The rhizobial effector NopT targets Nod factor receptors to regulate symbiosis in Lotus japonicus
Figures
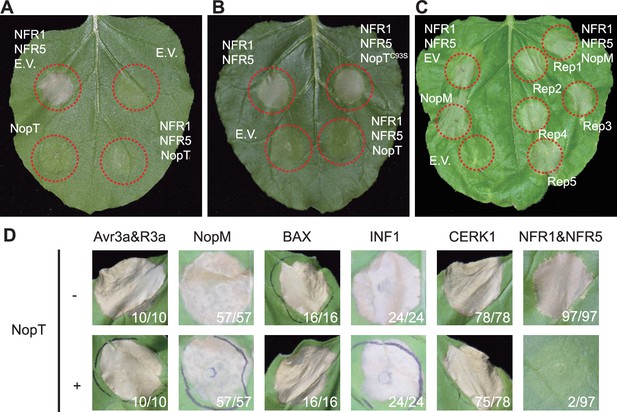
NopT specifically suppresses NFR1/NFR5-triggered cell death in N. benthamiana.
Agrobacterium strains harboring plasmid DNA encoding NopT or the empty vector (EV) were infiltrated into N. benthamiana leaves. At 12 hpi, a second infiltration was performed with an Agrobacterium strain containing a plasmid with NFR1/NFR5 genes. Dashed red lines indicate leaf discs where different proteins were expressed. NopT (A), but not NopTC93S (B), a protease-inactive version of NopT, suppressed NFR1–NFR5-induced cell death. (C) Expression of NopM-induced cell death in N. benthamiana. (D) Expression of NopT could not suppress the cell death response triggered by expression of Avr3a/R3a, NopM, BAX, INF1, or AtCERK1. Numbers in (D) represent the number of leaf discs showing cell death and total leaf discs tested. Cell death in leaf disc results in the formation of necrotic plaques, which restrains pathogens within deceased cells. These plaques commonly manifest as leaf dehydration, frequently accompanied by a translucent appearance. Brown and shriveled leaf discs serve as indicators of cell death. The pictures shown in this figure are representative of at least three independent biological replicates.
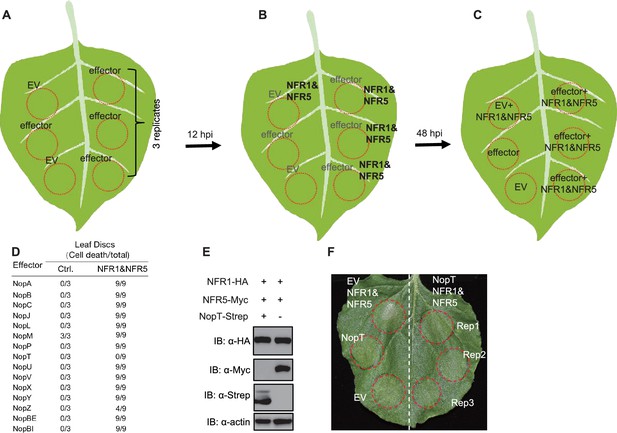
Analyses of the functions of all 15 effectors from S. fredii NGR234 in preventing NFR1- and NFR5-induced cell death in N. benthamiana leaves.
15 genes from S. fredii NGR234 encoding effector proteins were amplified and cloned into binary vector for expression of Strep-tagged proteins in N. benthamiana. NFR1 and NFR5 from L. japonicus were C-terminally tagged with HA and Myc, respectively, and cloned into binary vectors for expression in N. benthamiana. All plasmids were electroporated into Agrobacterium tumefaciens EHA105. (A) Agrobacterium strain harboring each effector gene or control empty vector (EV) were infiltrated into N. benthamiana. Twelve-hour post infiltration (hpi), Agrobacterium strains harboring NFR1 and NFR5 were mixed and hand-infiltrated in the same leaf discs as shown in (B). 48 hpi, cell death phenotypes were evaluated. (C) At least three replicates in the same leaf were performed to test the function of each effector in suppressing cell death triggered by NFR1 and NFR5. (D) Results of all 15 effector proteins in suppressing programmed cell death by over-expression of NFR1 and NFR5. (E) Abundance of NFR1, NFR5, and NopT proteins, as measured by immunoblot with specific antibodies. Leaf discs expressing NFR1, NFR5, and NopT or NFR1 and NFR5 and with EV were used for immunoblot analyses. Actin was used as loading control. (F) Another example of suppression of programmed cell death by NopT in the leaf discs co-expressing with NFR1 and NFR5. Rep1, Rep2, and Rep3 represent three technical repeats in the same leave.
-
Figure 1—figure supplement 1—source data 1
Original files for western blot analysis displayed in Figure 1—figure supplement 1E.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig1-figsupp1-data1-v1.zip
-
Figure 1—figure supplement 1—source data 2
PDF file containing original western blots for Figure 1—figure supplement 1E, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig1-figsupp1-data2-v1.pdf
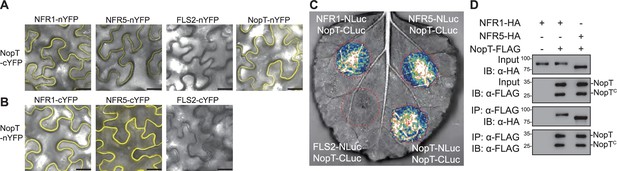
NopT interacts with NFR1 and NFR5.
Interactions between NopT and NFR1 or NFR5 were detected using bimolecular fluorescence complementation (BiFC) (Split-YFP) (A, B), Split-LUC complementation (C) and co-IP (D) assays in N. benthamiana leaves. (A, B) For BIFC analysis, nYFP and cYFP tags were fused at the C-terminus of NopT, NFR1, NFR5, and the flagellin receptor FLS2 (negative control). YFP fluorescence signals represent protein–protein interactions. Scale bar = 25 μm. (C) For the Split-LUC complementation assay, the NLuc and CLuc tags were fused at the C-terminus of NopT, NFR1, NFR5, and FLS2 (negative control). Luminescence signals represent protein–protein interactions. (D) For the co-IP assay, HA-tagged NFR1 or NFR5 and FLAG-tagged NopT were expressed in N. benthamiana cells followed by immunoprecipitation using an anti-FLAG antibody. Immunoblot analysis was performed using anti-HA and anti-FLAG antibodies (NopTC denotes the truncated version of NopT after autocleavage). The images and immunoblots shown in this figure are representative of three biological replicates.
-
Figure 2—source data 1
Original files for western blot analysis displayed in Figure 2D.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-data1-v1.zip
-
Figure 2—source data 2
PDF file containing original western blots for Figure 2D, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-data2-v1.pdf

NopT interacts with both NFR1 and NFR5.
The full blot of co-IP assay as shown in Figure 2D. The smaller bands are marked with asterisks. Compared to NFR1, NFR5 might be more susceptible to degradation during protein extraction from N. benthamiana leaves.
-
Figure 2—figure supplement 1—source data 1
Original files for western blot analysis displayed in Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-figsupp1-data1-v1.zip
-
Figure 2—figure supplement 1—source data 2
PDF file containing original western blots for Figure 2—figure supplement 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-figsupp1-data2-v1.pdf
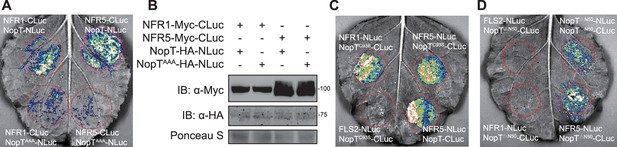
Split-LUC assay testing the interactions between different NopT mutants and NFRs.
The interaction between NopT with mutated acylation site and NFR1/NFR5. NopTG50A/C51A/C52A represents a modified NopT form lacking acylation sites. (B) The protein expressing of indicating genes in the leaves of (A). (C) NopC93S interacts with NFR1 and NFR5. (D) The interaction between NopTΔN50 and NFR5. NopTΔN50 represents a truncated NopT with deletion of 50 amino acid residues from its N-terminus.
-
Figure 2—figure supplement 2—source data 1
Original files for western blot analysis displayed in Figure 2—figure supplement 2B.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-figsupp2-data1-v1.zip
-
Figure 2—figure supplement 2—source data 2
PDF file containing original western blots for Figure 2—figure supplement 2B, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig2-figsupp2-data2-v1.pdf
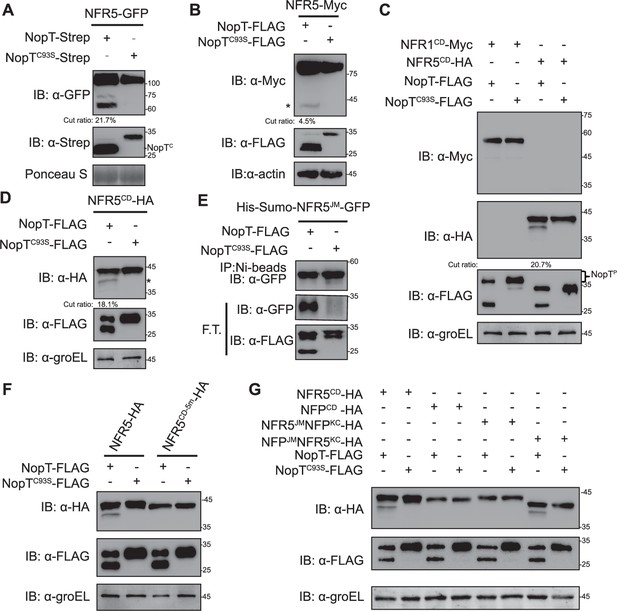
NopT proteolyzes NFR5 at its juxtamembrane (JM) domain.
Proteins with indicated tags were expressed in N. benthamiana (A), L. japonicus (B), or E. coli cells (C–G) and detected by immunoblotting. (A) NopT but not NopTC93S (a protease-dead version of NopT) cleaves NFR5-GFP protein expressed in N. benthamiana cells by releasing NFR5CD-GFP (NopTC denotes autocleaved NopT). The cleavage efficiency was marked under the lane. (B) NopT but not NopTC93S cleaves NFR5-Myc expressed in hairy roots of L. japonicus by releasing NFR5CD-Myc. The cleavage efficiency was marked under the lane. The asterisk indicates the HA-tagged NFR5 cleavage product containing the kinase domain and a C-terminal tail region. (C) Analysis of the CDs of NFR1 and NFR5 co-expressed with NopT or NopTC93S in E. coli cells. Cleavage of NFR5CD was observed for NopT but not NopTC93S, while NFR1CD was not proteolyzed by NopT. In the presence of NFR1CD, a slower migrating band was observed, possibly representing phosphorylated NopT (NopTP). The cleavage efficiency was marked under the lane. (D) A repeat experiment confirmed that NopT is able to cleave NFR5CD (the asterisk indicates the HA-tagged NFR5 cleavage product containing the kinase domain and a C-terminal tail region). The cleavage efficiency was marked under the lane. (E) NopT cleaves His-SUMO-NFR5JM-GFP (His-SUMO and GFP linked by the juxtamembrane [JM] domain of NFR5) in vitro. F.T. indicates proteins in flow through samples after purification with Ni-beads. (F) NopT expressed in E. coli cells was unable to cleave co-expressed NFR5CD-5m-HA, a modified version of NFR5CD-HA in which five amino acids of the JM were substituted by other residues (S283Y, G294Q, Y303S, A310I, and T311Y). (G) NopT expressed in E. coli cells was unable to cleave co-expressed M. truncatula NFPCD-HA and the NFR5JM-NFPKC-HA fusion protein, while NFPJM-NFR5KC was proteolyzed (KC stands for the kinase domain and a C-terminal tail region, modified Nod factor receptors without the JM).
-
Figure 3—source data 1
Original files for western blot analysis displayed in Figure 3A–G.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-data1-v1.zip
-
Figure 3—source data 2
PDF file containing original western blots for Figure 3A–G, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-data2-v1.zip
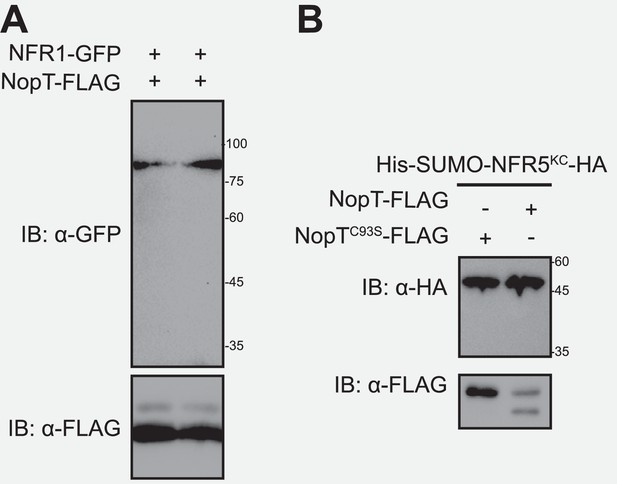
NopT cleaves NFR5 but not NFR1.
Proteins with indicated tags were expressed in either N. benthamiana or E. coli cells and detected by immunoblotting. (A) NopT did not cleave NFR1-GFP expressed in N. benthamiana leaves. (B) NopT expressed in E. coli cells did not cleave co-expressed SUMO-NFR5KC-HA. KC: kinase domain and C-terminal domain, a modified NFR5CD without the juxtamembrane (JM).
-
Figure 3—figure supplement 1—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 1A and B.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp1-data1-v1.zip
-
Figure 3—figure supplement 1—source data 2
PDF file containing original western blots for Figure 3—figure supplement 1A and B, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp1-data2-v1.zip

NopT interacts with the juxtamembrane (JM) domain of NFR5.
NopT interacts with NFR5JM in in vitro pull-down assay. IB: immunoblotting, IP: immunoprecipitation. His-SUMO-NFR5JM-GFP: a recombinant protein containing His tag, SUMO tag, the JM domain of NFR5, and GFP.
-
Figure 3—figure supplement 2—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 2.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp2-data1-v1.zip
-
Figure 3—figure supplement 2—source data 2
PDF file containing original western blots for Figure 3—figure supplement 2, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp2-data2-v1.pdf
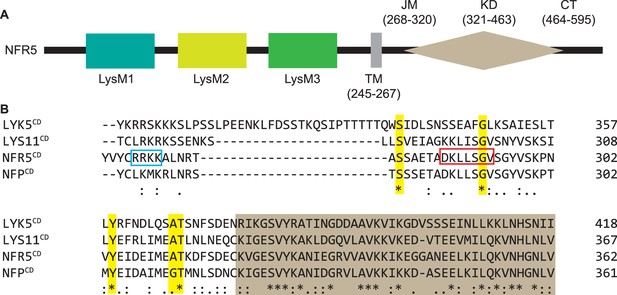
Conserved domains and residues of NFR5 and related proteins.
(A) Domain and motif annotation of NFR5. LysM: Lysin-Motif; TM: transmembrane domain; JM: juxtamembrane domain; KD: kinase domain; CT: C-terminal domain. (B) Alignment of amino acid residues of AtLYK5, LjLYS11, NFR5, and MtNFP close to the RRKK motif in the JM domain of NFR5. Five highly conserved residues marked by yellow coloration (S283, G294, Y304, A310, T311 in NFR5) and the brown box indicates the start region of kinase domain. The blue frame delineates the RRKK motif in NFR5 and the red-framed residues are similar to the NopT autocleavage region.
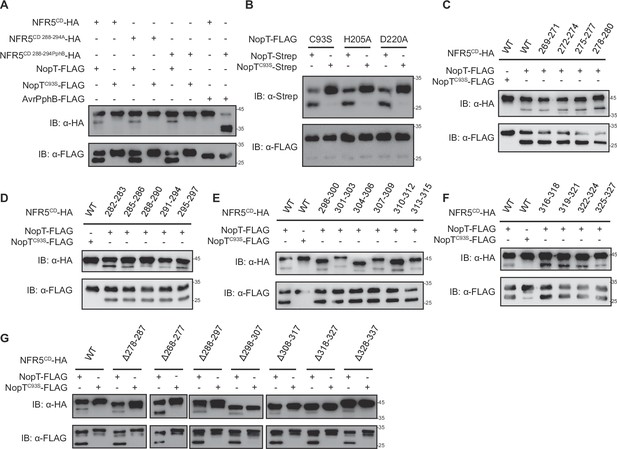
NopT cleaves NFR5 at the juxtamembrane domain.
(A) Cleavage assay using two mutant versions of NFR5CD in the presence of NopT or AvrPphB. NFR5CD288-294A represents a mutant NFR5CD with residues 288–294 replaced with seven alanines; NFR5CD288-294PphB represents a mutant NFR5CD with residues 288–294 replaced with AvrPphB recognition sites. (B) Autocleavage assay, using different NopT variants with single amino acid mutations. H205A and D220A indicate His-205 and Asp-220 replaced with alanine, respectively. (C–F) NopT cleavage assay using 19 variants of NFR5CD with three adjacent amino acids replaced with three alanines. The numbers indicate the location of the three residues replaced with alanines. (G) NopT cleavage assay using seven truncated variants of NFR5CD with 10 adjacent amino acids deleted. The numbers indicate the location of the 10 deleted residues.
-
Figure 3—figure supplement 4—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 4A–G.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp4-data1-v1.zip
-
Figure 3—figure supplement 4—source data 2
PDF file containing original western blots for Figure 3—figure supplement 4A–G, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp4-data2-v1.zip
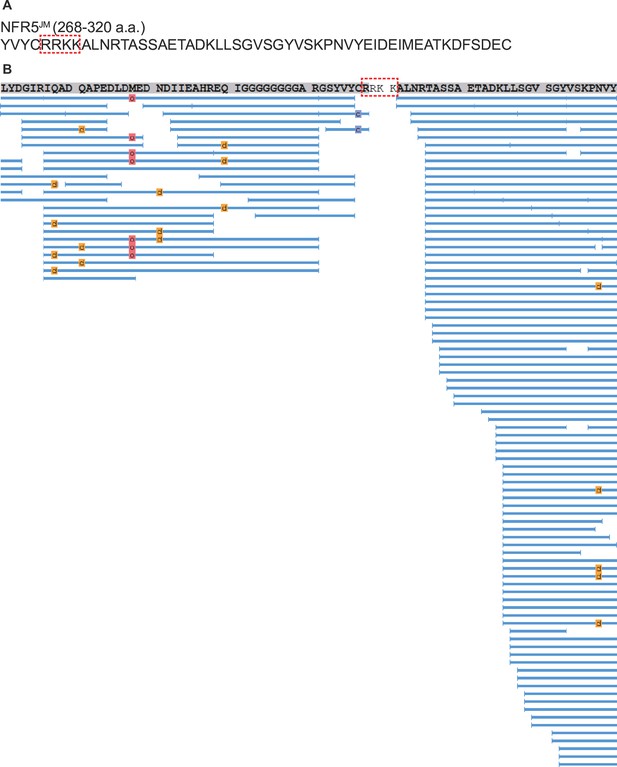

Mass spectrometry analysis of cleavage site of NFR5 by NopT.
(A) The sequence of NFR5 juxtamembrane domain (286–320 a.a.). (B) Mass spectrometry analysis of peptides of NFR5CD after proteolysis by NopT. Blue lines indicated the peptides characterized by mass spectrometry. Red frame delineated cleavage site of NFR5 identified using mass spectrometry.
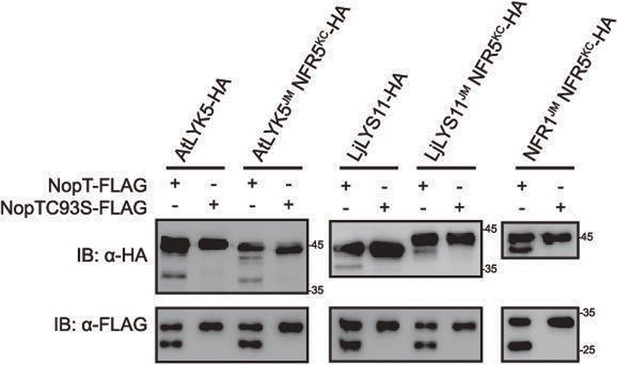
NopT cleaves the NFR5 homolog proteins at the juxtamembrane (JM) domain.
NopT but not NopTC93S could cleave the CDs of AtLYK5 and LjLYS11 expressed in E. coli cells. NopT could cleave both AtLYK5JM-NFR5KC, LjLYS11JM-NFR5KC, and NFR1JM-NFR5KC, three recombinant proteins expressed in E. coli cells. At LYK5JM-NFR5KC, LjLYS11JM-NFR5KC, and NFR1JM-NFR5KC recombinant NFR5 proteins with the JM were replaced with the JMs from AtLYK5, LjLYS11, and NFR1, respectively. KC: kinase domain and C-terminal domain as illustrated in Figure S4B.
-
Figure 3—figure supplement 6—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 6.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp6-data1-v1.zip
-
Figure 3—figure supplement 6—source data 2
PDF file containing original western blots for Figure 3—figure supplement 6, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp6-data2-v1.pdf

NopT cleaves the juxtamembrane (JM) of MtNFP.
NopT cleaves a recombinant protein where His-tagged SUMO and GFP is bridged with the JM domain of NFP. F.T. represents flow through sample after Ni-beads purification. IB: immunoblotting.
-
Figure 3—figure supplement 7—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 7.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp7-data1-v1.zip
-
Figure 3—figure supplement 7—source data 2
PDF file containing original western blots for Figure 3—figure supplement 7, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp7-data2-v1.pdf

NopT cleaves recombinant proteins NFR5268-445-NFP458-595 and NFP270-457-NFR5456-595.
NopT but not NopTC93S could cleave recombinant proteins NFR5268-445-NFP458-595 and NFP270-457-NFR5456-595 detected by immunoblotting (IB).
-
Figure 3—figure supplement 8—source data 1
Original files for western blot analysis displayed in Figure 3—figure supplement 8.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp8-data1-v1.zip
-
Figure 3—figure supplement 8—source data 2
PDF file containing original western blots for Figure 3—figure supplement 8, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig3-figsupp8-data2-v1.pdf
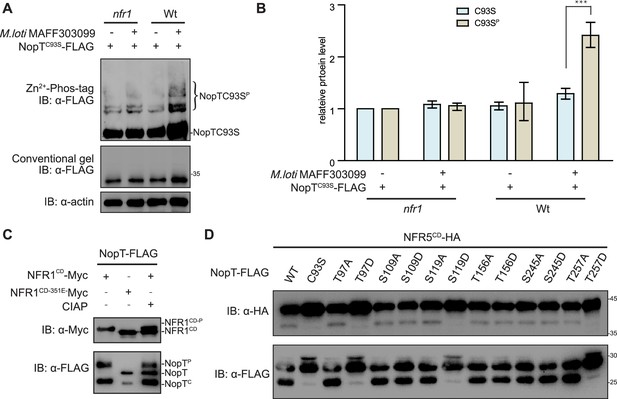
Phosphorylation of NopT by NFR1 suppresses its proteolytic activity.
(A) In vivo phosphorylation assay with proteins expressed in L. japonicus roots (wild-type and nfr1-1 mutant plants) using Zn2+-Phos-tag SDS–PAGE. Phosphorylation of NopTC93S was induced by inoculation with rhizobia (Mesorhizobium loti MAFF303099) and was largely dependent on NFR1. (B) The relative protein amount of each lane, as shown in (A), was quantified with ImageJ software (three biological replicates, student’s t-test; *** indicates statistically significant differences at p<0.01). The value of the control band in each gel was set to 1 for comparison. Values are means ± SEM. (C) NFR1CD but not NFR1CD-K351E phosphorylates NopT in E. coli cells. The phosphorylated full-length form of NopT could be dephosphorylated by calf intestinal alkaline phosphatase (CIAP) (gel band shift). Abbreviations: NFR1CD-P, autophosphorylated NFR1; NopTP, phosphorylated NopT; NopT, nonphosphorylated NopT; NopTC, autocleaved NopT. (D) The phosphorylation sites of NopT identified by liquid chromatography–mass spectrometry (LC–MS) were either substituted to alanine (A) or aspartate (D). The indicated NopT variants were subsequently tested for autocleavage and NFR5CD proteolysis in E. coli cells. Wild-type NopT (WT) and protease-dead NopTC93S were included into the analysis.
-
Figure 4—source data 1
Original files for western blot analysis displayed in Figure 4A, C and D.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig4-data1-v1.zip
-
Figure 4—source data 2
PDF file containing original western blots for Figure 4A, C and D, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig4-data2-v1.zip

NFR1CD phosphorylates NFR5CD in vitro.
The cytoplasmic domains (CDs) of NFR1, NFR5, and kinase-dead NFR1CD-K351E were used for kinase assays in E. coli cells. Proteins were detected by immunoblotting using anti-HA and anti-Myc antibodies. Asterisk indicated the band retardation on the gel representing the phosphorylated NFR5CD proteins.
-
Figure 4—figure supplement 1—source data 1
Original files for western blot analysis displayed in Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig4-figsupp1-data1-v1.zip
-
Figure 4—figure supplement 1—source data 2
PDF file containing original western blots for Figure 4—figure supplement 1, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig4-figsupp1-data2-v1.pdf
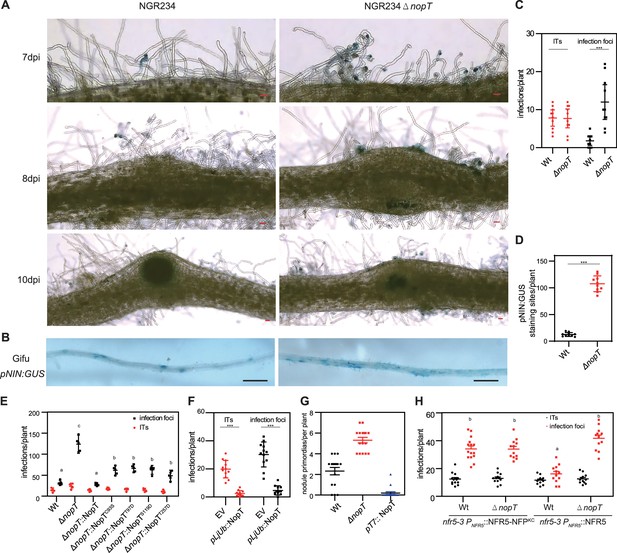
NopT regulates rhizobial infection in L. japonicus.
(A) Analysis of rhizobial infection in L. japonicus roots inoculated with GUS-labeled S. fredii NGR234 (wild-type; WT) or a nopT knockout mutant (NGR234ΔnopT; abbreviated as ΔnopT in other panels). Scale bar = 100 µm. Infection represents both infection focus and infection thread. The infection threads in the root hairs were shown in the left image in the upper panel, while the infection foci were shown in the left image in the middle panel. (B) GUS staining pictures showing roots of L. japonicus plants expressing GUS expression under control of the NIN promoter (pNIN:GUS). Plants were inoculated with NGR234 or NGR234ΔnopT and analyzed at 7 dpi. Scale bar = 1 mm. (C) Infection data (ITs, infection threads) for roots shown in (A) at 7 dpi (n = 10, Student’s t-test; *** indicates statistically significant differences at p < 0.01). (D) Quantification of GUS staining sites for roots shown in (B) (n = 10, Student’s t-test, p < 0.01). (E) Infection data for wild-type roots inoculated with NGR234 (WT), NGR234ΔnopT, and NGR234ΔnopT expressing indicated NopT variants at 7 dpi (n = 5, Student’s t-test: p < 0.01). (F) Analysis of rhizobial infection in hairy roots of L. japonicus (wild-type) expressing GFP (EV, empty vector control) or NopT. Plants were inoculated with DsRed-labeled M. loti MAFF303099 and analyzed at 5 dpi (n = 8, Student’s t-test: p < 0.01). (G) Nodule primordia formation in L. japonicus wild-type roots inoculated with NGR234 (WT), NGR234ΔnopT, or NGR234 over-expressing nopT (pT7:NopT). Roots were analyzed at 14 dpi (n = 16, Student’s t-test: p < 0.01). (H) Expression of NFR5 and NFR5-NFPKC in hairy roots of nfr5-3 mutant plants. Plants were inoculated with GFP-labeled NGR234 (WT) or NGR234ΔnopT and analyzed at 8 dpi. Roots expressing NFR5-NFPKC showed high numbers of infection foci for both strains whereas significant differences were observed for NFR5 expressing roots (n > 10, Student’s t-test: p < 0.01).
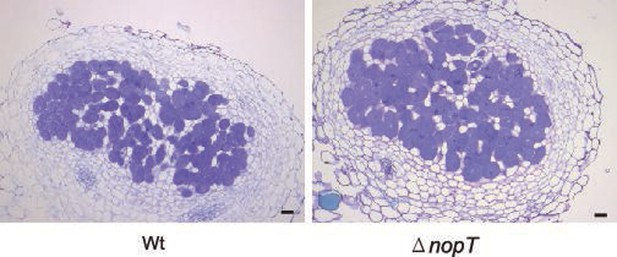
Cross-section images of nodules at 14 dpi after toluidine blue staining.
NGR234 and NGR234ΔnopT mutants inoculated L. japonicus Gifu at 14 dpi. Scale bars correspond to 100 μm.
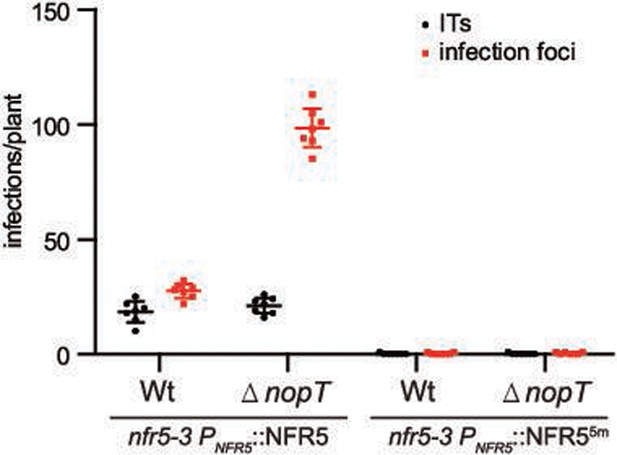
Rhizobial infection in the mutant versions of NFR5.
NFR55m failed to form rhizobial infection (n = 7, Student’s t-test: p < 0.01). The transgenic roots expressing NFR5 or NFR55m in the nfr5-3 mutant under the control of native promoter were inoculated with GFP-labeled wild-type NGR234 and NGR234ΔnopT, respectively. The data were collected 9 days post inoculation. NFR55m represents a mutant version of NFR5 with S283Y, G294Q, Y304S, A310I, and T311Y point mutation.
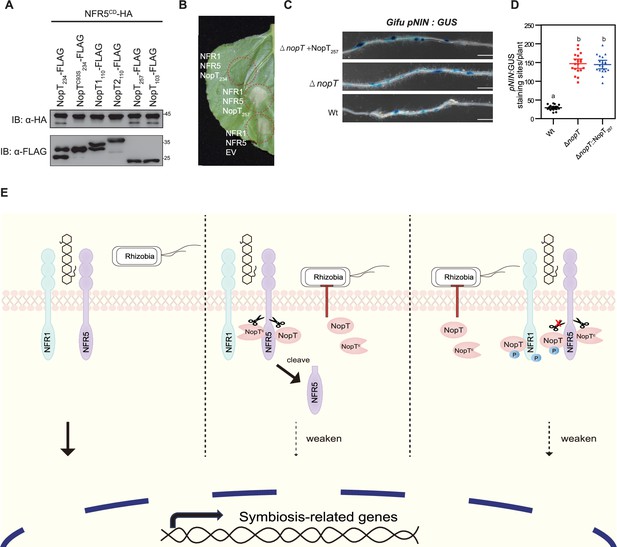
S. fredii NopT proteins cleave NFR5 and working model for NopT of NGR234.
(A) NopT of S. fredii NGR234 and homologs from other rhizobial strains were co-expressed with NFR5CD in E. coli cells (NopT234, NopT of NGR234; NopT1110 and NopT2110, NopT proteins of B. diazoefficiens USDA110; NopT257, NopT of S. frediiUSDA257; NopT103, NopT of S. fredii HH103). Immunoblot analysis indicated NFR5 cleavage by NopT proteins from S. fredii strains. (B) Expression of NopT257 in N. benthamiana could not inhibit the cell death triggered by co-expressed NFR1 and NFR5. (C) L. japonicus Gifu pNIN:GUS plants were inoculated with S. fredii, NGR234, NGR234ΔnopT, and NGR234ΔnopT expressing NopT257 (ΔnopT + NopT257). Roots were subjected to GUS staining at 7 dpi. Scale bar = 2.5 mm. (D) Quantitative analysis of GUS-stained roots shown in panel C (n = 19, Student’s t-test: p < 0.01). (E) A proposed model for NopT of NGR234 interacting with NFRs. NopT and NopTC (autocleaved NopT) proteolytically cleave NFR5 at the juxtamembrane (JM) domain to release the intracellular domain of NFR5 (cleaved NFR5). NFR1 phosphorylates full-length NopT to block its proteinase activity. NopTC cannot be phosphorylated by NFR1.
-
Figure 6—source data 1
Original files for western blot analysis displayed in Figure 6A.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig6-data1-v1.zip
-
Figure 6—source data 2
PDF file containing original western blots for Figure 6A, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/97196/elife-97196-fig6-data2-v1.pdf
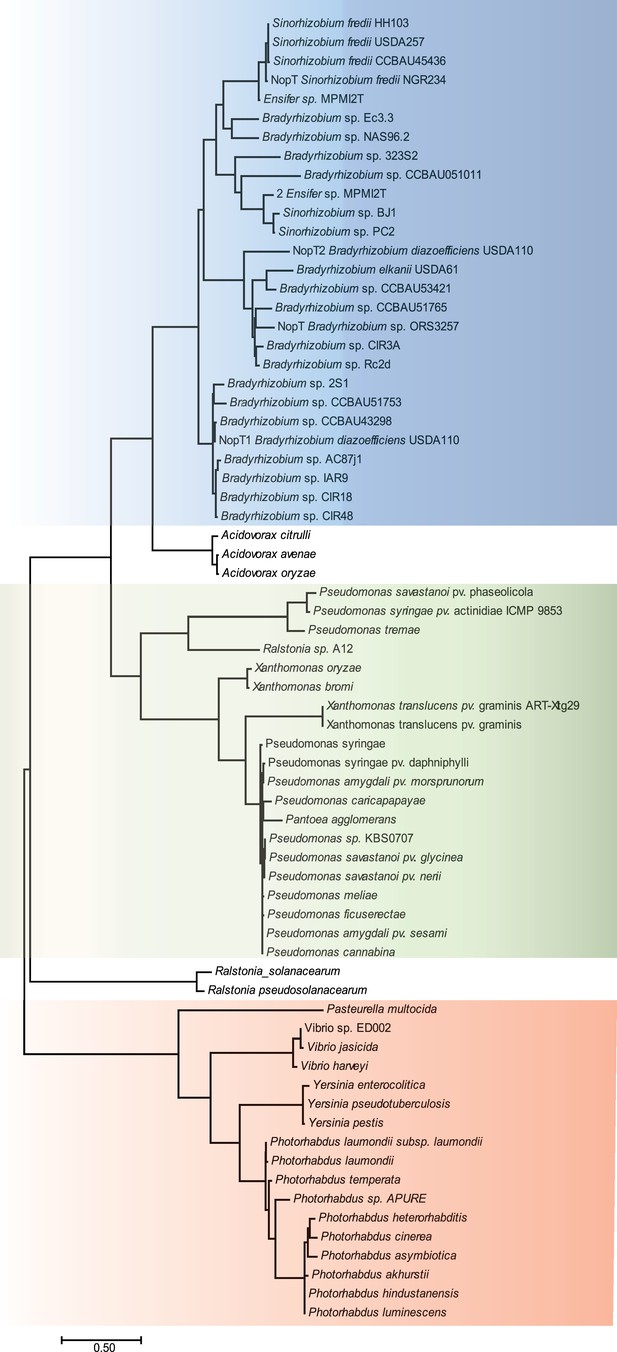
Phylogenetic tree based on the amino acid sequence of NopT homologs from different bacterial species.
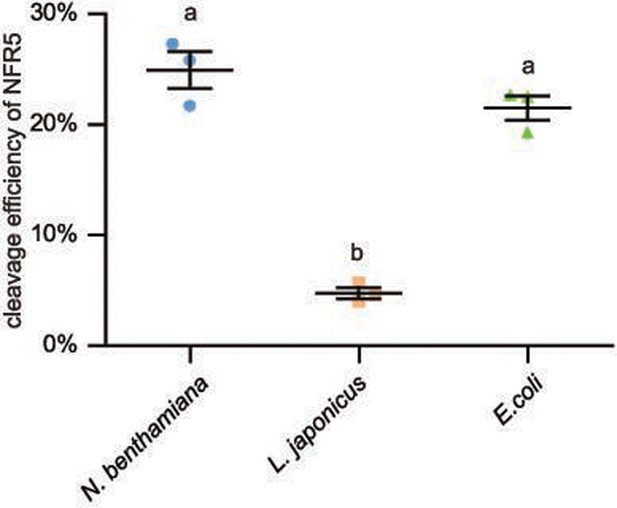
The cleavage efficiency of NFR5 in different system.
The relative cleavage efficiency for experiment displayed in Figure 3A (N. benthamiana), Figure 3B (L. japonicus), and Figure 3D (E. coli) quantified from three biological replicates using ImageJ software. Values are means ± SE (n = 3, Student t-test, letters represent significant differences, p < 0.05).
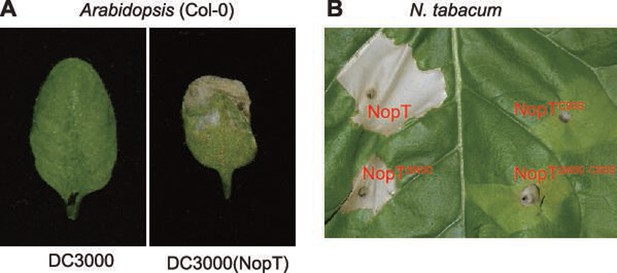

Transient expression of NopT triggers cell death in Arabidopsis thaliana and Nicotiana tabacum.
Pseudomonas syringae pv. tomato DC3000 harboring control plasmids (left) or expressing NopT (right) were infiltrated into Arabidopsis leaves. Pictures were taken 3 days post inoculation. (B) Agrobacterium strains harboring NopT, NopT with an N-terminal 50-amino acid deletion, NopTC93S, or NopTC93S with an N-terminal 50-amino acid deletion were infiltrated into N. tabacum leaves. Pictures were taken 3 days post inoculation.
Additional files
-
Supplementary file 1
Phosphopeptide identification and oligonucleotide used in the study.
(A) Phosphopeptides identified by liquid chromatography–mass spectrometry. (B) Oligonucleotides used in the study.
- https://cdn.elifesciences.org/articles/97196/elife-97196-supp1-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/97196/elife-97196-mdarchecklist1-v1.docx





