Compositional editing of extracellular matrices by CRISPR/Cas9 engineering of human mesenchymal stem cell lines
Figures
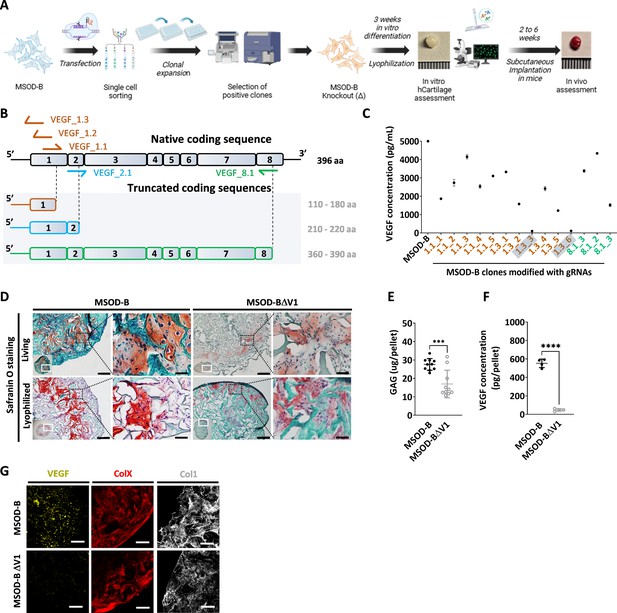
CRISPR/Cas9 editing of mesenchymal cell lines lead to efficient vascular endothelial growth factor (VEGF) knockout in cartilage tissues.
(A) Experimental scheme depicting the generation of CRISPR/Cas9-edited mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) lines, and the subsequent in vitro and in vivo tissue formation assessment. (B) Overview of the native human VEGF coding sequence composed of eight exons. Designed guide RNAs (gRNAs) and their targeted binding sites are illustrated, as well as the corresponding expected impact on the coding sequence. gRNAs targeting exon 1 (orange) disrupt translation initiation and inhibit protein expression, gRNA targeting exon 2 (blue) disrupts VEGF receptor binding, and gRNA targeting exon 8 (green) alters the C-terminal sequence and represses activation of protein. (C) ELISA-based quantitative analysis of VEGF protein content in cell culture supernatant from expanded single cell colonies. From all clones, only two had no detectable level of VEGF (1.3_3 and 1.3_6). These clones were subsequently defined as MSOD-BΔV1 and MSOD-BΔV2. (D) Histological assessment of living and lyophilized in vitro differentiated constructs using Safranin O staining (scale bars=100 µm and 20 µm for magnified areas). Both the MSOD-B and MSOD-BΔV1 displayed glycosaminoglycans (GAG) (orange to reddish in Safranin O), indicating successful cartilage formation. Left bottom inserts show the whole tissue section. (E) Quantitative assessment of the total GAG content in MSOD-B and MSOD-BΔV1 in vitro differentiated constructs, post-lyophilization. Unpaired t-test, n=10 biological replicates, ***p<0.001. (F) ELISA-based quantitative assessment of VEGF protein in in vitro differentiated constructs, post-lyophilization. Unpaired t-test, n=3–4 biological replicates, ****p<0.0001. (G) Immunofluorescence images of MSOD-B and MSOD-BΔV1 tissues, post-lyophilization. Displayed images consist of 3D-stacks from 80- to 100-µm-thick sections, stained for VEGF (yellow), Collagen Type X (COLX, red), and Collagen Type I (COL1, gray). A clear reduction in the VEGF signal could be observed in the MSOD-BΔV1 tissues, indicating a successful VEGF knockdown (scale bars=80 µm). All error bars in the figures indicate the standard deviation (SD). Panel A was created with BioRender.com.
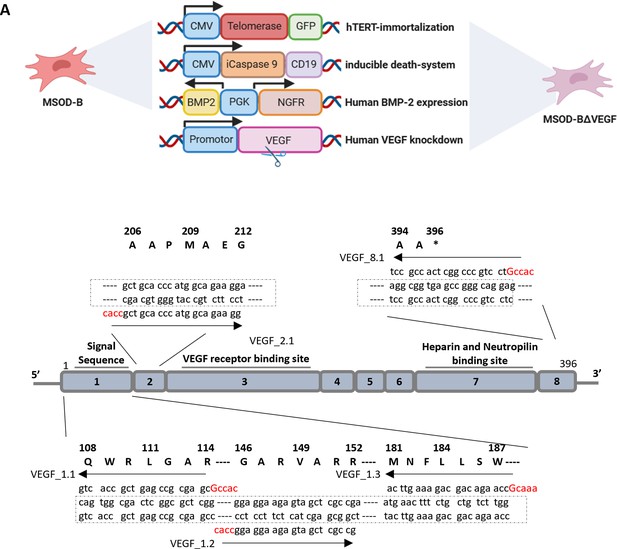
Implemented genetic elements of the mesenchymal sword of Damocles vascular endothelial growth factor (VEGF) (MSOD-BΔV) cells consisting of the human telomerase reverse transcriptase (hTERT), an inducible caspase 9 (iCaspase) death system, the bone morphogenetic protein type-2 and VEGF knockdown and the overview of the human VEGF exon structure, nucleotide sequences of targeting guide RNAs (gRNAs) and their corresponding amino acid sequences. Panel A was created with BioRender.com.
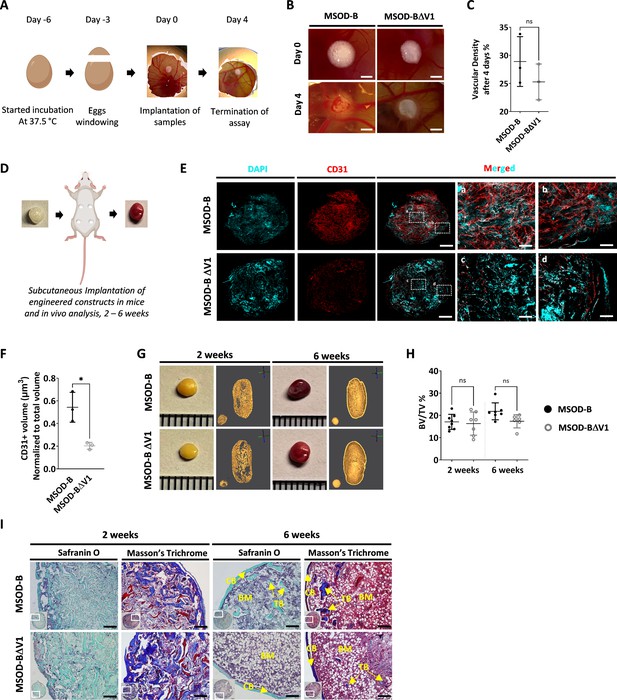
Vascular endothelial growth factor (VEGF) knockout cartilage tissues retain bone remodeling capacity despite reduced early-stage vascularization.
(A) Experimental scheme of the chorioallantoic membrane (CAM) assay, for ex vivo evaluation of angiogenic potential. (B) Macroscopic comparison of in vitro mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and MSOD-BΔV1 constructs at day 0 (day of implantation) and day 4 (4 days postimplantation), illustrating robust de novo vessel formation perfusing the tissues (scale bars at 1 mm). (C) Quantitative analysis of vascular density in MSOD-B and MSOD-BΔV1. Vascular densities were quantified from macroscopic images obtained on day 4 using ImageJ. Unpaired t-test, n=3–4 biological replicates, n.s.=not significant. (D) Overview of the subcutaneous implantation procedure in mice and subsequent in vivo evaluation of vascularization from 2 to 6 weeks postimplantation. (E) Immunofluorescence images of MSOD-B and MSOD-BΔV1 tissues 2 weeks post-in vivo implantation. Displayed images consist of 3D-stacks from 80 to 100-µm-thick sections, vessels stained with mouse CD31 (red) and nuclei with DAPI (cyan) (scale bars (for DAPI, CD31 and MERGED) at 500 µm except for magnified white inserts at 80 µm). Box ‘a’ and ‘c’ display the periphery whereas Box ‘b’ and ‘d’ show the central region of MSOD-B and MSOD-BΔV1 constructs respectively. A reduction in tissue vascularization is observed in MSOD-BΔV1 samples. (F) Quantitative analysis of the CD31 signal using an isosurface-based strategy (IMARIS software). Unpaired t-test, n=3–4 biological replicates, *p<0.05. (G) Representative macroscopic and microtomography images of in vivo constructs retrieved at 2 and 6 weeks postimplantation. (H) Microtomography-based quantification of the sample’s bone/ mineralized volume over their total volume (ratio). No significant differences between MSOD-B and MSOD-BΔV1 could be observed (BV: bone volume, TV: total volume). Ordinary one-way ANOVA, n=8 biological replicates, n.s.=not significant. (I) Histological analysis of in vivo tissues using Safranin O and Masson’s trichrome stains, at 2 (2W) and 6 weeks (6W) postimplantation. Both sample types underwent full remodeling into a bone organ after 6 weeks, with presence of bone structures and a bone marrow compartment (scale bars=200 µm). CB – cortical bone; BM – bone marrow; TB – trabecular bone. All error bars in the figures indicate the standard deviation (SD).
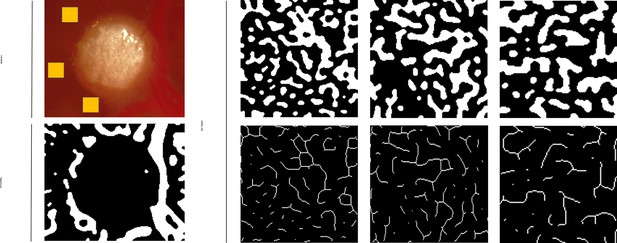

Sequential processing of a vascular tissue section using Q-VAT, depicting the original image, binary mask creation, and vessel segmentation for quantitative vascular density analysis.
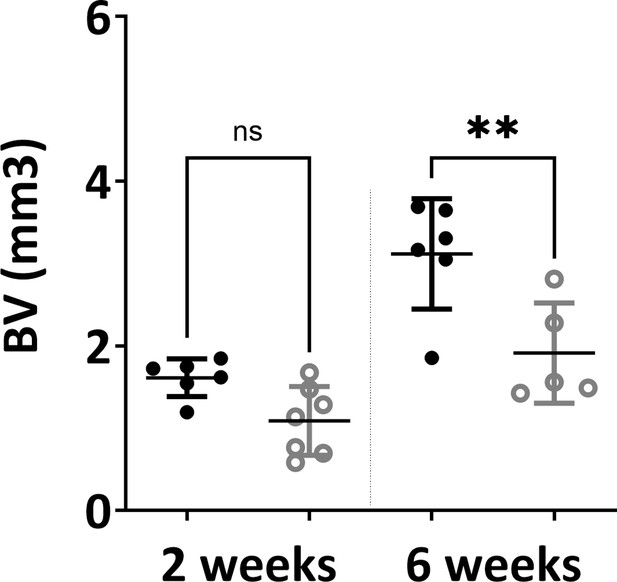
Microtomography analysis (bone/mineralized volume) showed significant differences between in vivo constructs at both 2 weeks and 6 weeks time points (one-way ANOVA, n=3, **p<0.01, n.s.=not significant).
The error bars in the figures indicate the standard deviation (SD).
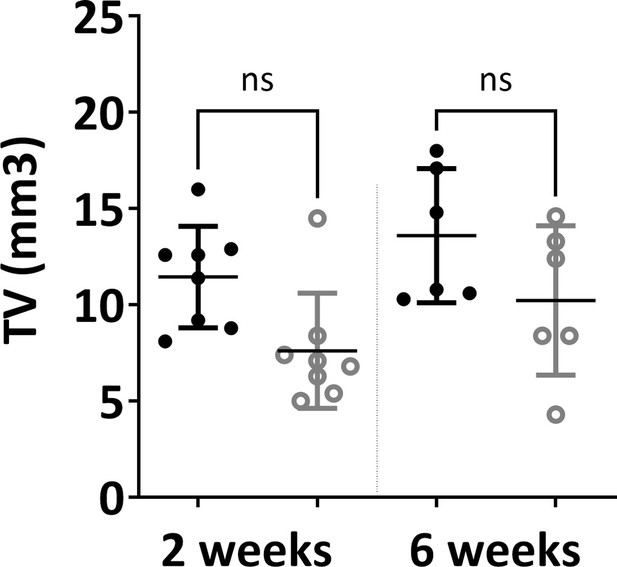
Microtomography analysis (total volume) did not show significant difference between in vivo constructs at either 2 weeks or 6 weeks time points (one-way ANOVA, n=3, n.s.=not significant) (BV: bone volume, TV: total volume).
The error bars in the figures indicate the standard deviation (SD).
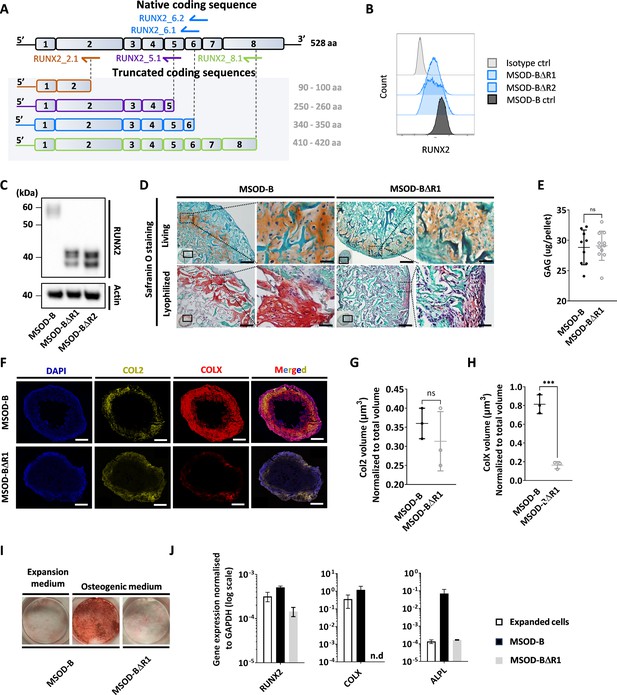
Runt-related transcription factor 2 (RUNX2) knockout does not prevent chondrogenic differentiation but impairs hypertrophy.
(A) Overview of the human RUNX2 coding sequence comprising eight exons. Guide RNAs (gRNAs) and their corresponding expected protein structure. The gRNA targeting exon 2 (orange) disrupts the DNA binding domain, gRNA targeting exon 5 (violet) disrupts nuclear translocation, gRNAs targeting exon 6 (blue) disrupt the transcriptional activation domain, and gRNAs targeting exon 8 (green) disrupt the nuclear matrix targeting signal and repress the protein function. (B) Intracellular flow cytometry for RUNX2 detection in mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and RUNX2-edited clones. A clear protein reduction could be observed in the 6.1_1 and 6.1_23 clones. (C) Western blot analysis of RUNX2 in cultured MSOD-B and RUNX2-edited cells. The genetic editing of RUNX2 is confirmed by the detection of the truncated proteins. Actin is used as a control to normalize the protein content. (D) Histological analysis of living and lyophilized in vitro differentiated constructs using Safranin O staining (scale bars=100 µm and 20 µm for magnified areas), indicating the presence of cartilage matrices. (E) Quantitative assessment of the total GAG content in corresponding in vitro generated lyophilized tissues. Unpaired t-test, n=10–11 biological replicates **p<0.01. (F) Immunofluorescence images of MSOD-B and MSOD-BΔR1 lyophilized tissues. Displayed images consist of 3D-stacks from 80 to 100 µm thick sections, stained for DAPI (blue), Collagen Type II (COL2, yellow), and Collagen Type X (COLX, red). A clear reduction in the COLX signal could be observed in the MSOD-BΔR1 tissues, indicating impaired hypertrophy (scale bars=500 µm). (G) Isosurface-based quantification of the COL2 immunofluorescent signal using the IMARIS software. No significant difference between groups confirms the retention of cartilage formation in RUNX2-edited constructs. Unpaired t-test, n=3 biological replicates, n.s.=not significant. (H) Isosurface-based quantification of the COLX immunofluorescent signal using the IMARIS software, confirming the disruption of hypertrophy in the MSOD-BΔR1 constructs. Unpaired t-test, n=3, ***p<0.001. (I) Alizarin Red staining evidencing a lack of mineralization in the MSOD-BΔR1 culture compared to the MSOD-B. (J) Quantitative polymerase chain reaction (PCR) analysis displaying the relative expression levels of osteogenesis-related genes: RUNX2, COL1, and ALPL. The expression is normalized to GAPDH as housekeeping gene. n.d.=not detected. All error bars in the figures indicate either the standard deviation (SD) or the standard error of the mean (SEM); the specific metric used is consistent within each figure.
-
Figure 3—source data 1
Figure 3C annotated western blot analysis of Runt-related transcription factor 2 (RUNX2) editing.
Western blot analysis of RUNX2 in cultured mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) (MB) and RUNX2-edited cells (clones 6.1 and 6.23, respectively). The genetic editing of RUNX2 is confirmed by the detection of the truncated proteins. Actin is used as a control to normalize the protein content. Exposure was set at 13.7 s.
- https://cdn.elifesciences.org/articles/96941/elife-96941-fig3-data1-v1.pdf
-
Figure 3—source data 2
Figure 3C annotated western blot analysis of Runt-related transcription factor 2 (RUNX2) editing.
Non-annotated western blot pictures of RUNX2 (left part of the gel) in cultured mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) (line 3) and RUNX2-edited cells (clone 6.1 line 4 and clone 6.23 line 5, respectively). The genetic editing of RUNX2 is confirmed by the detection of the truncated proteins. Actin (right part of the gel) is used as a control to normalize the protein content.
- https://cdn.elifesciences.org/articles/96941/elife-96941-fig3-data2-v1.zip
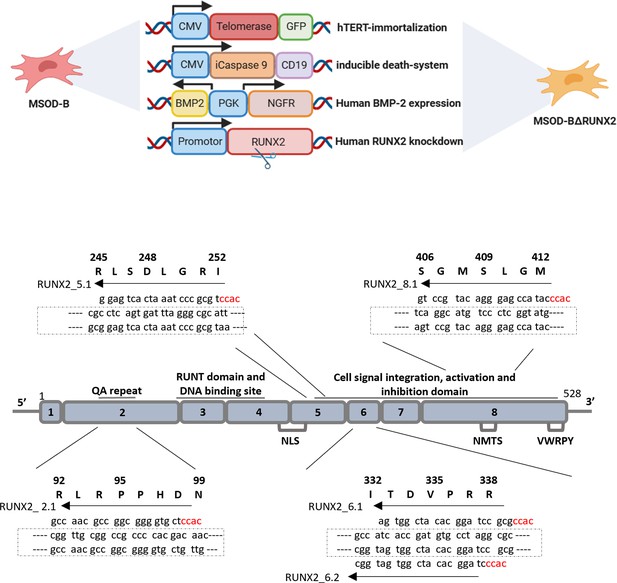
Implemented genetic elements of the mesenchymal sword of Damocles Runt-related transcription factor 2 (RUNX2) (MSOD-BΔR) cells consisting of the human telomerase reverse transcriptase (hTERT), an inducible caspase 9 (iCaspase) death system, the bone morphogenetic protein type-2 and RUNX2 knockdown and an overview of human RUNX2 exon structure, nucleotide sequences of targeting guide RNAs (gRNAs) and its corresponding amino acid sequence. Partly created with BioRender.com.
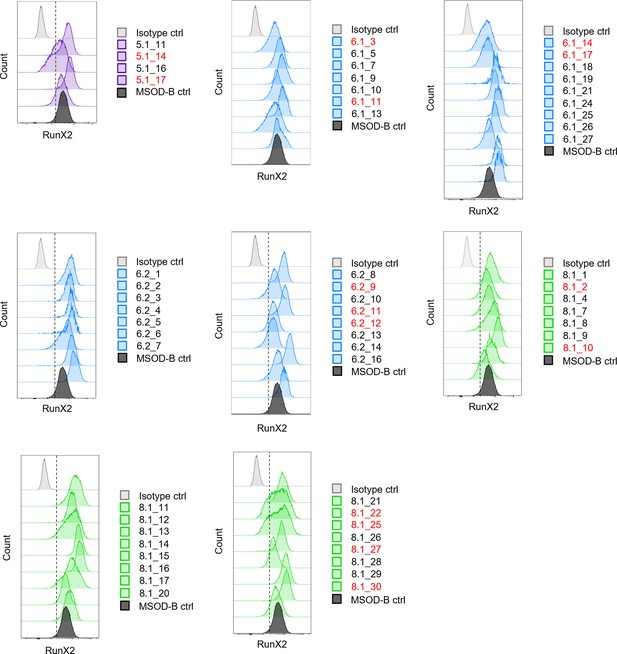
Intracellular flow cytometry for Runt-related transcription factor 2 (RUNX2) detection in mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and RUNX2-edited clones.
Clones whose peaks fell left of the dotted line were selected for sequencing.
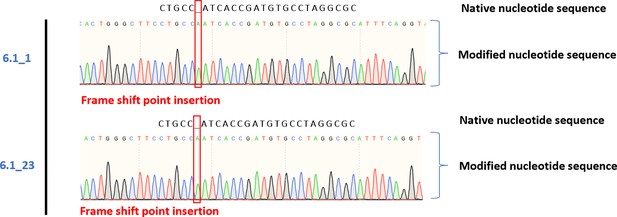
Sanger sequencing results of DNA extracted from mesenchymal sword of Damocles (MSOD)-BΔR samples and the modifications compared to MSOD-B DNA sequence.
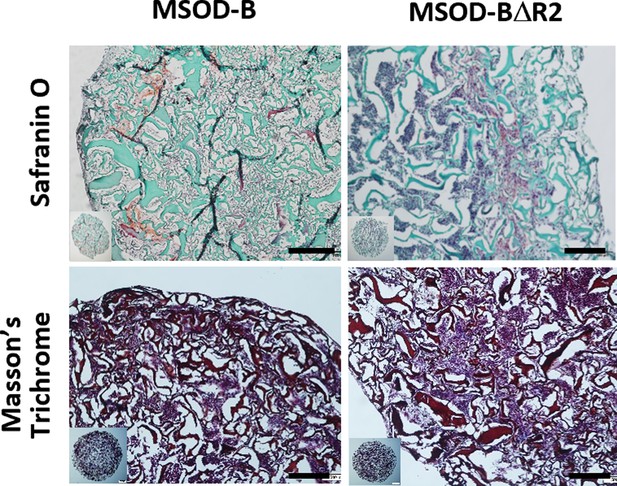
Histological analysis of in vitro constructs from using Safranin O and Masson’s trichrome staining displayed the presence of glycosaminoglycans (GAG) and collagen content, respectively, indicating the presence of cartilage formation in mesenchymal sword of Damocles (MSOD)-BΔR2 (scale bars=200 µm).
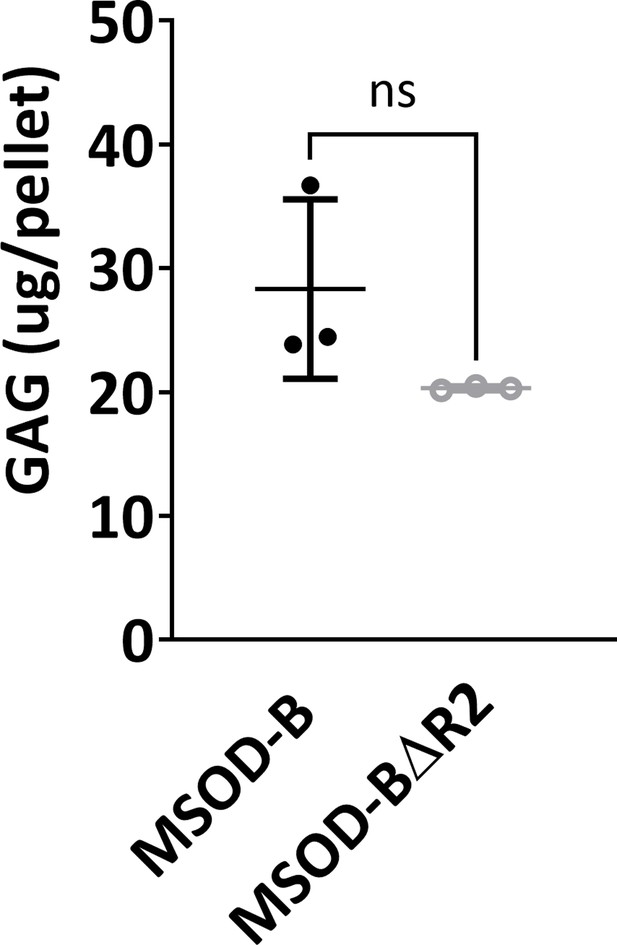
Quantitative assessment of the total GAG content confirms the presence of cartilage formation in both constructs thus confirming Runt-related transcription factor 2 (RUNX2) knockout in human mesenchymal stromal/stem cells (hMSCs) (mesenchymal sword of Damocles [MSOD]-BΔR2) retain cartilage formation (unpaired t-test, n=3, n.s.=not significant).
The error bars in the figures indicate the standard deviation (SD).
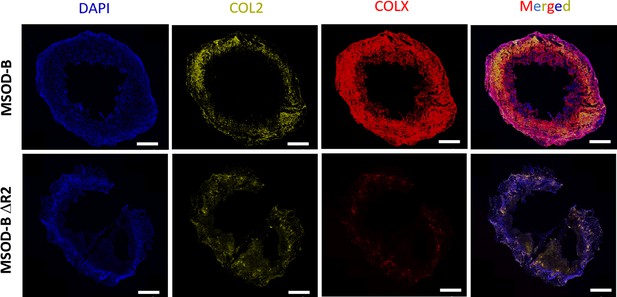
Reduction of Runt-related transcription factor 2 (RUNX2) expression in cartilage tissues lead to reduced hypertrophy.
Representative immunofluorescence staining images of COL2, COLX, and DAPI tissue section of in vitro cartilage constructs show reduction of COLX expression in knockout construct (mesenchymal sword of Damocles [MSOD]-BΔR2), indicating reduction of hypertrophy (scale bars=500 µm).
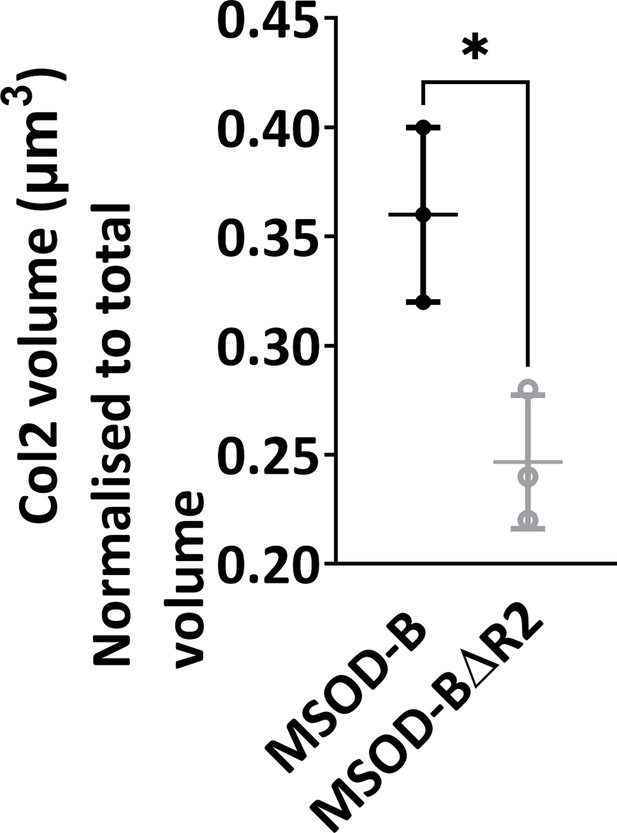
Quantitative analysis of the immunofluorescent staining sections using IMARIS software displayed significant difference in the expression of COL2, demonstrating reduction of cartilage formation in knockout constructs (mesenchymal sword of Damocles [MSOD]-BΔR2) (unpaired t-test, n=3, *p<0.05).
The error bars in the figures indicate the standard deviation (SD).
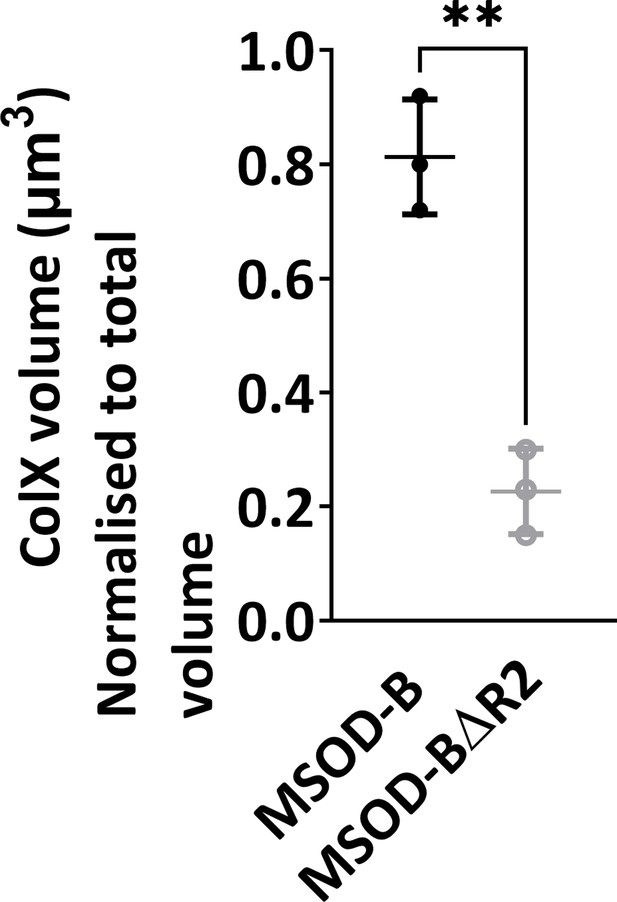
Quantitative analysis of the immunofluorescent staining sections using IMARIS software displayed a significant reduction in expression of COLX, confirming the disruption of hypertrophy in knockout constructs (unpaired t-test, n=3, **p<0.01).
The error bars in the figures indicate the standard deviation (SD).
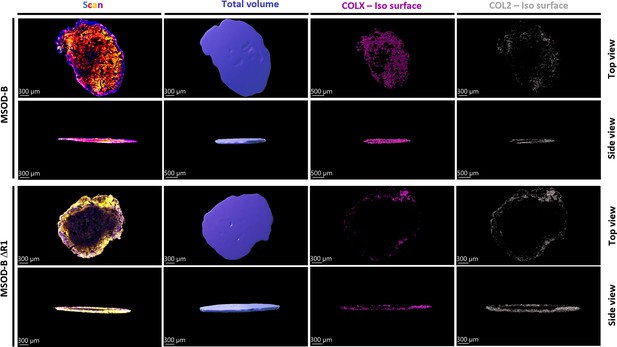
Representative isosurface images of immunofluorescent staining sections constructed using IMARIS software.
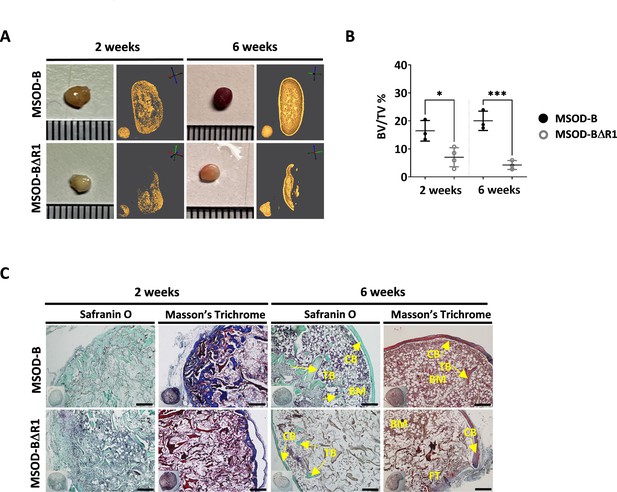
Runt-related transcription factor 2 (RUNX2) knockout in cartilage tissues disrupts effective ectopic bone formation.
(A) Representative macroscopic and microtomography images of in vivo constructs retrieved at 2 and 6 weeks postimplantation. (B) Microtomography-based quantification of the sample’s bone/mineralized volume over their total volume (ratio) (BV: bone volume, TV: total volume). A marked difference is observed as early as 2 weeks, with a clear lower mineral content in mesenchymal sword of Damocles (MSOD)-BΔR1 samples. Ordinary one-way ANOVA, n=3 biological replicates, *p<0.05, ***p<0.001. (C) Histological analysis of in vivo constructs using Safranin O and Masson’s trichrome stains. After 2 weeks (2W), a higher bone formation is already evident in the MSOD-B control group. The MSOD-BΔR1 samples explanted after 6 weeks (6W) displayed presence of cortical and trabecular bone, but also large amount of fibrous tissue indicating an incomplete remodeling (scale bars=200 µm). The error bars in the figures indicate the standard deviation (SD).
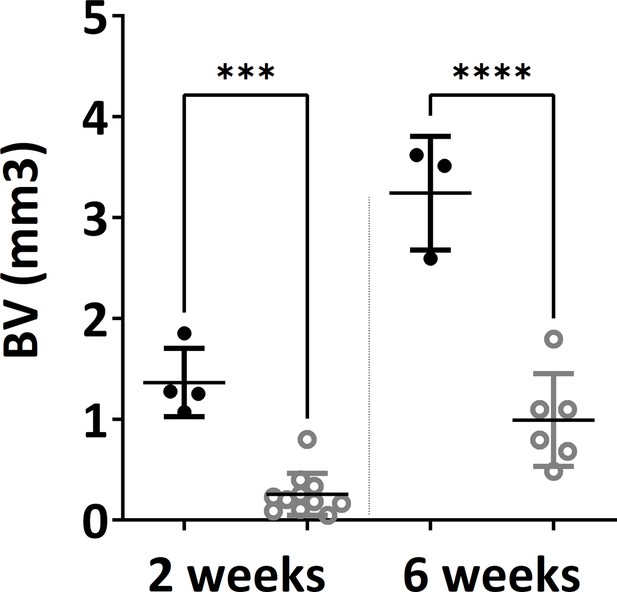
Microtomography analysis (bone/mineralized volume) showed very high significant difference between in vivo constructs at both 2 weeks and 6 weeks time points (one-way ANOVA, n=3, ***p<0.001, ****p<0.0001).
The error bars in the figures indicate the standard deviation (SD).
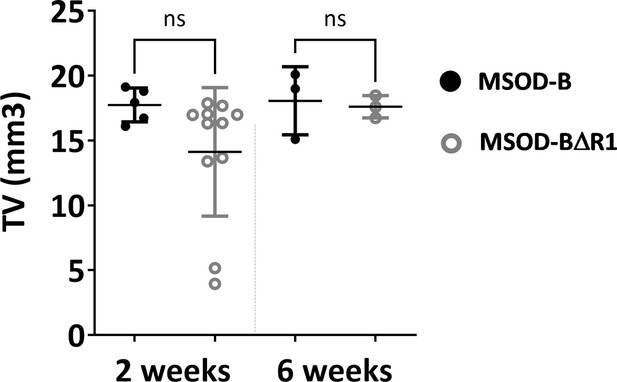
Microtomography analysis (total volume) showed significant difference between in vivo constructs at 2 weeks time point but no significant difference at 6 weeks time point (one-way ANOVA, n=3, n.s.=not significant) (BV: bone volume, TV: total volume).
The error bars in the figures indicate the standard deviation (SD).
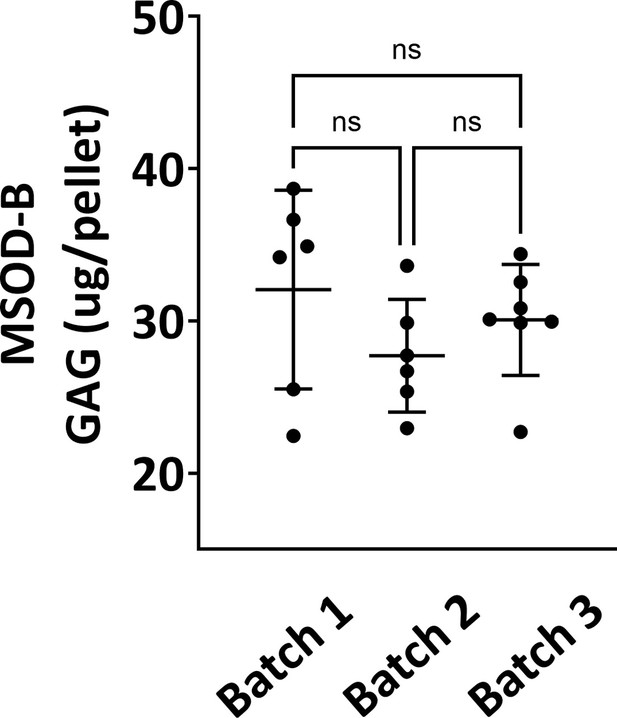
Quantitative assessment of the total GAG content in mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) in vitro differentiated constructs across three different batch productions.
One-way ANOVA test, n=6–7 biological replicates, n.s.=not significant. The error bars in the figures indicate the standard deviation (SD).
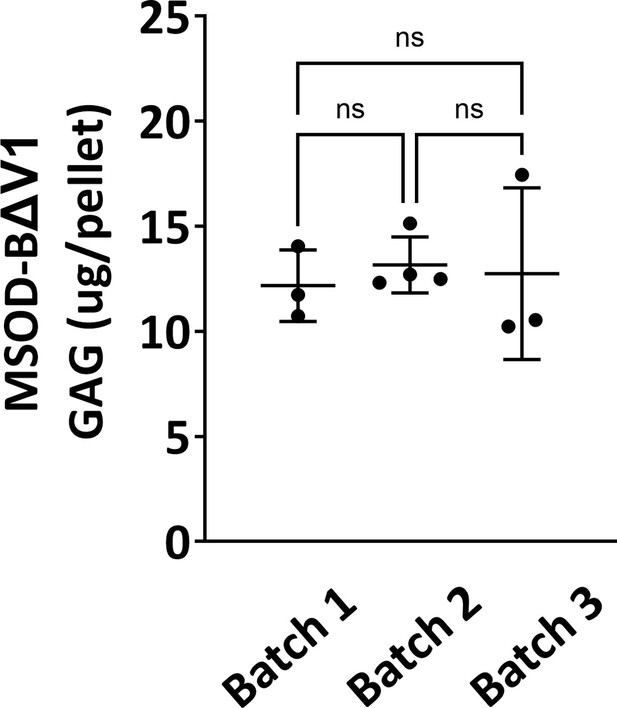
Quantitative assessment of the total GAG content in mesenchymal sword of Damocles (MSOD)-BΔV1 in vitro differentiated constructs across three different batch productions.
One-way ANOVA test, n=3–4 biological replicates, n.s.=not significant. The error bars in the figures indicate the standard deviation (SD).
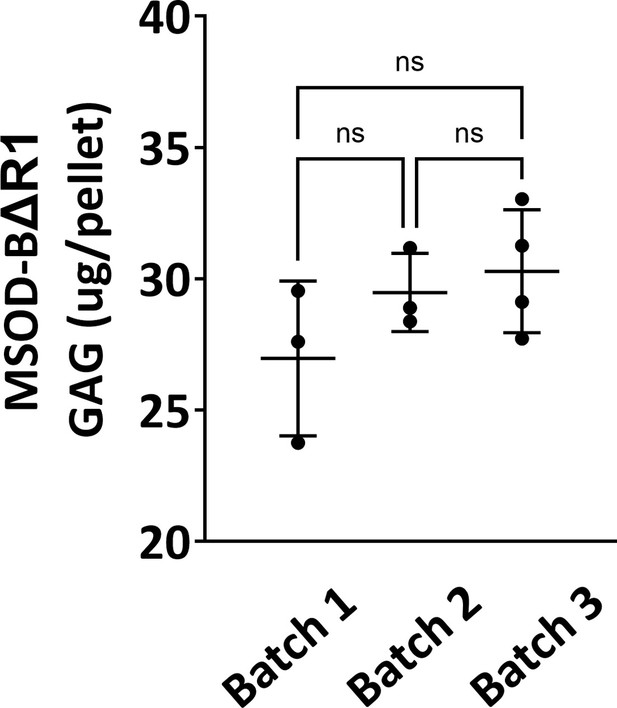
Quantitative assessment of the total GAG content in mesenchymal sword of Damocles (MSOD)-BΔR1 in vitro differentiated constructs across three different batch productions.
One-way ANOVA test, n=3 biological replicates, ***p<0.001, ****p<0.0001, n.s.=not significant. The error bars in the figures indicate the standard deviation (SD).
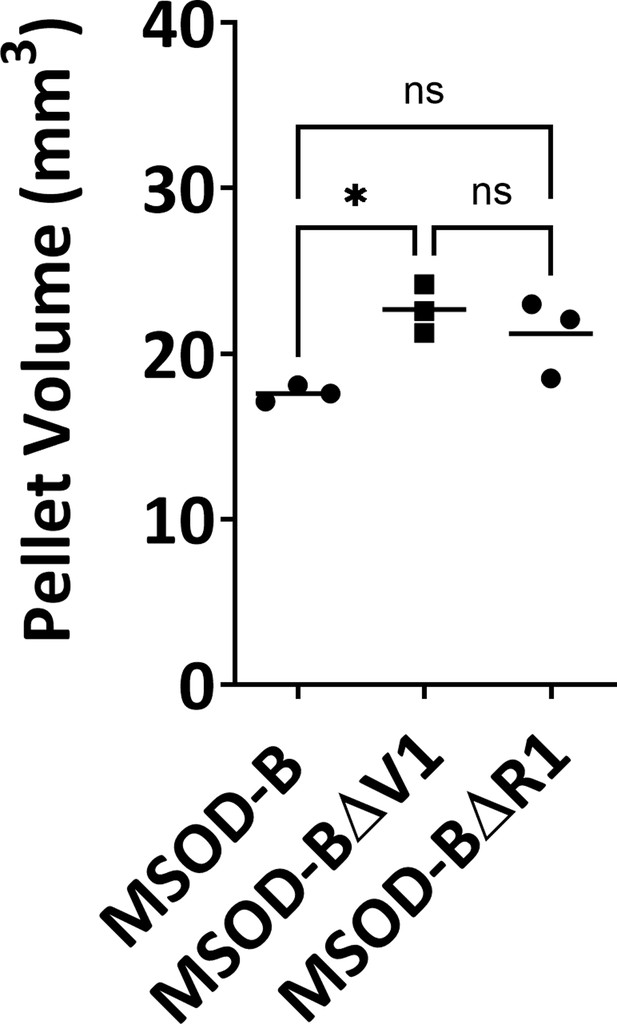
Quantitative assessment of cartilage tissue volume in mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B), MSOD-BΔV1, and MSOD-BΔR1 tissues.
One-way ANOVA test, n=3 biological replicates, *p<0.05, n.s.=not significant. The error bars in the figures indicate the standard deviation (SD).
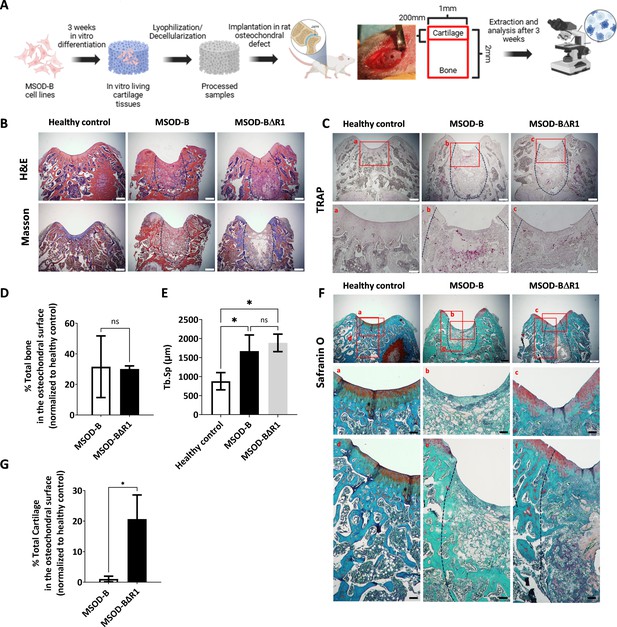
Runt-related transcription factor 2 (RUNX2) knockout in cartilage tissues leads to better cartilage regeneration and maintenance in osteochondral defect in rats.
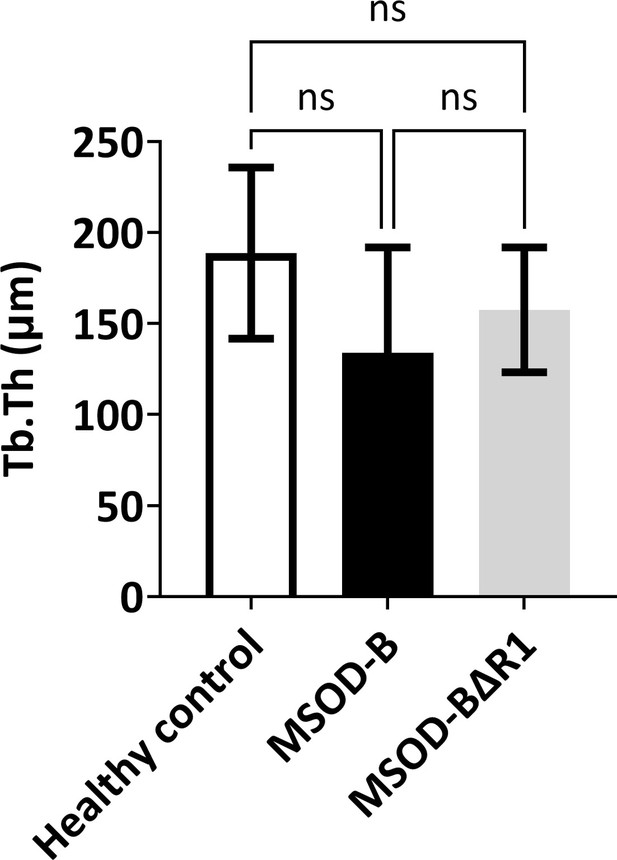
(A) Experimental scheme for the regenerative potential assessment of mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and MSOD-BΔR1 cartilage tissues in a rat osteochondral defect. (B) Histological analysis of the osteochondral defects for each group using hematoxylin and eosin (H&E) and Masson’s trichrome stains, at 3 weeks postimplantation. The dash-line marks the defect area (scale bars=500 µm). (C) Histological analysis of the osteochondral defects using tartrate-resistant acid phosphatase (TRAP) staining reporting osteoclastic activity, at 3 weeks postimplantation. The dash-line marks the defect area (scale bars=500 µm and 100 µm for magnified areas). (D) Microtomography-based quantification of the sample’s total bone/mineralized volume normalized to the healthy control in percentage. Unpaired t-test, n=3 biological replicates, n.s.=not significant. (E) ImageJ-based quantification of trabecular separation (Tb.Sp). One-way ANOVA test, n=3 biological replicates, *p<0.05. (F) Histological analysis of the osteochondral defects using Safranin O staining. After 3 weeks, a higher regeneration of the surface cartilage is evident in the MSOD-BΔR1 group (a,b,c) (scale bars=500 µm and 100 µm for magnified areas). The magnified regions of the subchondral area (d,e,f) show higher cartilage remnants and integration in the MSOD-BΔR1 group. (G) Quantitative analysis of cartilage regeneration in the osteochondral surface as compared to the healthy control (100%). Unpaired t-test, n=3 biological replicates, *p<0.05. All error bars in the figures indicate the standard deviation (SD). Panel A was created using BioRender.com.
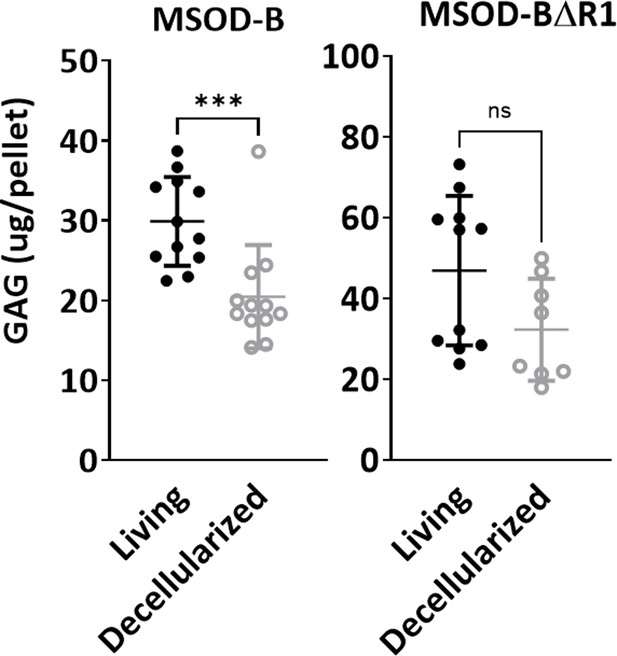
Quantitative assessment of the total GAG content in mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and MSOD-BΔR1 in vitro differentiated constructs before (living) and after (decellularized) the decellularization process (unpaired t-test, n=12 biological replicates, ***p<0.001, n.s.=not significant).
The error bars in the figures indicate the standard deviation (SD).
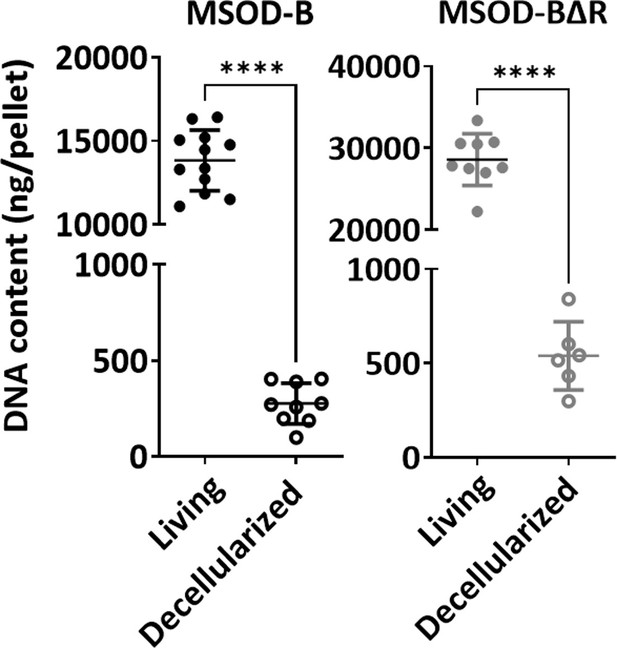
DNA quantification of in vitro engineered cartilage grafts before (living) and after (decellularized) the decellularization process (unpaired t-test, n=12 biological replicates, ****p<0.0001).
The error bars in the figures indicate the standard deviation (SD).
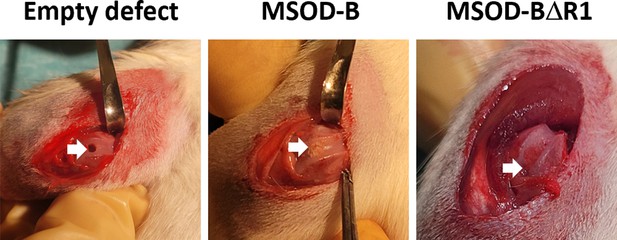
Macroscopic images of osteochondral defect model in rats depicting in of empty defect, defect filled with mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B), and defect filled with MSOD-BΔR1, respectively.
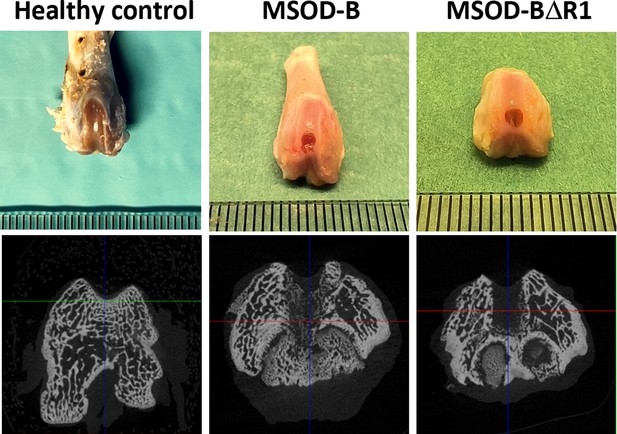
Macroscopic images of extracted rat knees depicting healthy control without any defect, defect filled with mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B), and defect filled with MSOD-BΔR1 respectively and microtomography images of same respective constructs.
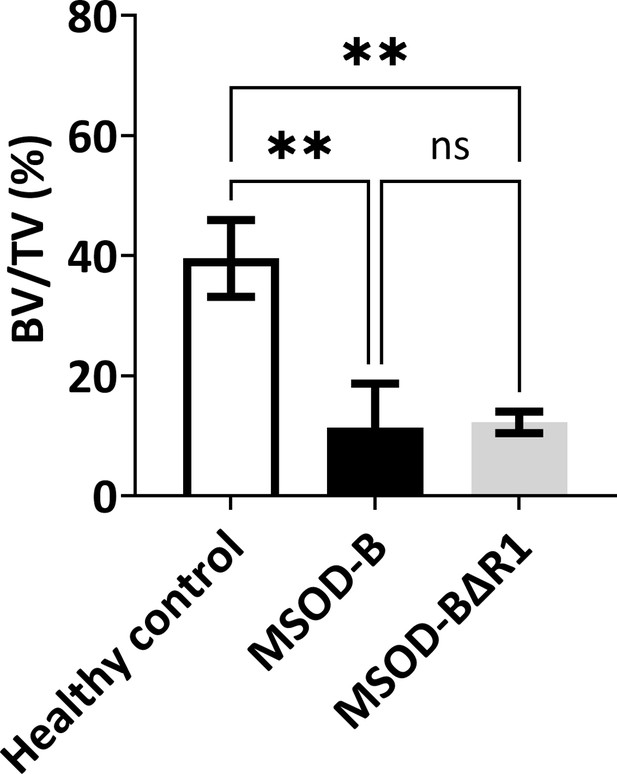
No significant difference in bone volume/total volume (BV/TV) is observed between mesenchymal sword of Damocles bone morphogenetic type-2 (MSOD-B) and MSOD-BΔR1 samples despite significantly reduced BV/TV ratio in both samples compared to healthy control.
One-way ANOVA test, n=3 biological replicates, **p<0.01. The error bars in the figures indicate the standard deviation (SD).
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | MSOD-B | University/Hospital of Basel | Pigeot S, Klein T, Gullotta F, et al. doi:10.1002/adma.202103737 | Parental human mesenchymal stromal/stem cell line used for eECM production |
| Cell line (Homo sapiens) | MSOD-BΔV1 | MSOD-B (this paper) | MSOD-BΔV1 | CRISPR/Cas9‐edited derivative with VEGF knockout. This cell line was created using our MSOD-B line as base. |
| Cell line (Homo sapiens) | MSOD-BΔR1 | MSOD-B (this paper) | MSOD-BΔR1 | CRISPR/Cas9‐edited derivative with RUNX2 knockout; exhibits impaired hypertrophy. This cell line was created using our MSOD-B line as base. |
| Transfected construct (Homo sapiens) | pU6-(BbsI)_CBh-Cas9-T2A-mCherry | Addgene | Plasmid #64324 | Vector for CRISPR/Cas9 editing; encodes SpCas9 and mCherry reporter |
| Transfected construct (Homo sapiens) | VEGF gRNAs (VEGF_1.1, 1.2, 1.3, 2.1, 8.1) | This paper | pU6-(BbsI)_CBh-Cas9-T2A-mCherry-VEGF_1.1, 1.2, 1.3, 2.1, 8.1 | Same plasmid but with guide RNAs targeting all isoforms of VEGF |
| Transfected construct (Homo sapiens) | RUNX2 gRNAs (RUNX2_2.1, 5.1, 6.1, 6.2, 8.1) | This paper | pU6-(BbsI)_CBh-Cas9-T2A-mCherry-RUNX2_2.1, 5.1, 6.1, 6.2, 8.1 | Same plasmid but with guide RNAs targeting conserved regions of RUNX2 |
| Antibody | Anti-RUNX2 (Rabbit polyclonal) | Thermo Fisher Scientific | Cat# PA5-82787 | Rabbit polyclonal; used for intracellular flow cytometry (1:100) |
| Antibody | Anti-RUNX2 (D1L7F) (Rabbit monoclonal) | Cell Signaling Technology | Cat# 12556 | Used for western blot analysis (1:1000) |
| Antibody | Anti-Actin (Mouse monoclonal) | BD Biosciences | Cat# 612656 | Loading control for western blotting (1:200) |
| Antibody | Anti-Collagen II (Mouse monoclonal) | Invitrogen | Cat# MA137493 | Detects cartilage matrix in immunofluorescence (1:200) |
| Antibody | Anti-Collagen I (Rabbit monoclonal) | Abcam | Cat# ab138492 | Used for extracellular matrix immunostaining (1:200) |
| Antibody | Anti-Collagen X (Rabbit polyclonal) | abbexa | Cat# abx101469 | Marker of hypertrophic cartilage (assesses tissue maturation) (1:200) |
| Antibody | Anti-CD31 (Mouse monoclonal) | R&D Systems | Cat# 11-0319-42 | Vascular endothelial marker; used in immunofluorescence for vessel detection (1:200) |
| Antibody | Anti-VEGF (Rabbit polyclonal) | Bioss Antibodies | Cat# bs-0279R | Used to detect VEGF deposition within engineered ECM (1:200) |
| Sequence-based reagent | RUNX2 | Thermo Fisher Scientific | Hs00298328_s1 | qPCR primer |
| Sequence-based reagent | COLX | Thermo Fisher Scientific | Hs00166657_m1 | qPCR primer |
| Sequence-based reagent | ALPL | Thermo Fisher Scientific | Hs01029144_m1 | qPCR primer |
| Sequence-based reagent | Forward primer | IDT | PCR primer | AACGCTTTGTGCTATTTAAGGC |
| Sequence-based reagent | Reverse primer | IDT | PCR primer | AAGAAAGGAACACAAGCAGAGG |
| Sequence-based reagent | Forward primer | IDT | Sequencing primer | TCCCTGTTTTTCTGCTTTTTCC |
| Sequence-based reagent | Reverse primer | IDT | Sequencing primer | TAACTGGGCGGCATTAAATACC |
| Sequence-based reagent | VEGF_1.1F | IDT | gRNA | caccGCGAGCGCCGAGT CGCCACTG |
| Sequence-based reagent | VEGF_1.2 | IDT | gRNA | caccGGAGGAAGAGTAG CTCGCCG |
| Sequence-based reagent | VEGF_1.3 | IDT | gRNA | caccGCCAAGACAGCAG AAAGTTCA |
| Sequence-based reagent | VEGF_2.1 | IDT | gRNA | caccGCTGCACCCATGG CAGAAGG |
| Sequence-based reagent | VEGF_8.1 | IDT | gRNA | caccGTCCTGCCCGGCT CACCGCCT |
| Sequence-based reagent | RUNX2_2.1 | IDT | gRNA | caccTCGTGGGGCGGCC GCAACCG |
| Sequence-based reagent | RUNX2_5.1 | IDT | gRNA | caccTGCGCCCTAAATC ACTGAGG |
| Sequence-based reagent | RUNX2_6.1 | IDT | gRNA | caccGCGCCTAGGCACATC GGTGA |
| Sequence-based reagent | RUNX2_6.2 | IDT | gRNA | caccCTAGGCACATCGGT GATGGC |
| Sequence-based reagent | RUNX2_8.1 | IDT | gRNA | caccCATACCGAGGGAC ATGCCTG |
Additional files
-
Supplementary file 1
Scoring rate for individual categories.
Semiquantitative analysis utilizing a modified histological grading system, including parameters such as cellular morphology, matrix staining, surface regularity, thickness of cartilage, subchondral bone formation, and integration of adjacent cartilage, resulting in a comprehensive assessment of tissue regeneration and integration
- https://cdn.elifesciences.org/articles/96941/elife-96941-supp1-v1.docx
-
Supplementary file 2
Histological grading system.
The histological graded system is modified and adapted from previously established model as mentioned in Materials and methods section. This modified scale incorporates six distinct parameters to comprehensively evaluate the quality of tissue repair.
- https://cdn.elifesciences.org/articles/96941/elife-96941-supp2-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/96941/elife-96941-mdarchecklist1-v1.pdf
-
Source data 1
Compiled source data for all figures.
- https://cdn.elifesciences.org/articles/96941/elife-96941-data1-v1.zip