Rab10 regulates neuropeptide release by maintaining Ca2+ homeostasis and protein synthesis
Figures
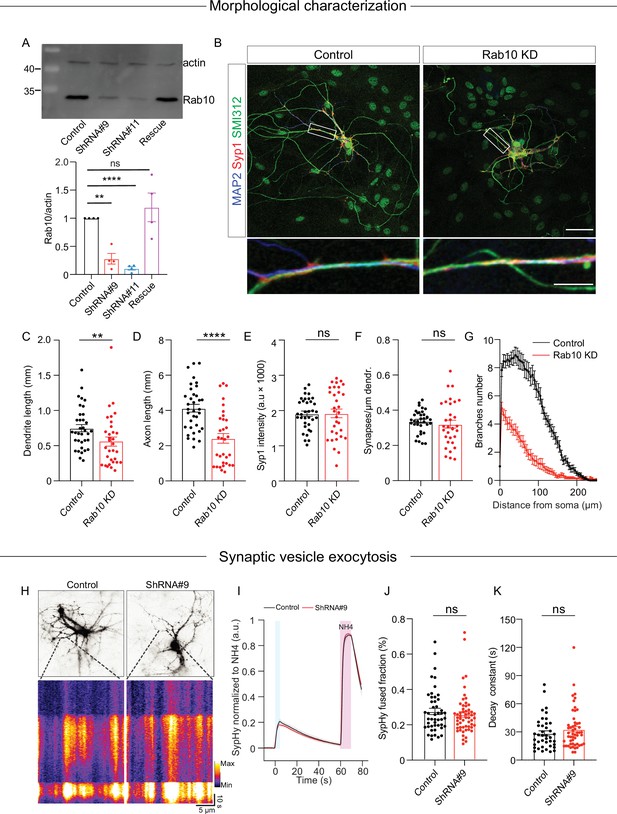
Rab10 is required for neuronal outgrowth but dispensable for synaptic vesicle (SV) exocytosis evoked by intense stimulations.
(A) Representative immunoblotting showing knockdown and rescue of Rab10 expression in cultured primary neurons infected with shRNA against Rab10 or rescue constructs (upper) and quantification of Rab10 levels (bottom). (B) Example images of control or Rab10 KD hippocampal neurons (days in vitro [DIV]14) stained for the dendrite marker MAP2 (blue), the synapse marker Syp1 (red), and the axonal marker SMI312 (green). Scale bar: 50 μm (upper) and 10 μm (bottom). (C) Quantification of the dendritic length (MAP2). (D) Quantification of the axonal length (SMI312). (E) Quantification of Syp1 intensity per synapse per neuron. (F) Quantification of the Syp1-positive synapse density in MAP2-positive dendrites. (G) Sholl analysis showing the mean number of dendritic branches against the distance from the soma. (H) Example neurons infected with the SV fusion marker SypHy (upper), typical kymographs of neurites showing SypHy intensity increase during stimulation and upon NH4Cl superfusion (bottom). (I) The average signal SypHy from active synapses, normalized from baseline to maximum fluorescence upon NH4Cl superfusion. (J) SV exocytosis determined as the ratio of the maximum SypHy intensity during stimulation to the maximum during NH4Cl stimulation. (K) SV endocytosis determined as the SypHy signal decay time constant τ in the 60 s after field stimulation. All data are plotted as mean ± s.e.m. (A) N=4, n=4, one-sample t-test. (C–G) Control: N=3, n=35; ShRNA#9: N=3, n=32. (J, K) Control: N=3, n=47; ShRNA#9: N=3, n=56. (C–F, J, K) A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, **=p<0.01, ns=not significant.
-
Figure 1—source data 1
PDF file containing original western blots for Figure 1A, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig1-data1-v1.zip
-
Figure 1—source data 2
Original files for western blot analysis displayed in Figure 1A.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig1-data2-v1.zip
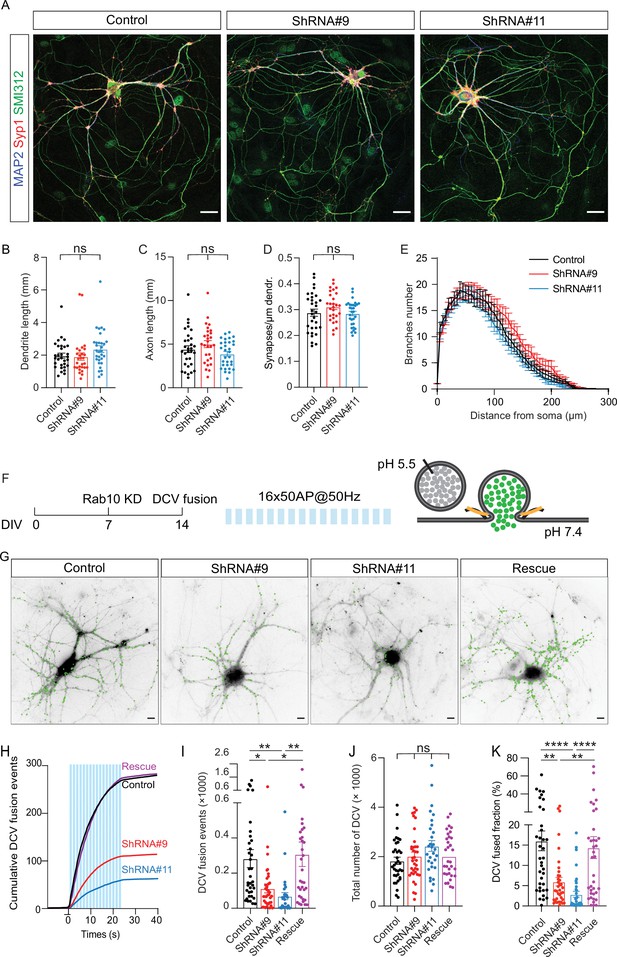
Rab10 is a major regulator of dense core vesicle (DCV) exocytosis.
(A) Example images of control and Rab10 KD hippocampal neurons (days in vitro [DIV]14) stained for MAP2 (blue), Syp1 (red), and SMI312 (green). Scale bar: 30 μm. (B) Quantification of the dendritic length (MAP2). (C) Quantification of the axonal length (SMI312). (D) Quantification of the Syp1-positive synapse density in MAP2-positive dendrites. (E) Sholl analysis showing the mean number of dendritic branches against the distance from the soma. (F) Schematic representation of DCV fusion assay. DCVs are labeled with NPY-pHluorin, and neurons are stimulated with one train of 16 bursts of 50 action potentials (APs) at 50 Hz (light blue bars). (G) Representative neurons during electrical stimulation superimposed with NPY-pHluorin fusion events (green dots). Scale bar: 5 μm. (H) Cumulative plot of DCV fusion events per cell. Light blue bars represent the stimulation trains. (I) Summary graph of DCV fusion events per cell. (J) The total number of DCVs (total pool) of neurons analyzed in H, measured as the number of NPY-pHluorin puncta upon NH4Cl perfusion. (K) Fraction of NPY-pHluorin-labeled DCV fusing during stimulation. All data are plotted as mean ± s.e.m. (B–D) Control: N=3, n=31; ShRNA#9: N=3, N=28; ShRNA#11: N=3, n=31. (I–K) Control: N=4, n=36; shRNA#9: N=4, N=37; shRNA#11: N=4, n=30; Rescue: N=4, n=34. A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, ***=p<0.001, **=p<0.01, *=p<0.05, ns=not significant.
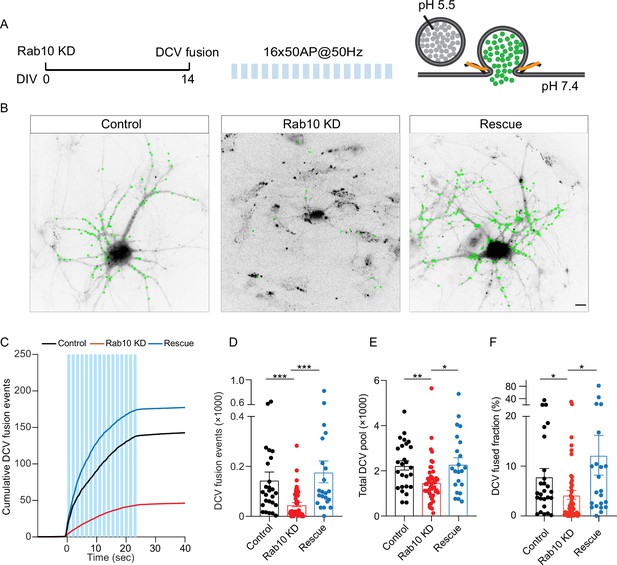
Rab10 depletion at day in vitro (DIV)0 impedes dense core vesicle (DCV) fusion.
(A) Schematic representation of DCV fusion assay. DCVs are labeled with NPY-pHluorin, and neurons are stimulated with one train of 16 bursts of 50 action potentials (APs) at 50 Hz (light blue bars). (B) Representative neurons during electrode stimulation superimposed with NPY-pHluorin fusion events (green dots). Scale bar: 10μm. (C) Cumulative plot of DCV fusion events per cell. (D) Summary graph of DCV fusion events per cell. (E) Total number of DCVs (total pool) of neurons, measured as the number of NPY-pHluorin puncta upon NH4Cl perfusion. (F) Fraction of NPY-pHluorin-labeled DCV fusing during stimulation. All data are plotted as mean ± s.e.m. (D–F) Control: N=3, n=26; Rab10 KD: N=3, n=47; Rescue: N=3, n=22. (D–F) A one-way ANOVA tested the significance of adding experimental group as a predictor. ***=p<0.001, **=p<0.01, *=p<0.05.
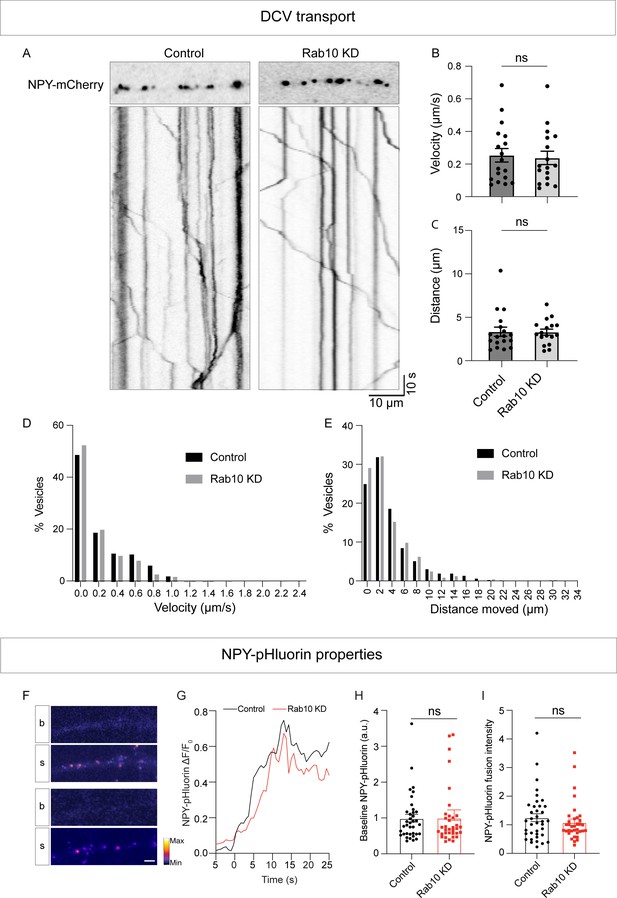
Rab10 depletion does not affect dense core vesicle (DCV) transport or cargo loading.
(A) Representative kymographs illustrating the transport of NPY-mCherry-labeled DCVs in control and Rab10 KD neurons. (B) Quantification of average velocity (µm/s) of control and Rab10 KD neurons. (C) Quantification of average distance moved from the start (µm) of control and Rab10 KD neurons. (D) Histogram of average velocity (µm/s) of control and Rab10 KD neurons. (E) Histogram of average distance moved from the start (µm) of control and Rab10 KD neurons. (F) Typical neurite expressing NPY-pHluorin during baseline (b) and during stimulation (s). Scale bar: 10μm. (G) Average traces of NPY-pHluorin fusion events aligned at the moment of fusion (0 s). (H) Quantification of NPY-pHluorin baseline fluorescence before stimulation. (I) Quantification of average NPY-pHluorin fusion intensity per cell. All data are plotted as mean ± s.e.m. (B, C) Control: N=3, n=18; Rab10 KD: N=3, n=17. (H, I) Control: N=3, n=37; Rab10 KD: N=3, n=35. A one-way ANOVA tested the significance of adding experimental group as a predictor. ns=not significant.
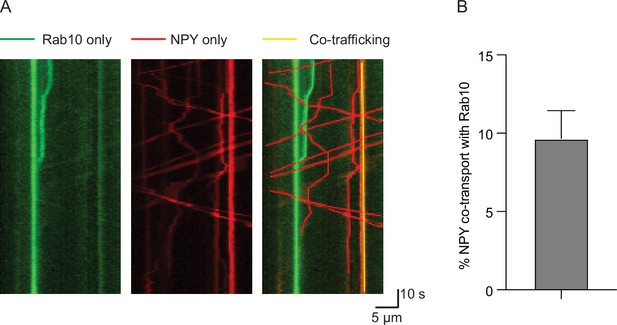
Rab10 does not typically co-transport together with dense core vesicles (DCVs).
(A) Representative kymographs of neurons co-infected with Rab10-GFP and NPY-mCherry. (B) Percentage moving DCVs that co-transport with Rab10. Data are plotted as mean ± s.e.m. (N=3, n=22). Data are plotted as mean ± s.e.m.
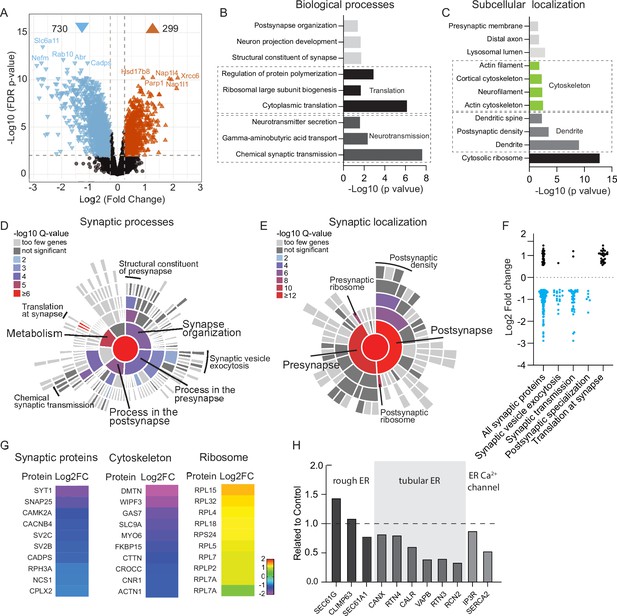
Depletion of Rab10 leads to dysregulation of proteins enriched in presynaptic transmission and cytosolic translation.
(A) Volcano plots showing significantly dysregulated proteins in Rab10-depleted neurons. (B) Gene Ontology (GO) enrichment analysis of functional pathways of the significant hits with ClueGO. Shown are the Bonferroni corrected p-values. (C) GO enrichment analysis of subcellular localization of the significant hits with ClueGO. Shown are the Bonferroni corrected p-values. (D) Sunburst plot showing the annotation in synaptic function of the altered proteins in Rab10-depleted neurons. (E) Sunburst plot showing the annotation in synaptic location of the altered proteins in Rab10-depleted neurons. (F) Log2 fold changes of synaptic proteins within SynGO terms. Downregulated proteins are shown in blue and upregulated proteins are shown in black. (G) Examples of proteins that are significantly affected by Rab10 depletion grouped by their subcellular localization. Heat maps represent the degree of up- or downregulation. (H) Selective MS data analysis of ER-related proteins in Rab10 KD neurons. Bars show the fold change of the indicated peptides compared to the control.
-
Figure 3—source data 1
Proteome analysis of neuronal cultures by mass spectrometry – complete list of proteins.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig3-data1-v1.xlsx
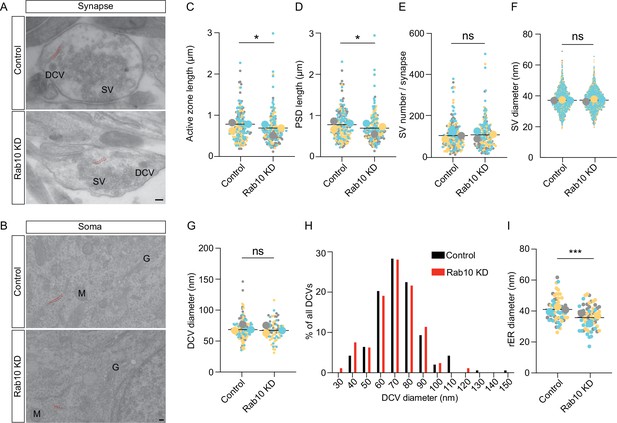
Rab10 regulates synapse size and endoplasmic reticulum (ER) morphology.
(A) Representative electron microscopy (EM) pictures showing the ultrastructure of synapses. Scale bar: 100 nm. Synaptic ER is indicated by red dotted lines. (B) Representative EM pictures showing the ultrastructure of soma. Rough ER (rER) is indicated by red dotted lines. M: mitochondrion, G: Golgi. Scale bar: 100 nm. (C) Quantification of the length of active zone and postsynaptic density (PSD). (D) Quantification of the length of PSD. (E) Quantification of synaptic vesicle (SV) number per synapse and SV diameter. (F) Quantification of SV diameter. (G) Quantification of dense core vesicle (DCV) diameter. (H) Frequency distribution of DCVs by diameter. (I) Quantification of the diameter of rER. Data are plotted with superplot (C–G, I), where averages from three independent cultures are shown as large circles and single observations are shown as dots. Horizontal lines represent the means of the averages from 3 weeks. Data from different cultures are grouped with different colors. (C–D) Control: N=3, n=184; shRNA#9: N=3, n=187. (E) Control: N=3, n=189; shRNA#9: N=3, n=188. (F) Control: N=3, n=1770; shRNA#9: N=3, n=1803. (G) Control: N=3, n=137; shRNA#9: N=3, n=122. (I) Control: N=3, n=63; shRNA#9: N=3, n=64. (C–G, I) Linear mixed model analysis. ***=p<0.001, *=p<0.05, ns=not significant.
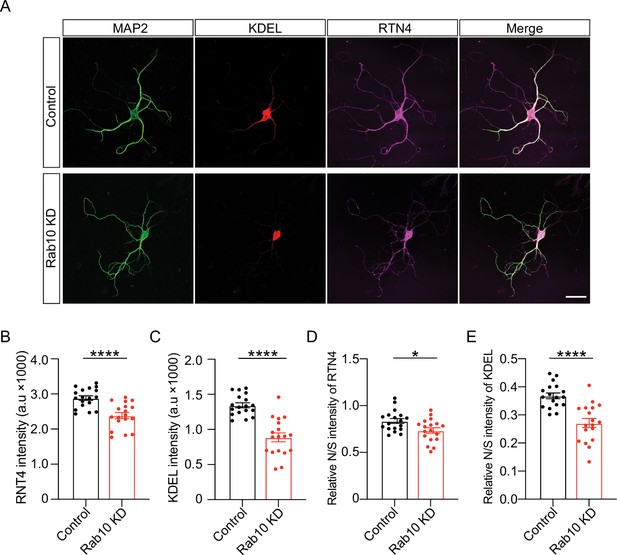
Altered endoplasmic reticulum (ER) morphology in Rab10 KD neurons.
(A) Example images of control or Rab10 KD hippocampal neurons (days in vitro [DIV]14) stained for the dendrite marker MAP2 (green), two ER markers KDEL (red) and RTN4 (magenta). Scale bar: 50 μm. (B) Quantification of RTN4 intensity in MAP2-positive dendrites. (C) Quantification of KDEL intensity in MAP2-positive dendrites. (D) The ratio of neuritic to somatic RTN4 intensity (N/S). (E) The ratio of neuritic to somatic KDEL intensity (N/S). All data are plotted as mean ± s.e.m. (B–D) Control: N=3, n=18; Rab10 KD: N=3, n=18. (B–D) A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, ***=p<0.001, **=p<0.01, *=p<0.05.
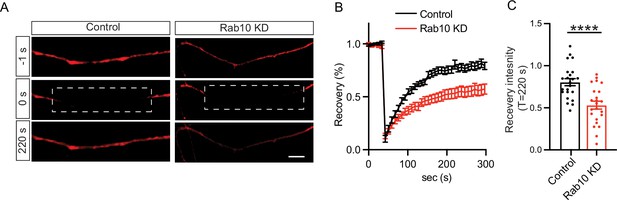
Impaired endoplasmic reticulum (ER) dynamics in Rab10 KD neurons.
(A) Representative time-lapse of ER-mCherry3 signal before (upper), upon (middle), and after (bottom) photobleaching. Scale bar: 20 μm. (B) Average normalized ER-mCherry3 fluorescence recovery after photobleaching in control and Rab10 KD hippocampal neurons. (C) Normalized ER-mCherry3 fluorescence recovery after photobleaching at T=220 s in control and Rab10 KD hippocampal neurons. All data are plotted as mean ± s.e.m. (B, C) Control: N=3, n=23; Rab10 KD: N=3, n=23. (B, C) A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001.
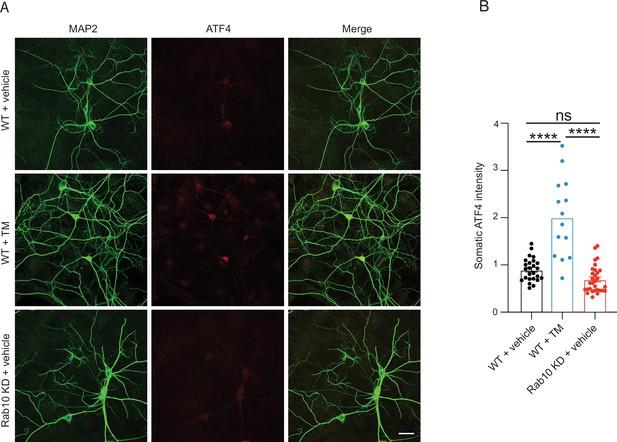
Rab10 depletion does not increase endoplasmic reticulum (ER) stress.
(A) Representative images of wild-type (WT) neurons treated with vehicle (top) or tunicamycin (TM, middle) and Rab10 KD neurons treated with vehicle (bottom). Neurons were stained for ATF4 and MAP2. Scale bar: 50μm. (B) Quantification of ATF4 intensity in soma from each condition. All data are presented as mean ± s.e.m. WT+vehicle: N=2, n=25; WT+vehicle: N=2, n=30; Rab10 KD+vehicle: N=2, n=14. A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, ns=not significant.
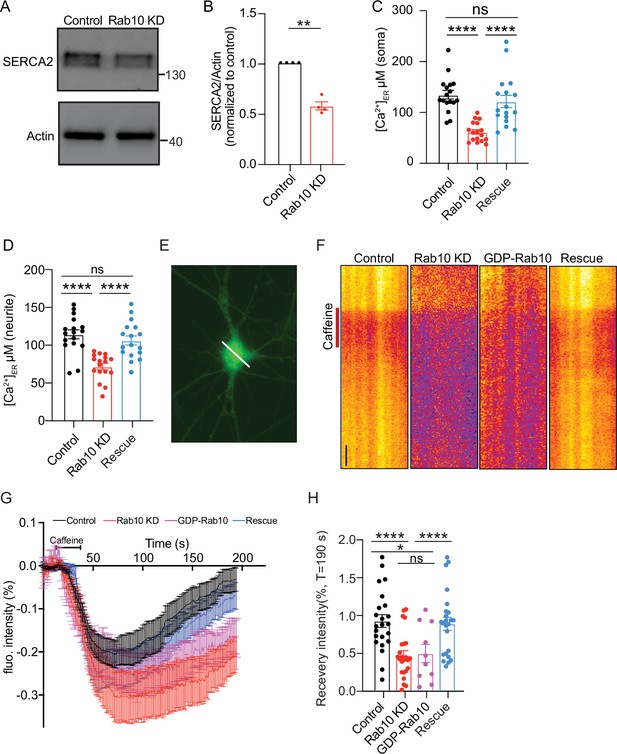
Reduced SERCA2 levels and impaired endoplasmic reticulum (ER) Ca2+ homeostasis in Rab10 KD neurons.
(A) Typical immunoblot showing reduced SERCA2 levels in Rab10 KD hippocampal neurons. (B) Quantification of protein levels in Rab10 KD neurons normalized to control. (C) Quantification of somatic ER Ca2+ concentration. (D) Quantification of dendritic ER Ca2+ concentration. (E) Representative image of a neuron infected with ER-GCaMP6-150 displayed with a pseudo line. Scale bar: 3 μm. (F) Typical kymographs of the somatic intensity of ER-GCaMP6-150 showing the intensity decrease upon caffeine superfusion (red line) and the recovery in intensity after caffeine washout. Scale bar: 10 s. (G) Average normalized ER-GCaMP6-150 fluorescence recovery after caffeine treatment. (H) Normalized ER-GCaMP6-150 fluorescence recovery after caffeine treatment at T=190 s. All data are plotted as mean ± s.e.m. (B) Control: N=4, n=4; Rab10 KD: N=4, n=4; (C-D) Control: N=3, n=17; Rab10 KD: N=3; n=17; Rescue: N=3, n=17; (H) Control: N=3, n=23; Rab10 KD: N=3; n=24; GDP-Rab10: n=3, n=10; Rescue: N=3, n=24. A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, ***=p<0.001, **=p<0.01, ns=not significant.
-
Figure 5—source data 1
PDF file containing original western blots for Figure 5A, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig5-data1-v1.zip
-
Figure 5—source data 2
Original files for western blot analysis displayed in Figure 5A.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig5-data2-v1.zip
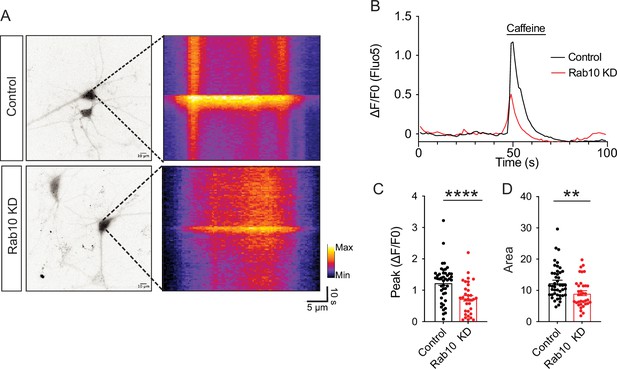
Caffeine triggers less endoplasmic reticulum (ER) Ca2+ release in Rab10 KD neurons.
(A) Left: representative cytosolic Fluo-5 AM signals upon caffeine perfusion. Right: representative kymographs of cytosolic Fluo-5 AM signals upon caffeine perfusion in somas. (B) Average traces of Fluo-5 AM signals. (C) Quantification of the peak values of the Fluo-5 AM fluorescence traces upon caffeine perfusion. (D) Quantification of the area under the curve (AUC) of the Fluo-5 AM fluorescence traces upon caffeine perfusion. All data are plotted as mean ± s.e.m. (C, D) Control: N=3, n=44; Rab10 KD: N=3, n=35. (C, D) A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, **=p<0.01.
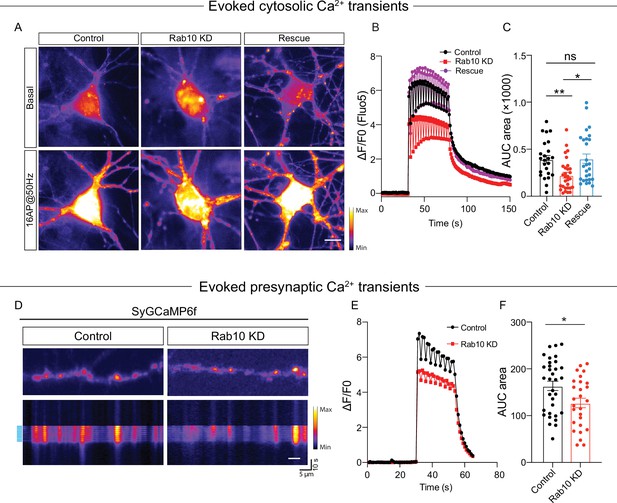
Impaired neuronal Ca2+ influx triggered by electrical stimulation.
(A) Representative time-lapse of cytosolic Fluo-5 AM upon electrical stimulation (16 action potentials [APs], 50 Hz) in somas of hippocampal neurons. Scale bar: 10 μm. (B) Average normalized response of somatic Fluo-5 AM fluorescence upon stimulation (16 APs, 50 Hz) in hippocampal neurons. (C) Quantification of the area under the curve (AUC) of the Fluo-5 AM fluorescence traces. (D) Typical neurons infected with Synaptophysin-GCaMP6 (upper), typical kymograph of a neurite (bottom) showing Synaptophysin-GCaMP6 intensity increase upon electrical stimulation (16 APs, 50 Hz, blue bars). Scale bar: 5 μm. (E) Average normalized response of Synaptophysin-GCaMP6 fluorescence intensity at presynaptic boutons upon stimulation (16 APs, 50 Hz) in hippocampal neurons. (F) Quantification of the AUC of the Synaptophysin-GCaMP6 fluorescence traces in control and Rab10 KD neurons. All data are plotted as mean ± s.e.m. (C) Control: N=4, n=24; Rab10 KD: N=4, n=30; Rescue: N=4, n=27. (F) Control: N=3, n=33; Rab10 KD: N=3; n=27. A one-way ANOVA tested the significance of adding experimental group as a predictor. **=p<0.01, *=p<0.05, ns=not significant.
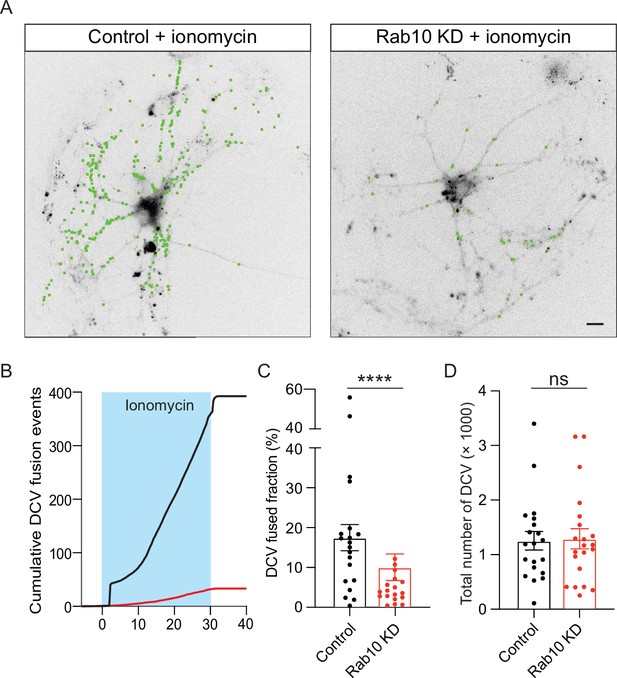
Impaired dense core vesicle (DCV) fusion induced by ionomycin in Rab10 KD neurons.
(A) Representative neurons during electrical stimulation superimposed with NPY- pHluorin fusion events (green dots). Scale bar: 10 μm. (B) Cumulative plot of DCV fusion events per cell. (C) Fraction of NPY-pHluorin-labeled DCVs fusing during stimulation. (D) The total number of DCVs (total pool) of neurons analyzed in B, measured as the number of NPY-pHluorin puncta upon NH4Cl perfusion. All data are plotted as mean ± s.e.m. (C, D) Control: N=3, n=20; Rab10 KD: N=3, n=21. (C, D) A one-way ANOVA tested the significance of adding experimental group as a predictor. *=p<0.05, ns=not significant.

Leucine supplementation ameliorates the deficits in protein synthesis and neuropeptide release in Rab10 KD neurons.
(A) Representative western blot showing puromycinilated proteins as a measure for de novo protein synthesis in each condition. (B) Quantification of puromycin intensity in each condition. (B) Representation of the dense core vesicle (DCV) fusion assay. Leucine (5 μM) was added to the culture media and incubated for 72 hr before DCV fusion assay. DMSO (1‰) was used as a control. (C) Cumulative plot of DCV fusion events per cell. (D) Fraction of NPY-pHluorin-labeled DCVs fusing during stimulation. (E) The total number of DCVs (total pool) of neurons analyzed in D, E, measured as the number of NPY-pHluorin puncta upon NH4Cl perfusion. All data are plotted as mean ± s.e.m. (B) All: N=3, n=3; (E, F) Control: N=3, n=47; Control+leu: N=3, n=45; Rab10 KD: N=3; n=61; Rab10+leu: N=3, n=54. Rab10 KD+Rab10: N=3, n=24. (B) One-sample t-test. (E, F) A one-way ANOVA tested the significance of adding experimental group as a predictor. **=p<0.01, *=p<0.05, ns=not significant.
-
Figure 8—source data 1
PDF file containing original western blots for Figure 8A, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig8-data1-v1.zip
-
Figure 8—source data 2
Original files for western blot analysis displayed in Figure 8A.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig8-data2-v1.zip
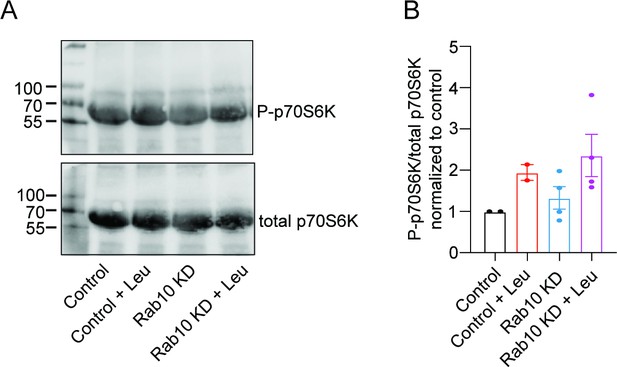
Rab10 depletion does not upregulate mTORC1 pathway.
(A) Typical immunoblot showing pS6K1 levels in each condition. (B) Quantification of relative pS6K1 levels in each condition. All data are plotted as mean ± s.e.m. (C) Control, Control+Leu: N=2, n=2, Rab10 KD, Rab10 KD+Leu: N=2, n=4.
-
Figure 8—figure supplement 1—source data 1
PDF file containing original western blots for Figure 8—figure supplement 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig8-figsupp1-data1-v1.zip
-
Figure 8—figure supplement 1—source data 2
Original files for western blot analysis displayed in Figure 8—figure supplement 1.
- https://cdn.elifesciences.org/articles/94930/elife-94930-fig8-figsupp1-data2-v1.zip
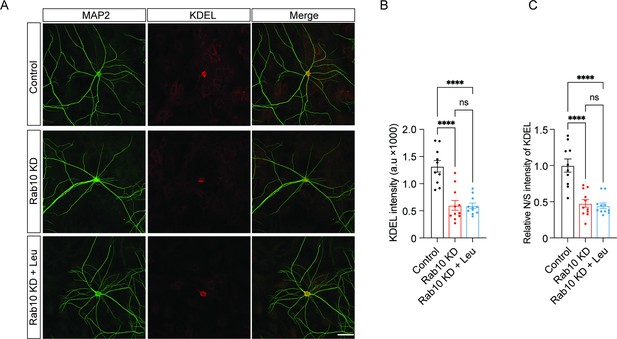
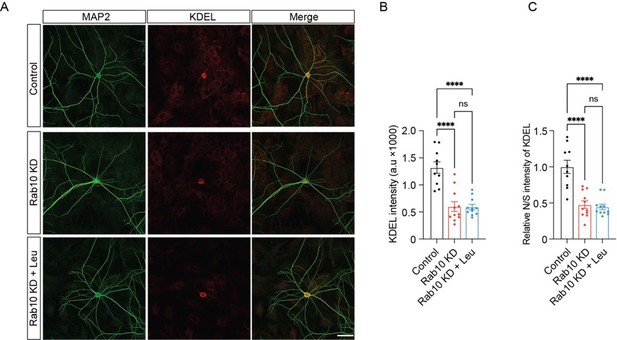
Leucine supplementation does not rescue endoplasmic reticulum (ER) morphological deficiency in Rab10 KD neurons.
(A) Typical examples showing the KDEL signals in each condition. Scale bar: 50 μm. (B) Quantification of RTN4 intensity in MAP2-positive dendrites. (C) The ratio of neuritic to somatic RTN4 intensity (N/S). All data are plotted as mean ± s.e.m. (B, C) Control: N=3, n=10; Rab10 KD: N=3, n=11; Rab10 KD+Leu: N=3; n=11. A one-way ANOVA tested the significance of adding experimental group as a predictor. ****=p<0.0001, ns=not significant.
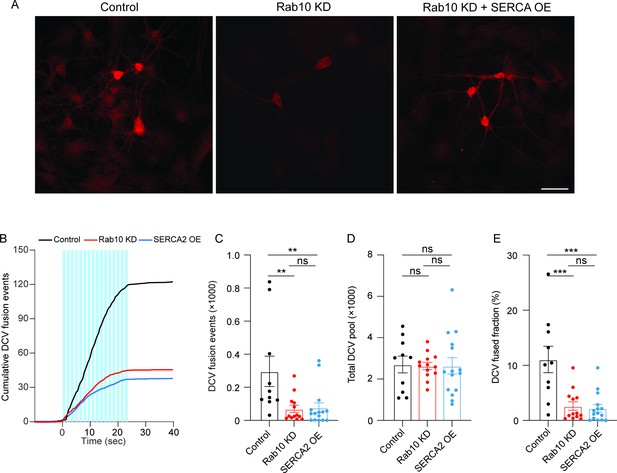
Overexpression of SERCA2 does not rescue the dense core vesicle (DCV) fusion deficits in Rab10 KD neurons.
(A) Typical examples showing the SERCA2 signals in each condition. Scale bar: 50 μm. (B) Cumulative plot of DCV fusion events per cell. (C) Summary graph of DCV fusion events per cell. (D) Total number of DCVs (total pool) of neurons, measured as the number of NPY-pHluorin puncta upon NH4Cl perfusion. (E) Fraction of NPY-pHluorin-labeled DCV fusing during stimulation. All data are plotted as mean ± s.e.m. (C–E) Control: N=2, n=10; Rab10 KD: N=2, n=13; SERCA2 OE: N=2; n=15. A one-way ANOVA tested the significance of adding experimental group as a predictor. ***=p<0.001, **=p<0.01, ns=not significant.
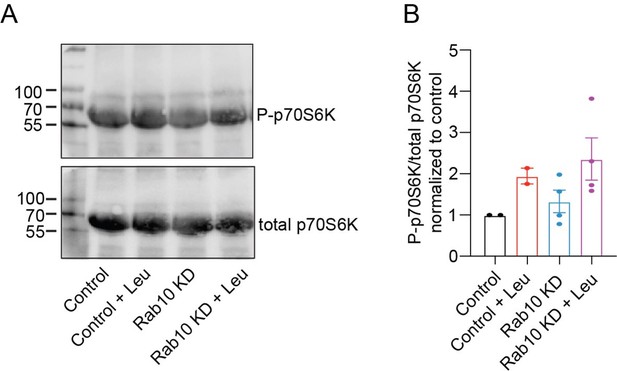
Rab10 depletion does not upregulate mTORC1 pathway.
(A)Typical immunoblot showing pS6K1 levels in each condition. (B) Quantification of relative pS6K1 levels in each condition. All Data are plotted as mean± s.e.m. (C) Control, Control + Leu: N = 2, n = 2, Rab10 KD, Rab10 KD + Leu: N = 2, n = 4.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Mus musculus) | Rab10 | NCBI | 74173 | |
| Genetic reagent (Mus musculus) | C57BL/6J | Charles River | Strain code 631 | |
| Genetic reagent (Rattus norvegicus) | Wistar (Crl:WI) | Charles River | Strain code 003 | |
| Antibody | MAP2 (chicken polyclonal) | Abcam | ab5392 RRID:AB_2138153 | 1:200 (IF) |
| Antibody | SMI312 (mouse polyclonal) | Eurogentec | SMI-312P-050 | 1:500 (IF) |
| Antibody | Synaptophysin 1 (guinea pig polyclonal) | Synaptic Systems | 101004 RRID:AB_1210382 | 1:500 (IF) |
| Antibody | KDEL (mouse monoclonal) | Enzo Life Sciences | ADI-SPA-827-D RRID:AB_2039327 | 1:200 (IF) |
| Antibody | RTN4 (rabbit polyclonal) | Novus Biologicals | NB100-56681 RRID:AB_838641 | 1:200 (IF) |
| Antibody | SERCA2 (mouse monoclonal) | Santa Cruz | sc-376235 RRID:AB_10989947 | 1:200 (IF) |
| Antibody | Rab10 (rabbit polyclonal) | Protein Tech | 11808-1-AP RRID:AB_2173442 | 1:2000 (WB) |
| Antibody | Rab10 (mouse monoclonal) | Abcam | Ab104859 RRID:AB_10711207 | 1:2000 (WB) |
| Antibody | Actin (mouse monoclonal) | Chemicon | MAB1501 RRID:AB_2223041 | 1:4000 (WB) |
| Antibody | Puromycin (mouse monoclonal) | Bio Connect | MABE343 RRID:AB_2566826 | 1:2500 (WB) |
| Antibody | Phospho-p70 S6 Kinase (rabbit monoclonal) | Cell Signaling Technology | 9234S RRID:AB_2269803 | 1:1000 (WB) |
| Antibody | p70 S6 kinase (rabbit polyclonal) | Cell Signaling Technology | 9202S RRID:AB_331676 | 1:1000 (WB) |
| Transfected construct (Mus musculus) | shRNA#9 | This paper | – | Lentiviral construct to transfect and express the shRNA (see Materials and methods) |
| Transfected construct (Mus musculus) | shRNA#11 | This paper | – | Lentiviral construct to transfect and express the shRNA (see Materials and methods) |
| Transfected construct (Mus musculus) | Control | This paper | – | Lentiviral construct to transfect and express the control (see Materials and methods) |
| Recombinant DNA reagent | pLenti-Syn(pr)-NPY-pHluorin | PMID:31679900 | – | – |
| Recombinant DNA reagent | pLenti-Syn(pr)-NPY-mCherry | PMID:31679900 | – | – |
| Recombinant DNA reagent | pLenti-Syn(pr)-Synaptophysin-pHluorin | PMID:34020952 | – | – |
| Recombinant DNA reagent | pLenti-Syn(pr)-Synaptophysin-GCaMP6 | This paper | – | Generation of this reagent is described in Materials and methods |
| Recombinant DNA reagent | ER-GCaMP6-150 | Addgene | RRID:Addgene_86918 | – |
| Recombinant DNA reagent | mCherry-ER3 | Addgene | RRID:Addgene_55041 | – |
| Recombinant DNA reagent | EGFP-Rab10T23N | Addgene | RRID:Addgene_86918 | – |
| Peptide, recombinant protein | pLenti-Syn(pr)- Rab10-EGFP | This paper | – | Generation of this reagent is described in Materials and methods |
| Peptide, recombinant protein | 2.5% trypsin | Gibco | 15090046 | – |
| Peptide, recombinant protein | Poly-L-ornithine | Worthington Biochemical Corporation | LS003127 | – |
| Peptide, recombinant protein | Laminin | Sigma-Aldrich | L2020 | – |
| Peptide, recombinant protein | Poly-D-lysine | Sigma-Aldrich | P6407 | – |
| Peptide, recombinant protein | L-Leucine | Sigma-Aldrich | L8000 | – |
| Peptide, recombinant protein | Tunicamycin | Sigma-Aldrich | T7765-10MG | – |
| Chemical compound, drug | Puromycin | Merck/Millipore | 540222-25MG | – |
| Chemical compound, drug | Ionomycin | Fisher Emergo | 10429883 | – |
| Chemical compound, drug | TCE | Sigma-Aldrich | 115-20-8 | – |
| Software, algorithm | MATLAB | MathWorks | RRID:SCR_001622 | – |
| Software, algorithm | Prism | GraphPad | RRID:SCR_002798 | – |
| Other | Fiji/ImageJ | NIH | RRID:SCR_002285 | – |
-
WB: western blot; IF: immunofluorescence.
Summary of statistical analyses.
| Figure | Dataset | Groups | n-number* | Statistical test | p-value |
|---|---|---|---|---|---|
| 1 A | Band intensity of Rab10 | Control ShRNA#9 ShRNA#11 Rescue | 4 cultures | One sample t-test (compare to 100%) | PshRNA#9=0.0046 (**) PshRNA#11<0.0001 (****) Prescue =0.5034 (ns) |
| 1 C | Dendritic length (MAP2) | Control ShRNA#9 | 3 (35) 3 (32) | ANOVA model comparison for nested linear models | P=0.0093 (**) |
| 1D | Axonal length (SMI312) | Control ShRNA#9 | 3 (35) 3 (32) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 1E | Syp1 intensity per synapse per neuron | Control ShRNA#9 | 3 (35) 3 (32) | ANOVA model comparison for nested linear models | P=0.4975 (ns) |
| 1 F | Syp1-positive synapse density in MAP2-positive dendrites | Control ShRNA#9 | 3 (35) 3 (32) | ANOVA model comparison for nested linear models | P=0.4975 (ns) |
| 1 J | SypHy fused fraction | Control ShRNA#9 | 3 (47) 3 (56) | ANOVA model comparison for nested linear models | P=0.9496 (ns) |
| 1 K | Decay content | Control ShRNA#9 | 3 (47) 3 (56) | ANOVA model comparison for nested linear models | P=0.2910 (ns) |
| 2B | Dendritic length (MAP2) | Control ShRNA#9 ShRNA#11 | 3 (31) 3 (28) 3 (31) | One-way ANOVA | P=0.1818 (ns) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.9771 (ns); pControl vs ShRNA#11=0.3004 (ns); p ShRNA#9 vs ShRNA#11=0.2276 (ns); | ||||
| 2 C | Axonal length (SMI312) | Control ShRNA#9 ShRNA#11 | 3 (31) 3 (28) 3 (31) | One-way ANOVA | P=0.0936 (ns) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.5037 (ns); pControl vs ShRNA#11=0.5313 (ns); p ShRNA#9 vs ShRNA#11=0.0823 (ns); | ||||
| 2D | Syp1-positive synapse density in MAP2-positive dendrites | Control ShRNA#9 ShRNA#11 | 3 (31) 3 (28) 3 (31) | One-way ANOVA | P=0.2126 (ns) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.3405 (ns); pControl vs ShRNA#11=0.9788 (ns); p ShRNA#9 vs ShRNA#11=0.2503 (ns); | ||||
| 2I | DCV fusion events/neuron | Control ShRNA#9 ShRNA#11 Rescue | 3 (36) 3 (37) 3 (30) 3 (34) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.0450 (*); pControl vs ShRNA#11=0.0105 (**); p ShRNA#11vs Rescue=0.0021 (**); pShRNA#9 vs Rescue=0.0100 (*); | ||||
| 2 J | Total DCV pool/neuron | Control ShRNA#9 ShRNA#11 Rescue | 3 (36) 3 (37) 3 (30) 3 (34) | One-way ANOVA | P=0.1014 (ns) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.7669 (ns); pControl vs ShRNA#11=0.0584 (ns); p ShRNA#11vs Rescue=0.4978 (ns); pShRNA#9 vs Rescue=0.9969 (ns); | ||||
| 2 K | DCV fusion fraction | Control ShRNA#9 ShRNA#11 Rescue | 3 (36) 3 (37) 3 (30) 3 (34) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs ShRNA#9=0.0014 (**); pControl vs ShRNA#11=0.0001 (****); pControl vs Rescue=0.9902 (ns); pShRNA#9 vs Rescue>0.0048 (**); | ||||
| 2 suppl 1D | DCV fusion events/neuron | Control Rab10 KD Rescue | 3 (26) 3 (47) 3 (22) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.001 (***); pControl vs Rescue>0.9999 (ns); pRab10 KD vs Rescue=0.0008 (***); | ||||
| 2 suppl 1E | Total DCV pool/neuron | Control Rab10 KD Rescue | 3 (26) 3 (47) 3 (22) | One-way ANOVA | P=0.0021 |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0098(**); pControl vs Rescue=0.9699 (ns); pRab10 KD vs Rescue=0.0138 (*); | ||||
| 2 suppl 1 F | DCV fusion fraction | Control Rab10 KD Rescue | 3 (26) 3 (47) 3 (22) | One-way ANOVA | P<0.002 (**) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0435 (*); pControl vs Rescue=0.6189 (ns); pRab10 KD vs Rescue=0.0031 (**); | ||||
| 2 suppl 2B | DCV transport velocity | Control Rab10 KD | 3 (18) 3 (17) | ANOVA model comparison for nested linear models | P=0.8028(ns) |
| 2 suppl 2 C | DCV transport distance | Control Rab10 KD | 3 (18) 3 (17) | ANOVA model comparison for nested linear models | P=0.9131 (ns) |
| 2 suppl 2 H | Baseline NPY-phluorin intensity | Control Rab10 KD | 3 (37) 3 (35) | ANOVA model comparison for nested linear models | P=0.2734 (ns) |
| 2 suppl 2I | NPY-phluorin fusion intensity | Control Rab10 KD | 3 (37) 3 (35) | ANOVA model comparison for nested linear models | P=0.3385 (ns) |
| 4 C | Active zone length | Control Rab10 KD | 3 cultures | Linear mixed model | P=0.023 (*) |
| 4D | PSD length | Control Rab10 KD | 3 cultures | Linear mixed model | P=0.020 (*) |
| 4E | SV number per synapse | Control Rab10 KD | 3 cultures | Linear mixed model | P=0.746 (ns) |
| 4 F | SV diameter | Control Rab10 KD | 3 cultures | Linear mixed model | P=0.612 (ns) |
| 4 G | DCV diameter | Control Rab10 KD | 3 cultures | Linear mixed model | P=0.260 (ns) |
| 4I | rER diameter | Control Rab10 KD | 3 cultures | Linear mixed model | P<0.001 (***) |
| 4 suppl 1B | RTN4 intensity | Control Rab10 KD | 3 (18) 3 (18) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 4 suppl 1 C | KDEL intensity | Control Rab10 KD | 3 (18) 3 (18) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 4 suppl 1D | Relative N/S intensity of RTN4 | Control Rab10 KD | 3 (18) 3 (18) | ANOVA model comparison for nested linear models | P=0.01551 (*) |
| 4 suppl 1E | Relative N/S intensity of KDEL | Control Rab10 KD | 3 (18) 3 (18) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 4 suppl 2 C | Recovery intensity of mCherry-ER3 after photobleaching at T=220 s | Control Rab10 KD | 3 (23) 3 (23) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 4 suppl 3B | ATF4 intensity | Control Rab10 KD TM | 2 (25) 2 (30) 2 (14) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.1874 (ns); pControl vs TM<0.0001 (****); pRab10 KD vs TM<0.0001 (****); |
| 5B | Band intensity of SERCA2 | Control Rab10 KD | 4 cultures | One sample t-test (compare to 100%) | P=0.0017 (**) |
| 5 C | Somatic ER Ca2+ | Control Rab10 KD Rescue | 3 (17) 3 (17) 3 (17) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD<0.0001 (****); pControl vs Rescue>0.5242 (ns); pRab10 KD vs Rescue<0.0001 (****); | ||||
| 5D | Neuritic ER Ca2+ | Control Rab10 KD Rescue | 3 (17) 3 (17) 3 (17) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD<0.0001 (****); pControl vs Rescue>0.5360 (ns); pRab10 KD vs Rescue<0.0001 (****); | ||||
| 5 H | Recovery intensity of Fluo-5 AM | Control Rab10 KD GDP-Rab10 Rescue | 3 (23) 3 (24) 3 (10) 3 (24) | One-way ANOVA | P<0.0002 (***) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0005 (****); pControl vs Rescue>0.9999 (ns); pRab10 KD vs Rescue=0.0013 (****); pControl vs GDP-Rab10=0.0307 (*); | ||||
| 5 suppl 1 C | ER Ca2+ release triggered by caffeine (peak) | Control Rab10 KD | 3 (44) 3 (35) | ANOVA model comparison for nested linear models | P<0.0001 (****) |
| 5 suppl 1D | ER Ca2+ release triggered by caffeine (area) | Control Rab10 KD | 3 (44) 3 (35) | ANOVA model comparison for nested linear models | P=0.0025 (**) |
| 6 C | Evoked cytosolic Ca2+ influx | Control Rab10 KD Rescue | 3 (24) 3 (30) 3 (27) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0062 (**); pControl vs Rescue=0.9891 (ns); pRab10 KD vs Rescue=0.0128 (*); |
| 6 F | Evoked presynaptic Ca2+ influx | Control Rab10 KD | 3 (33) 3 (27) | ANOVA model comparison for nested linear models | P=0.0146 (*) |
| 7 C | Ionomycin-induced DCV fused fraction | Control Rab10 KD | 3 (20) 3 (21) | ANOVA model comparison for nested linear models | P=0.0009 (****) |
| 7D | Total DCV pool/neuron | Control Rab10 KD | 3 (20) 3 (21) | ANOVA model comparison for nested linear models | P=0.8821 (ns) |
| 8B | Band intensity of puromycin | Control Rab10 KD Rab10T23N KD +Leucine | 3 cultures | One sample t-test (compare to 100%) | PRab10 KD=0.0354 (*) PRab10 T23N=0.0053 (**) pKD+Leucine=0.1486 (ns) |
| 8E | DCV fused fraction | Control Control +Leu Rab10 KD Rab10+Leu Rab10 KD +Rab10 | 3 (47) 3 (45) 3 (61) 3 (54) 3 (24) | One-way ANOVA | P<0.0001 (****) |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD<0.0001 (****); pControl + Leu vs Rab10 KD<0.0001 (****); pRab10 KD vs Rab10 KD + Rab10<0.0001 (****); pRab10 KD vs Rab10 KD + Leu<0.0001 (****); pcontrol vs Rab10 KD + Leu=0.577 (ns) | ||||
| 8 F | Total DCV pool/neuron | Control Control +Leu Rab10 KD Rab10+Leu Rab10 KD +Rab10 | 3 (47) 3 (45) 3 (61) 3 (54) 3 (24) | One-way ANOVA | P=0.1035 |
| ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.2484 (ns); pControl + Leu vs Rab10 KD>0.9999 (****); pRab10 KD vs Rab10 KD + Rab10>0.9999 (ns); pRab10 KD vs Rab10 KD + Leu>0.9999 (ns); pcontrol vs Rab10 KD + Leu>0.9999 (ns) | ||||
| 8 suppl 2B | KDEL intensity | Control Rab10 KD Rab10+Leu | 3 (10) 3 (11) 3 (11) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD<0.0001 (****); pcontrol vs Rab10 KD + Leu<0.0001 (****); pRab10 KDvs Rab10 KD + Leu=0.9970(ns); |
| 8 suppl 2 C | Relative N/S intensity of KDEL | Control Rab10 KD Rab10+Leu | 3 (10) 3 (11) 3 (11) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD<0.0001 (****); pcontrol vs Rab10 KD + Leu<0.0001 (****); pRab10 KDvs Rab10 KD + Leu=0.9293(ns); |
| 8 suppl 3 C | DCV fusion events/neuron | Control Rab10 KD SERCA2 | 2 (10) 2 (13) 2 (15) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0084 (**); pControl vs SERCA2 =0.0095 (**); prab10 KD vs SERCA2 =0.0095 (**); |
| 8 suppl 3D | Total DCV pool/neuron | Control Rab10 KD SERCA2 | 2 (10) 2 (13) 2 (15) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.9988 (ns); pControl vs SERCA2 =0.9813 (ns); prab10 KD vs SERCA2 =0.9655 (ns); |
| 8 suppl 3E | DCV fused fraction | Control Rab10 KD SERCA2 | 2 (10) 2 (13) 2 (15) | ANOVA model comparison for nested linear models | pControl vs Rab10 KD=0.0003 (***); pControl vs SERCA2 =0.0001 (****); prab10 KD vs SERCA2 =0.9711 (ns); |