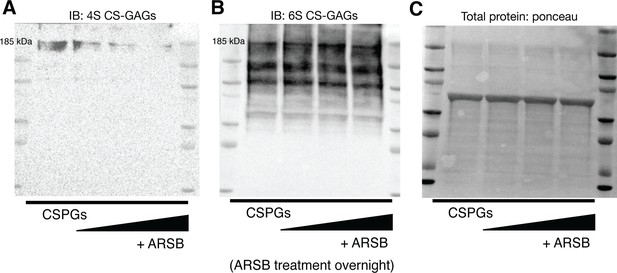
Arylsulfatase-B (ARSB) removes 4S CS-GAGs and leaves 6S CS-GAGs intact.
(A) Western blot of 4S content using purified chondroitin sulfate proteoglycans (CSPGs) upon treatment of increasing concentrations of ARSB; vehicle, 0.3, 0.6, and 1.2 μg/mL respectively left to right. (B) Western blot of 6S using purified CSPGs upon treatment of increasing concentrations of ARSB; vehicle, 0.3, 0.6, and 1.2 μg/mL respectively left to right. (C) Total protein loaded of purified CSPGs treated with increasing concentrations of ARSB; vehicle, 0.3, 0.6, and 1.2 μg/mL respectively left to right.
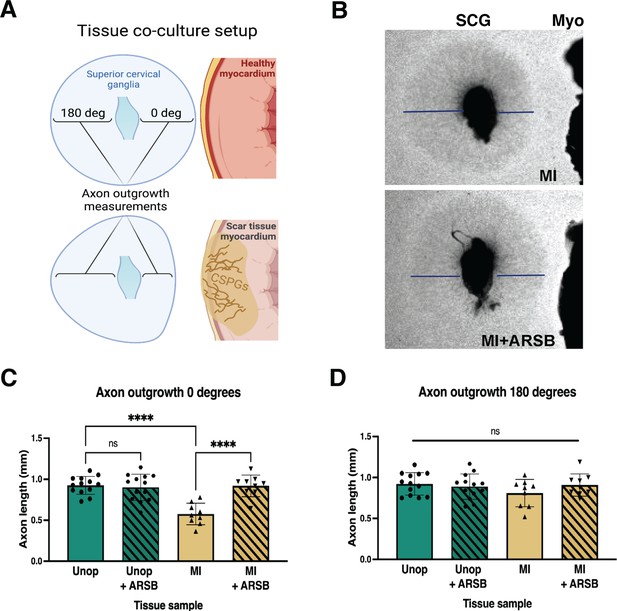
Reducing CS-GAG 4S in cardiac scar tissue ex vivo restores sympathetic axon outgrowth.
(A) Explant co-culture schematic. Ganglion axon extension toward (0 degree) and away from (180 degree) myocardium was measured. (B) Example images of superior cervical ganglion (SCG) axon outgrowth in the presence of cardiac scar tissue (Myo) treated with or without arylsulfatase-B (ARSB). Lines show example measurement of axon extension 48 hr after plating. (C) Quantification of ganglion axon outgrowth toward myocardium of either healthy tissue (Unop) or scar tissue (myocardial infarction [MI]) treated with vehicle (5% DMSO) or ARSB (0.6 μg/mL). (D) Quantification of ganglion axon outgrowth away from myocardium of either healthy tissue (Unop) or scar tissue (MI) treated with vehicle (5% DMSO) or ARSB (0.6 μg/mL). Data are mean axon length ± SD. Statistics for C, D: one-way ANOVA (Dunnett’s post-test), comparisons made to vehicle-treated MI tissue; 0 degree quantification ****p<0.0001, 180 degrees quantification ns – not significant p-value = 0.204, 0.467, 0.355 left to right respectively, n=13 control tissue and n=9 MI tissue.
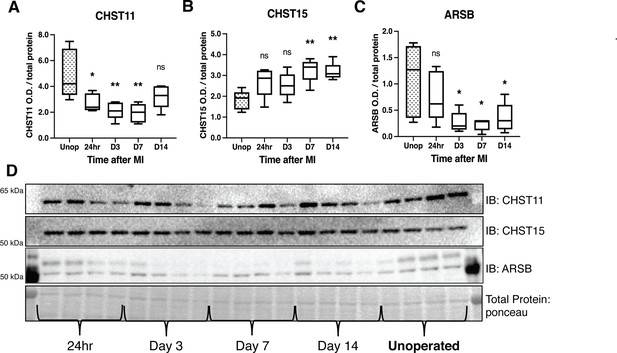
Chondroitin sulfate proteoglycans (CSPG) sulfation enzyme expression is altered after ischemia/reperfusion (I/R).
Western blot quantification of protein expression for (A) chondroitin-4-sulfotransferase (CHST11), (B) chondroitin-6-sulfotransferase (CHST15), and (C) 4-sulfatase (arylsulfatase-B [ARSB]) protein expression in control left ventricle (Unop), or in the days after myocardial infarction (MI) (24 hr, D3, D7, or D14). Data are mean optical density (OD) ± SD. Statistics: one-way ANOVA (Dunnett’s post-test) comparisons to unoperated tissue; CHST11 24 hr *p-value = 0.015, D3 **p-value = 0.002, D7 **p-value = 0.001, D14 ns – not significant p-value = 0.068; CHST15 24 hr ns p-value = 0.073, D3 ns p-value = 0.164, D7 **p-value = 0.002, D14 **p-value = 0.004; ARSB 24 hr ns p-value = 0.554, D3 *p-value = 0.018, D7 *p-value = 0.012, D14 *p-value = 0.035; n=5 animals per group. (D) Example western blot images for CSPG sulfation enzymes.
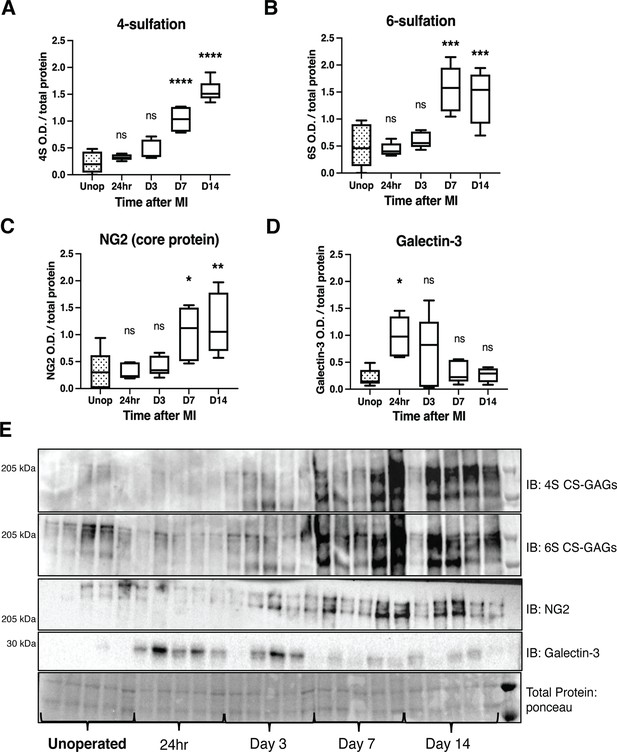
Time course of chondroitin sulfate proteoglycan (CSPG) sulfation and core protein expression after ischemia/reperfusion (I/R) in cardiac scar tissue.
Western blot quantification of (A) 4-sulfation (4S CS-GAGs), (B) 6-sulfation (6S CS-GAGs), (C) NG2 core protein, and (D) Galectin-3 in the days after MI. Data are mean optical density (OD) ± SD. Statistics: one-way ANOVA (Dunnett’s post-test), comparisons to unoperated tissue; 4S, 24 hr ns – not significant p-value = 0.804, D3 ns p-value = 0.124, D7 ****p-value < 0.0001, D14 ****p-value < 0.0001; 6S, 24 hr ns p-value = 0.999, D3 ns p-value = 0.905, D7 ***p-value = 0.0001, D14 ***p-value = 0.0003; NG2, 24 hr ns p-value = 0.999, D3 ns p-value = 0.977, D7 *p-value = 0.031, D14 **p-value = 0.007; Galectin-3, 24 hr *p-value = 0.013, D3 ns p-value = 0.172, D7 ns p-value = 0.974, D14 ns p-value = 0.998; n=5 animals per group. (E) Example western blot images of A–D.
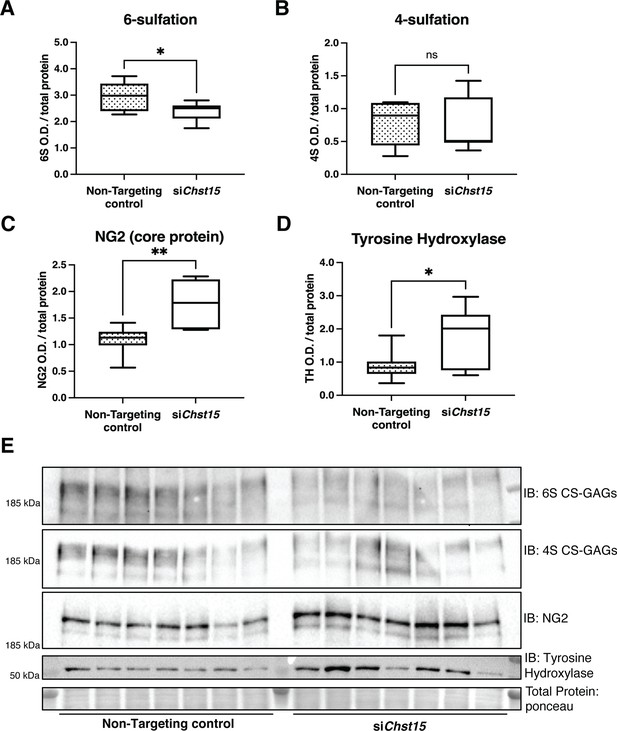
Reducing CS-GAG 4,6-tandem sulfation in vivo promotes sympathetic nerve regeneration into the cardiac scar after ischemia/reperfusion (I/R).
Western blot quantification of (A) 6-sulfation (6S CS-GAGs), (B) 4-sulfation (4S CS-GAGs), (C) NG2 core protein, and (D) the sympathetic neuron marker Tyrosine Hydroxylase (TH) in the cardiac scar after transient Chst15 knockdown or treatment with a non-targeting silencing RNA (siRNA) control. Tissue was collected 10 days after myocardial infarction (MI), n=7 animals per group. Data are mean optical density (OD) ± SD. Statistics: Student’s t-test (Welch’s test); 6S, *p-value = 0.028; 4S, ns – not significant p-value = 0.826; NG2, **p-value = 0.004; TH, *p-value = 0.043. (E) Western blot images of A–D.
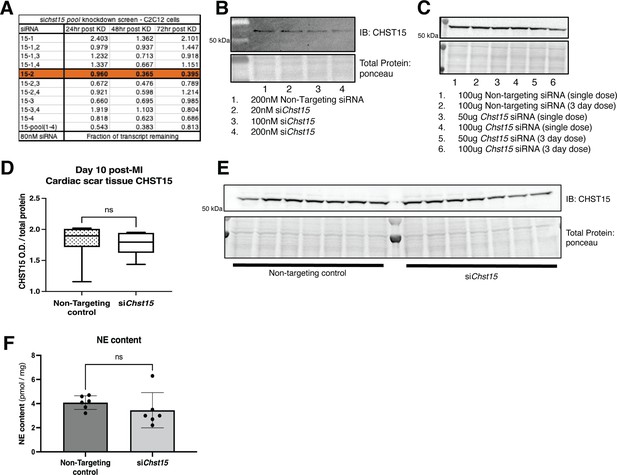
Identification of an effective silencing RNA (siRNA) against Chst15.
(A) qPCR knockdown of siRNA pool in C2C12 cells in 3 days after knockdown. Data shown fraction of transcript remaining, siChst15-2 was most effective in knockdown of Chst15. This transcript was selected for in vivo studies. (B) Western blot of CHST15 protein knockdown in C2C12 cells to confirm efficacy of siChst15-2, comparison to non-targeting controls. (C) Tail vein injection in mouse to determine ideal dosing for in vivo siChst15 knockdown. Western blot of tail vein injection dosing trial for siChst15 with either 1 or 3 days of injections. CHST15 protein in left ventricle (LV) 48 hr after final tail vein injection, comparison to non-targeting controls, all unoperated (non-myocardial infarction [MI]) animals. (D) Full siRNA CHST15 experimental trial, CHST15 protein expression on D10 post-MI quantified, siRNA injection D3, -5, -7 post-MI, n=7 animals per treatment group. Statistics: Student’s t-test (Welch’s test), ns – not significant p-value = 0.833. (E) Western blot of CHST15 protein expression D10 post-MI in siRNA-treated animals. (F) NE content in the cardiac scar following siRNA treatment. NE was not increased with reinnervation, consistent with previous studies showing suppression of NE synthesis and reuptake by inflammatory cytokines (Parrish et al., 2010). Quantification of n=6 animals for non-targeting controls and Chst15 siRNA treatment. Statistics: Student’s t-test (Welch’s test), ns p-value = 0.345.
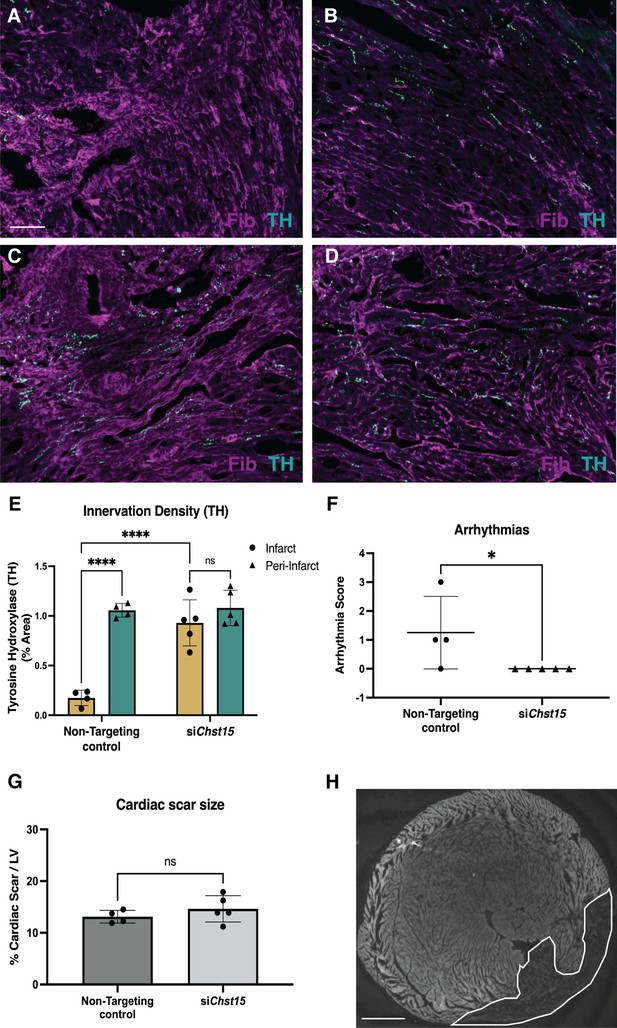
Reducing 4,6-tandem sulfation of CS-GAGs in vivo promotes sympathetic nerve regeneration into the cardiac scar.
Fibrinogen was used to label scar (magenta) and Tyrosine Hydroxylase (TH) was used to label sympathetic neurons (cyan). Example image of (A) denervated infarct in non-targeting control-treated animal versus (B) normal density of TH + fibers in the peri-infarct region adjacent to the scar. Systemic delivery of silencing RNA (siRNA) against Chst15 post-MI restored TH + fibers in the cardiac infarct (C) without altering nerve density outside the infarct (D). Scale bar, 100 μm. (E) Quantification of TH innervation density 10 days after MI. n=4 animals for non-targeting controls and n=5 for Chst15 siRNA-treated animals. Data are mean percent TH ± SD. Two-way ANOVA, Tukey’s post-test to compare all groups, select comparisons shown; siChst15 infarct vs. peri-nfarct ns – not significant p-value = 0.668, non-targeting infarct vs. peri-nfarct ****p-value < 0.0001, non-targeting infarct vs. siChst15 infarct ****p-value < 0.0001. (F) Arrhythmia scores based on the most severe arrhythmia observed in each heart after injection of isoproterenol and caffeine (0=no PVCs, 1=single PVCs, 2=bigeminy or salvos, 3=non-sustained ventricular tachycardia). See Materials and methods for details. Treatment with siRNA against Chst15 reduced arrhythmias compared to non-targeting controls. n=4 animals for non-targeting controls and n=5 animals for Chst15 siRNA. Data are arrhythmia score for each animal. Statistics: Student’s t-test (Mann-Whitney test), *p<0.047. (G) Cardiac scar size assessed as a percent area of total left ventricle (LV) area was unaltered in Chst15 siRNA-treated animals compared to non-targeting controls. Quantification of n=4 animals for non-targeting controls and n=5 for Chst15 siRNA-treated animals. Data are mean percent area scar ± SD. Statistics: Student’s t-test (Welch’s test), ns – not significant p-value = 0.283. (H) Example 2× image of cardiac scar in Chst15 siRNA-treated heart using autofluorescence, absence of autofluorescence indicates region of the infarct, outlined in white. Scale bar, 500 μm.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | C57BL/6J | Jackson Laboratories | 000664 | Male and female mice used for I/R surgeries |
| Strain, strain background (Rattus norvegicus) | Crl:CD(SD) Outbred | Charles River Laboratories | 001 | Newborn pups, P0-P2, male and female for sympathetic neuron cultures |
| Cell line (Mus musculus) | C2C12 myoblast | ATCC | CRL-1772 | siRNA knockdown pilot studies |
| Antibody | Anti-CHST11 (Rabbit polyclonal) | Invitrogen | PA5-68129 | WB (1:500) |
| Antibody | Anti-CHST15 (Rabbit polyclonal) | Proteintech | 14298-1-AP | WB (1:1000) |
| Antibody | Anti-ARSB (Rabbit polyclonal) | Proteintech | 13227-1-AP | WB (1:500) |
| Antibody | Anti-NG2 (Rabbit polyclonal) | Millipore | AB5320 | WB (1:1000) |
| Antibody | Anti-Tyrosine hydroxylase (Rabbit polyclonal) | Millipore | AB152 | WB (1:1000) IF (1:1000) |
| Antibody | Anti-Galectin-3 (Mouse monoclonal) | Abcam | AB2785 | WB (1:1000) |
| Antibody | Anti-chondroitin-4-sulfate (Mouse monoclonal) | Millipore | MAB2030 | WB (1:1000) |
| Antibody | Anti-chondroitin-6-sulfate (Mouse monoclonal) | Millipore | MAB2035 | WB (1:1000) |
| Antibody | Anti-Fibrinogen (Sheep polyclonal) | BioRad | 4400-8004 | IF (1:300) |
| Antibody | Anti-Rabbit IgG secondary (Goat polyclonal) | Molecular Probes | A-11034 | IF (1:1000) |
| Antibody | Anti-Sheep IgG secondary (Donkey polyclonal) | Molecular Probes | A-21099 | IF (1:1000) |
| Antibody | Anti-Rabbit HRP-conjugated secondary (Mouse polyclonal) | Thermo Fisher Scientific | A16104 | WB (1:10,000) |
| Antibody | Anti-Mouse HRP-conjugated secondary (Goat polyclonal) | Thermo Fisher Scientific | 31430 | WB (1:10,000) |
| Other | Ponceau S | Thermo Fisher Scientific | A40000278 | 3–5 min stain, de-stain with ddH2O |
| Peptide/ recombinant protein | PageRuler Plus prestained ladder | Thermo Fisher Scientific | 26619 | (7 μL) per well |
| Other | Poly-L-Lysine | Sigma-Aldrich | P8920 | (0.01%) for plate coating |
| Other | Laminin | RND Systems | 3400-010-02 | (1 μg/mL) |
| Other | Soluble CSPGs | Millipore | CC117 | (2 μg/mL) |
| Peptide/recombinant protein | NGF | Alomone Labs | N-100 | (10 ng/mL) |
| Peptide/recombinant protein | Recombinant ARSB | RND Systems | 4415-SU-010 | (0.6 μg/mL) |
| Peptide/recombinant protein | Recombinant Chondroitinase ABC | RND Systems | 6877 GH-020 | (100 μU/mL) |
| Chemical compound, drug | Dharmafect | Horizon Discovery | T-2001-02 | Use manufacturer’s protocol |
| Sequence-based reagent | siRNA: Chst15 | Horizon Discovery | J-059417-09 | (120 nM) – in vitro (100 μg) – in vivo Accell formulation |
| Sequence-based reagent | siRNA: Non-targeting | Horizon Discovery | D-001910–01 | (120 nM) – in vitro (100 μg) – in vivo Accell formulation |
| Sequence-based reagent | Chst15 primer | Thermo Fisher Scientific | 4331182 | Manufacturer’s protocol |
| Sequence-based reagent | Gapdh primer | Thermo Fisher Scientific | 4448489 | Manufacturer’s protocol |
| Other | C18 Column | Agilent | AG-588945-902 | (50 × 4.6 mm, 5 μm) |
| Chemical compound, drug | Isoproterenol | Sigma-Aldrich | CAS:5985-95-2 | (50 µg) |
| Chemical compound, drug | Caffeine | Sigma-Aldrich | CAS:58-08-2 | (3 mg) |
| Software, algorithm | Prism | GraphPad | Version 9 | Statistical analysis |
| Software, algorithm | ImageJ | NIH | https://imagej.net/ | Image analysis |
| Other | BZ-X 800 | Keyence | Imaging of co-culture (2×) and cardiac tissue sections (20×) | |
| Other | Incucyte | Essen Biosciences | Imaging neurite outgrowth (20×) |
Additional files
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/78387/elife-78387-transrepform1-v2.docx
-
Source data 1
Western blots.
- https://cdn.elifesciences.org/articles/78387/elife-78387-data1-v2.zip
-
Supplementary file 1
Supplemental data – additional controls and troubleshooting for siRNA studies.
- https://cdn.elifesciences.org/articles/78387/elife-78387-supp1-v2.pdf





