A direct experimental test of Ohno’s hypothesis
Figures
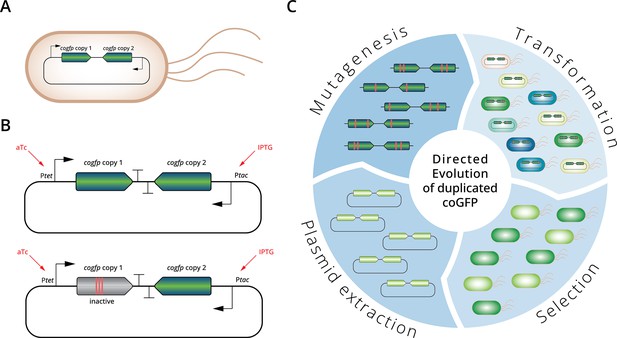
Experimental evolution of a duplicated fluorescent protein.
(A) E. coli cells carry a plasmid containing a duplicated gene coding for coGFP. (B) Upper panel: Plasmid with two copies of the cogfp gene, both under control of independently inducible promoters (Ptet and Ptac); lower panel: control single-copy plasmid with only one active gene copy, the other copy is not fluorescent due to mutations engineered into the chromophore. (C) Overview of the directed evolution experiment of the duplicated cogfp gene (see text for details).
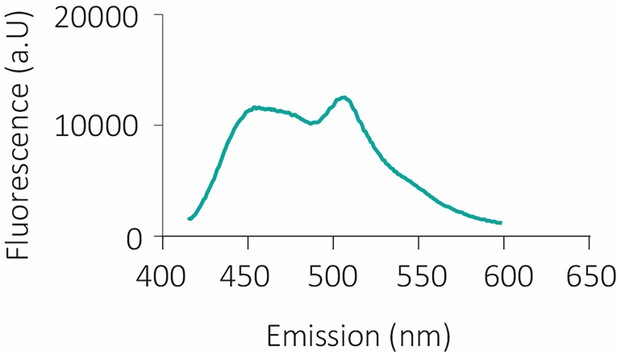
Dual-color-emitting fluorescent protein coGFP.
Emission spectra of coGFP_S147G upon 388 nm excitation and pH 7. Two emission peaks at 456 nm (blue) and 507 nm (green) are detected.
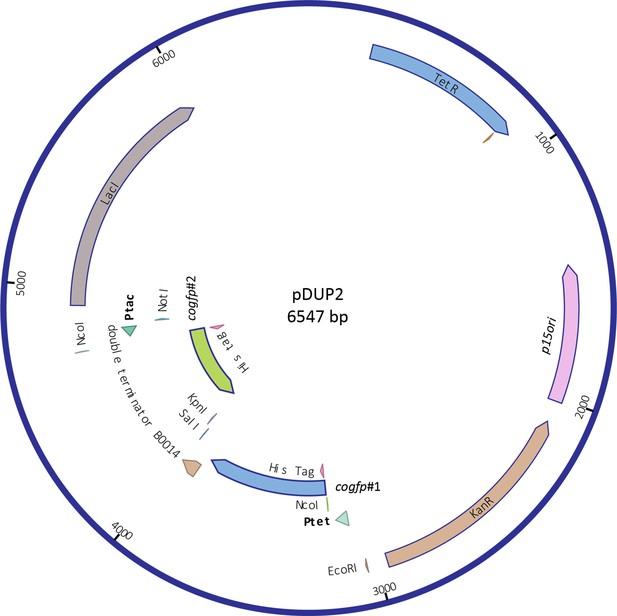
Plasmid map of pDUP2 carrying the duplicated cogfp gene.
The two copies of the cogfp gene are facing each other, separated by a bidirectional terminator. The pDUP2 plasmid also includes the transcriptional repressors LacI and TetR and a kanamycin resistance gene. (#1: cogfp under the Ptet, #2: cogfp under the Ptac).
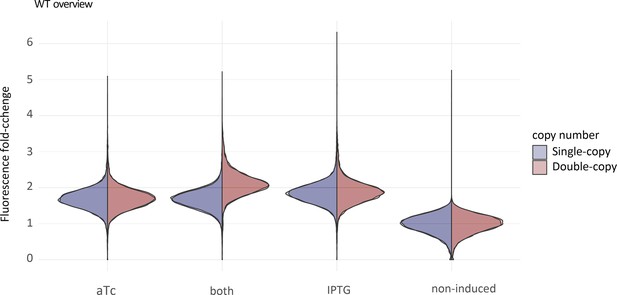
Expression levels of single-copy and double-copy constructs.
Green fluorescence distribution of the ancestral single-copy (blue) and double-copy (red) populations measured by flow cytometry upon the induction with anhydrotetracycline (aTc), isopropyl-β-D-1-thiogalactopyranosid (IPTG), or both normalized to the non-induced controls.
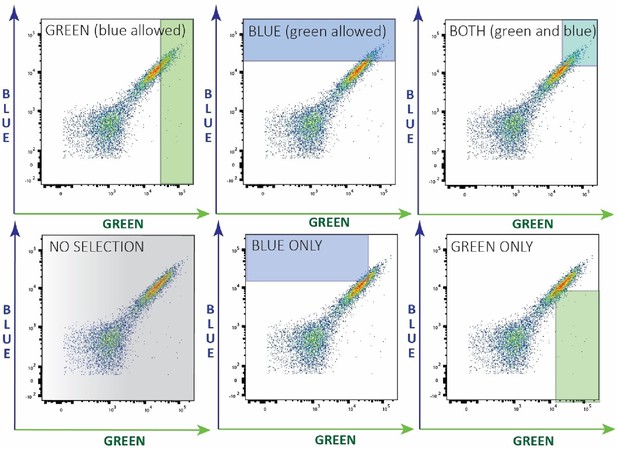
Selection regimes based on cell’s fluorescence phenotypes.
Flow cytometry plots of a first generation library after mutagenesis showing green (AmCyan) vs blue (DAPI) fluorescence. Highlighted regions indicate the gates used for the 6 different selection regimes: green: selection for green (no selection against blue fluorescence); blue: selection for blue (no selection against green fluorescence); green and blue; green-only (selection for green and against blue); blue-only (selection for blue and against green); and no selection for either fluorescence color.
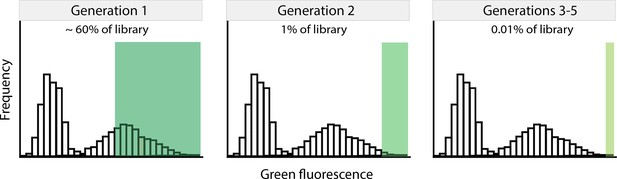
Applied selection stringencies.
Histograms are schemes representing fluorescence distributions of the libraries. Single-copy wild type was used to set the selection threshold in the first generation of the evolution experiment (selected top 60%). In the upcoming generations, the threshold was set on the mutant libraries: selected top 1% in the second and top 0.01% in the following rounds of the evolution experiment.
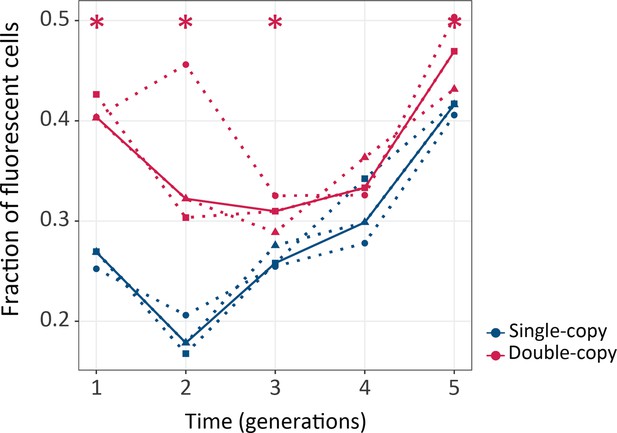
Gene duplication increases mutational robustness.
The vertical axis shows mutational robustness, measured as the percentage of cells that maintain their fluorescence after mutagenesis, as a function of time (in generations of directed evolution) on the horizontal axis. Thick blue and red lines stand for the median fraction of fluorescent cells for single-copy and double-copy mutant libraries, respectively, while dotted lines indicate data from the three biological replicates. One-tailed Mann-Whitney tests, *p≤0.05, n=3. (Figure 2—source data 1).
-
Figure 2—source data 1
Data plotted in Figure 2.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig2-data1-v1.xlsx
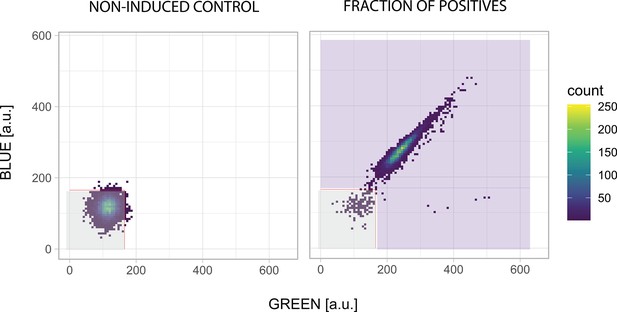
Quantification of fluorescent cells.
Flow cytometry plots showing green (AmCyan) vs blue (DAPI) fluorescence. A non-induced control was used to set the gate (gray). Cells outside this gate (violet area) are considered fluorescent.
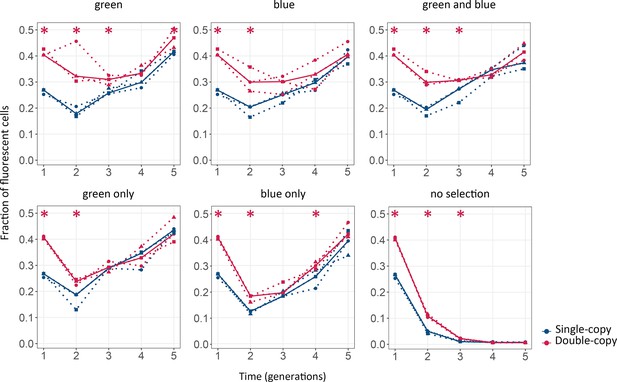
Gene duplication increases mutational robustness.
The vertical axis shows mutational robustness, measured as the percentage of cells that maintain their fluorescence after mutagenesis, as a function of time (in generations of directed evolution) on the horizontal axis. Thick blue and red lines stand for the median fraction of fluorescent cells for single-copy and double-copy mutant libraries, respectively, while dotted lines indicate data from the three biological replicates. The corresponding selection regime is indicated at the top. One-tailed Mann-Whitney tests, *p≤0.05, n=3.
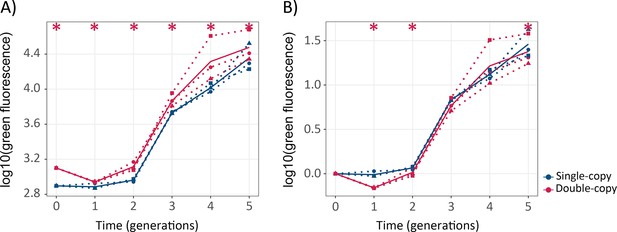
Gene duplication does not lead to significantly faster evolution of green fluorescence.
(A) The vertical axis shows green fluorescence (log10) as a function of time (generations) on the horizontal axis. (B) The vertical axis shows green fluorescence (log10) normalized to the fluorescence of its ancestral population as a function of time (generations) on the horizontal axis. Thick blue and red lines stand for the median fluorescence of single-copy and double-copy mutant populations, respectively, while dotted lines indicate data from the three biological replicates. One-tailed Mann-Whitney tests, *p≤0.05, n=3. (Figure 3—source data 1).
-
Figure 3—source data 1
Data plotted in Figure 3.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig3-data1-v1.xlsx
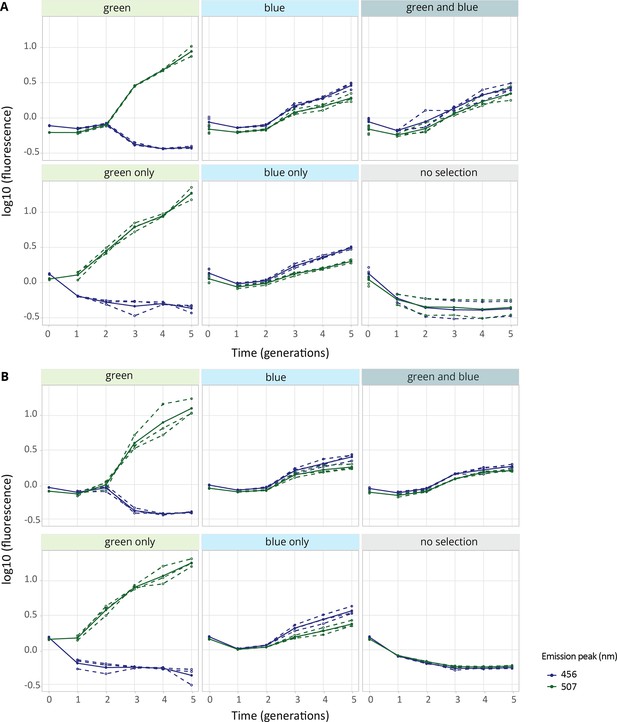
Fluorescence levels during evolution experiment.
Fluorescence levels of the (A) single- and (B) double-copy populations evolved under the indicated selection regimes throughout five generations of the evolution (1-5). Shown is the fluorescence (log10) at blue (456 nm) and green (507 nm) emission peaks upon excitation at 388 nm normalized to the ancestral population. Thick blue and green lines show the mean fluorescence of the populations, while dotted lines indicate data from the three biological replicates.
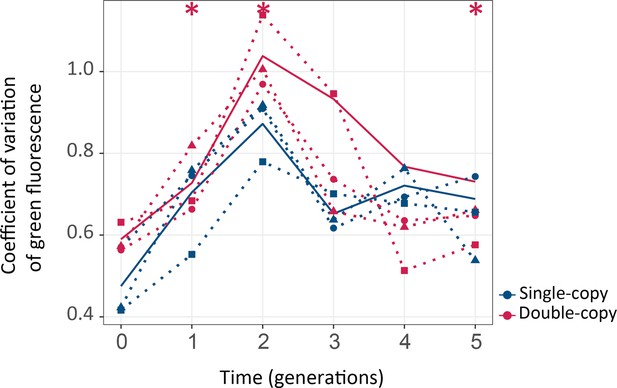
Coefficient of variation of fluorescence.
Populations evolved under selection for green. The vertical axis shows the coefficient of variation (CV) of fluorescence (standard deviation/mean) as a function of time (generations) on the horizontal axis. Thick blue and red lines stand for median variance for single or double-copy populations, respectively, while dotted lines indicate data from the three biological replicates. One-tailed Mann-Whitney tests, *p≤0.05, n=3.
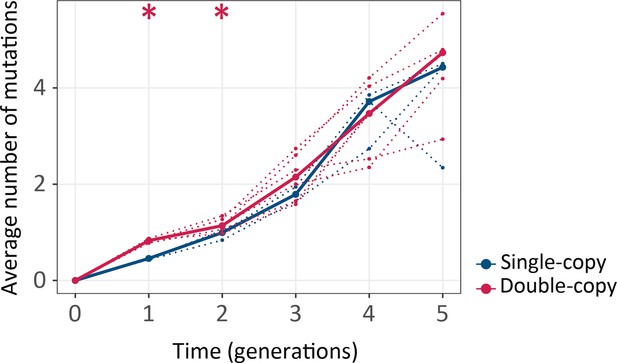
Double-copy populations accumulate more mutations per gene than single-copy populations during the first two generations.
The vertical axis shows the average number of non-synonymous mutations per cogfp gene, as a function of time (in generations of directed evolution) on the horizontal axis. Thick blue and red lines stand for the median numbers of single-copy and double-copy mutant populations respectively, while dotted lines indicate data from the three biological replicates. Mann-Whitney tests *p≤0.05, n=3. (Figure 4—source data 1).
-
Figure 4—source data 1
Data plotted in Figure 4.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig4-data1-v1.xlsx
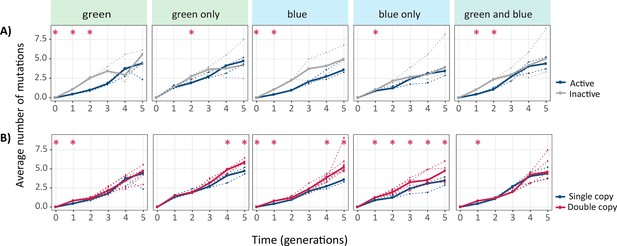
Average number of mutations for single- and double-copy populations evolved under the indicated selection regimes.
The vertical axes show the average number of non-synonymous mutations per cogfp gene, as a function of time (in generations of directed evolution) on the horizontal axes. (A) Average number of mutations per coGFP in single-copy populations (blue: active copy, gray: inactive copy), (B) average number of mutations per coGFP of the active copy in single-copy populations (blue) vs both copies in double-copy populations (pink). Thick lines stand for the median of populations, while dotted lines indicate data from the three biological replicates. Mann-Whitney tests, *p≤0.05, n=3.
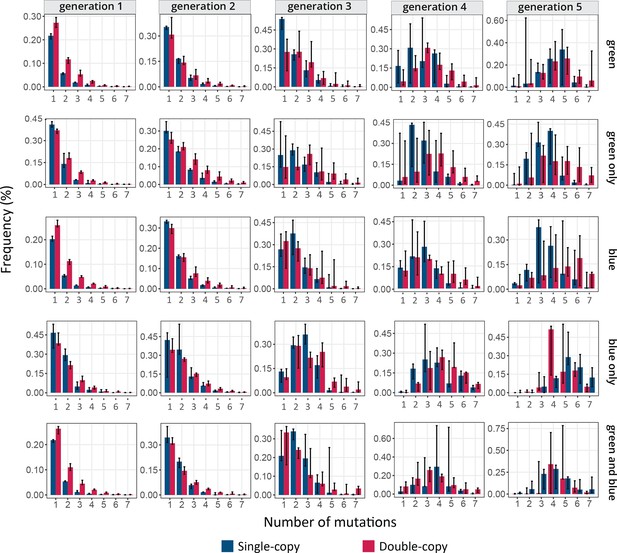
Frequency distribution of the number of mutations.
The vertical axes show the frequency of mutations, as a function of the number of mutations per cogfp gene on the horizontal axes. Blue and red bars stand for single-copy and double-copy populations, respectively. Generations of directed evolution are indicated at the top and selection regimes are indicated on the right. Average and standard deviation of three biological replicates are shown.
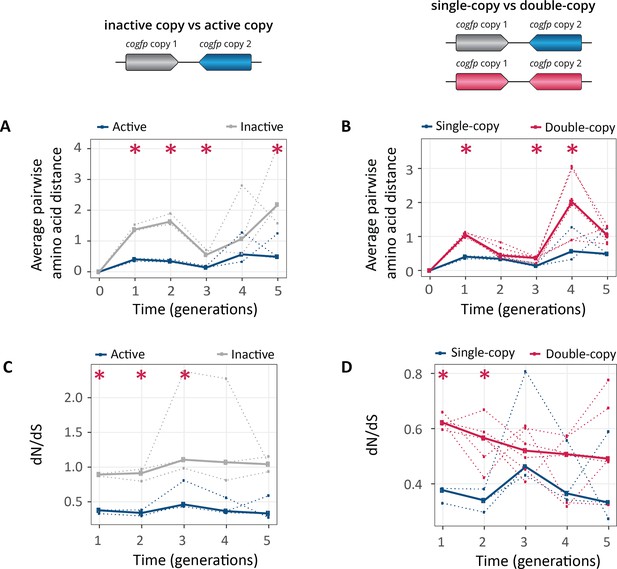
Populations with two gene copies show increased genetic diversity and dN/dS ratios.
The horizontal axes of all panels show time in generations during selection for green fluorescence. (A) Average pairwise amino acid distance for coGFP molecules in single-copy populations (blue: active copy, gray: inactive copy), (B) average pairwise amino acid distance for coGFP molecules of the active copy in single-copy populations (blue) vs both copies in double-copy populations (pink). (C) dN/dS ratio in single-copy populations (blue: active copy, gray: inactive copy), (D) dN/dS ratio of the active copy in single-copy populations (blue) vs both copies in double-copy populations (pink). Thick lines represent the median over three replicate populations, while dotted lines indicate data from the individual biological replicates. *p≤0.05 Mann-Whitney test, n=3. (Figure 5—source data 1).
-
Figure 5—source data 1
Data plotted in Figure 5.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig5-data1-v1.xlsx
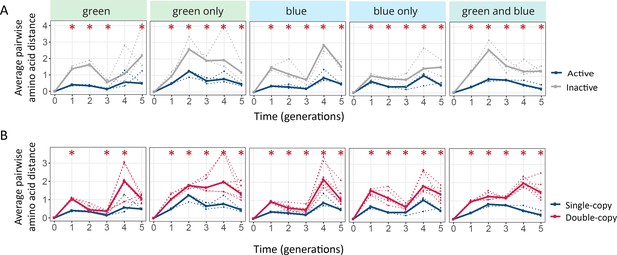
Populations with two gene copies are showing increased genetic diversity.
The horizontal axes of all panels show time in generations for the indicated selection regimes. (A) Average pairwise amino acid distance for coGFP molecules in single-copy populations (blue: active copy, gray: inactive copy), (B) Average pairwise amino acid distance for coGFP molecules of the active copy in single-copy populations (blue) vs both copies in double-copy populations (pink). One-tailed Mann-Whitney tests, *p≤0.05, n=3.
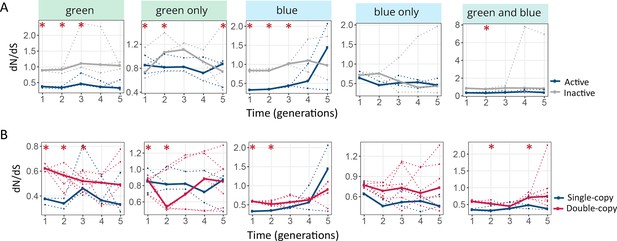
Populations with two gene copies are showing higher dN/dS ratios.
(A) dN/dS ratio in single-copy populations (blue: active copy, gray: inactive copy), (B) dN/dS ratio of the active copy in single-copy populations (blue) vs both copies in double-copy populations (pink). Thick lines represent the median over three replicate populations, while dotted lines indicate data from the individual biological replicates. One-tailed Mann-Whitney tests, *p≤0.05, n=3.
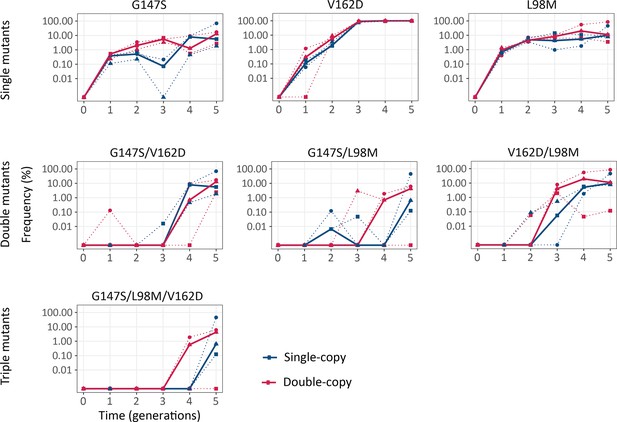
Combination of beneficial mutations emerged earlier in double-copy populations.
The vertical axis shows the frequency of the indicated mutations and combinations thereof in the populations under selection for green fluorescence, as a function of time (in generations of directed evolution) on the horizontal axis. Thick blue and red lines stand for the median frequencies for single-copy and double-copy populations, respectively, while dotted lines indicate data from the three biological replicates. Detailed statistics are reported in Appendix 1—table 2. (Figure 6—source data 1).
-
Figure 6—source data 1
Data plotted in Figure 6.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig6-data1-v1.xlsx
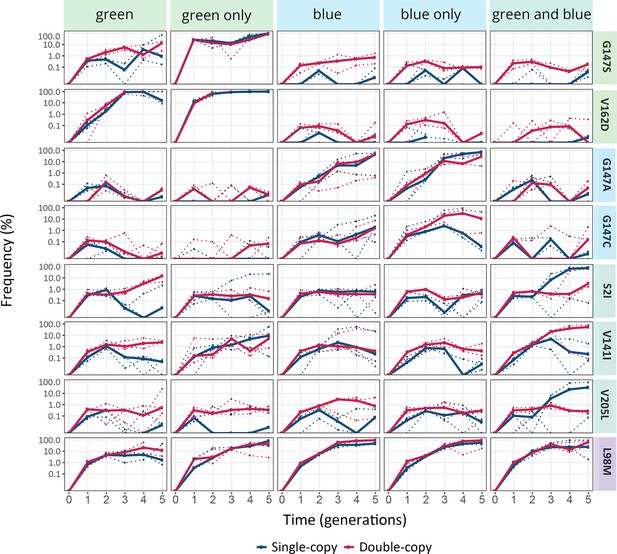
Enriched mutations.
The vertical axes show the frequency of indicated mutations in the populations under indicated selection regimes, as a function of time (in generations of directed evolution) on the horizontal axis. Thick blue and red lines stand for the median frequencies for single-copy and double-copy populations, respectively, while dotted lines indicate data from the three biological replicates.
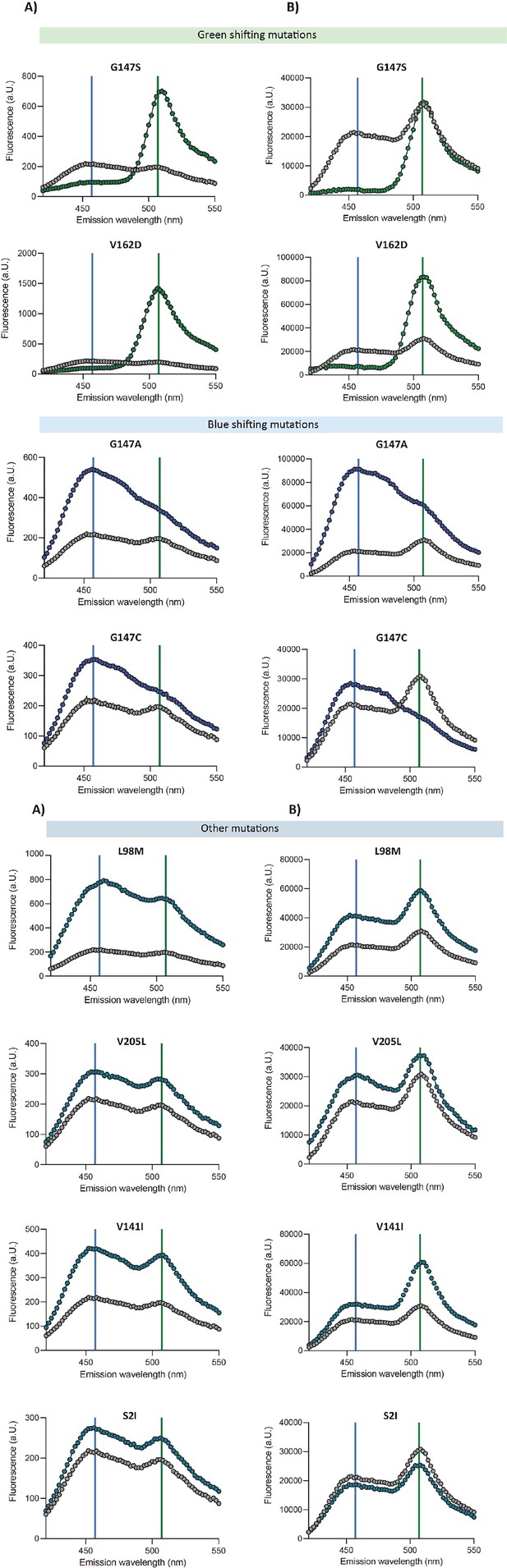
Emission spectra of engineered coGFP variants.
The vertical axes show the fluorescence intensities as a function of the emission wavelengths upon 388 nm excitation of the indicated coGFP variants. Vertical lines indicate blue (456 nm) and green (507 nm) emission peaks. In color, the engineered variants compared to the ancestral protein in gray. (A) Fluorescence of bacterial cells expressing a single copy of the indicated coGFP variant. (B) Fluorescence of purified protein of the indicated coGFP variant (0.05 mg/ml).
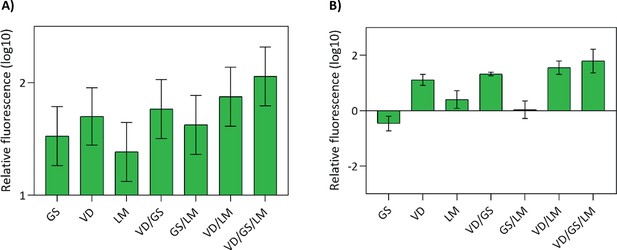
Relative green fluorescence of the engineered variants compared to the ancestral variant.
The vertical axes show the green (507 nm) fluorescence intensities (log10) of the indicated variant relative to that of the ancestral variant upon excitation at 388 nm. (A) Fluorescence of bacterial cells expressing a single copy of the indicated coGFP variant. (B) Fluorescence of purified coGFP protein after size-exclusion chromatography to remove unfolded protein. Tested variants: G147S (GS), V162D (VD), L98M (LM), G147S/V162D (GS/VD), G147S/L98M (GS/LM), V162D/L98M (VD/LM), G147S/V162D/L98M (GS/VD/LM).
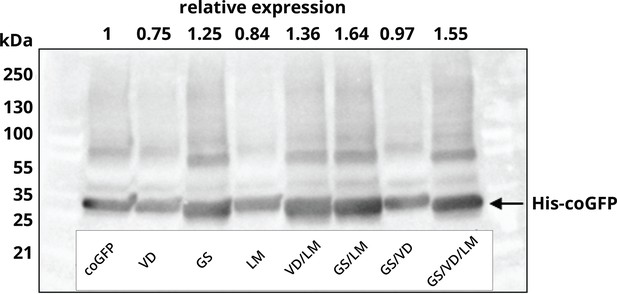

Expression levels of the engineered coGFP variants.
Western blot of the soluble cell lysate fraction using a primary mouse antibody against the His-tag on coGFP and a secondary goat anti-mouse antibody conjugated to horseradish peroxidase (HRP) for chemiluminescent detection. Relative expression levels compared to the ancestral variant coGFP 147G (wt) are indicated at the top. Tested variants: coGFP (wt), G147S (GS), V162D (VD), L98M (LM), G147S/V162D (GS/VD), G147S/L98M (GS/LM), V162D/L98M (VD/LM), G147S/V162D/L98M (GS/VD/LM).
-
Figure 6—figure supplement 4—source data 1
Raw and uncropped blots.
- https://cdn.elifesciences.org/articles/97216/elife-97216-fig6-figsupp4-data1-v1.zip
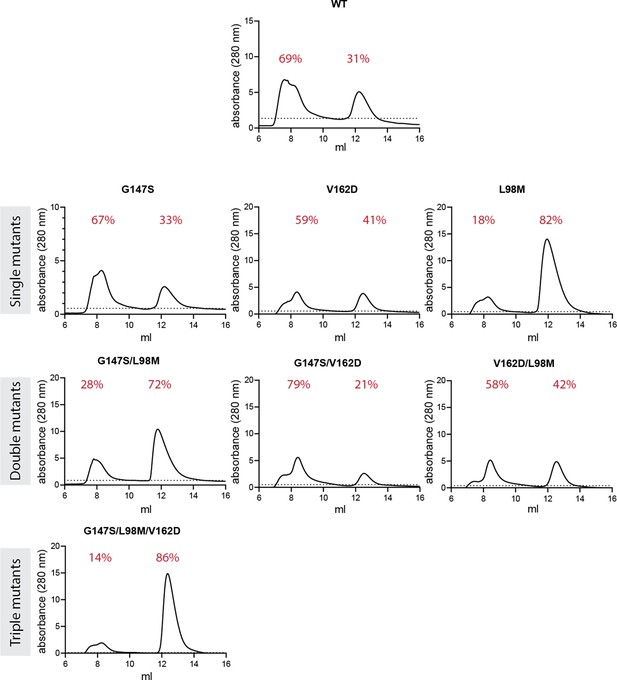
Analysis of fractions of folded protein.
Purified coGFP proteins were run on a size-exclusion column. The vertical axes show the absorbance (280 nm) - a proxy for protein concentration as a function of the elution volume (in ml). Two main peaks are detected: The first one (at ~8 ml) corresponds to aggregated unfolded proteins. The second one (at ~13 ml) corresponds to folded protein. Fractions of unfolded and folded protein are indicated in red numbers above the peaks.
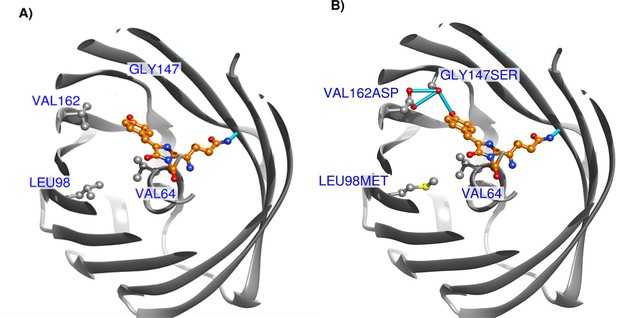
Structure of coGFP.
(A) Homology model of the ancestral coGFP with positions affected by key mutations presented in panel (B), top view. (B) Homology model of the coGFP triple mutant L98M, G147S, V162D chromophore region, top view. The top part of the protein is made invisible for clarity. Gray: coGFP in ribbon representation and carbon atoms, orange: mature chromophore; red: oxygen atoms; blue: nitrogen atoms; yellow: sulfur atoms; bright blue: hydrogen bonds.
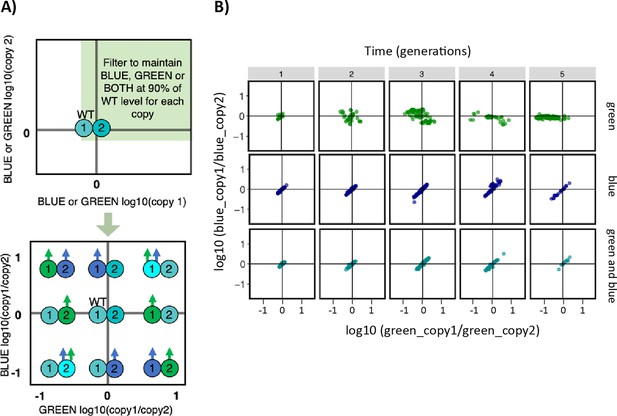
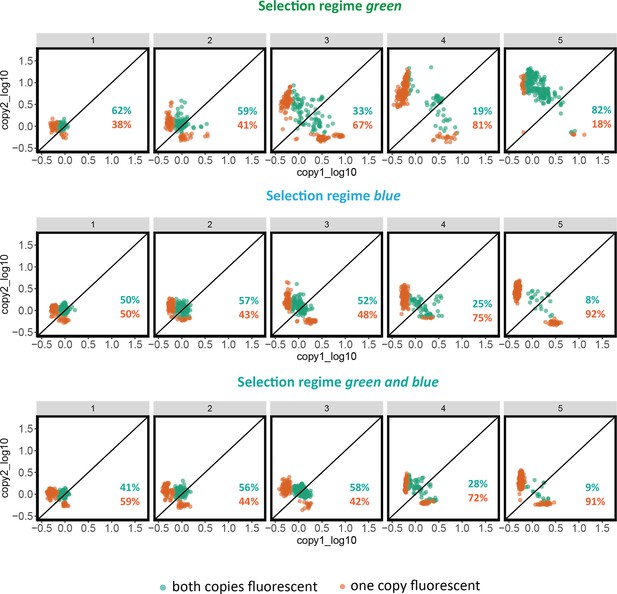
Analysis of variants with two active gene copies.
(A) Scheme to explain the data analysis. Both copies were individually induced, and blue and green fluorescence were measured. This analysis only looks at variants where both copies have green or blue fluorescence of at least 90% of the ancestral variant. For those, we plotted the ratio of copy1/copy2 for blue (x-axis) and green (y-axis) fluorescence (log10 scale). The bottom scheme explains where the different scenarios will lie. For example, data points in the upper left corner have copy 1 improved in green and copy 2 improved in blue compared to the ancestral (WT) protein. Data points in the upper right corner have copy 1 improved in blue and green fluorescence, while copy 2 did not change much. (B) Actual data as explained in (A). We did not find cases where one copy improves in green and the other copy improves in blue.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Cavernularia obesa) | Cogfp | PMID:23468077 | codon-optimised for E. coli | |
| Strain, strain background (E. coli) | NEB5α | New England Biolabs | ||
| Recombinant DNA reagent | pAND (plasmid) | PMID:24316737 | RRID:Addgene_49377 | |
| Recombinant DNA reagent | pAND-MCS (plasmid) | this paper | RRID:Addgene_223514 | sequence and plasmid available via Addgene |
| Recombinant DNA reagent | pDUP (plasmid) | this paper | RRID:Addgene_223515 | sequence and l plasmid available via Addgene |
| Recombinant DNA reagent | pDUP1 (plasmid) | this paper | RRID:Addgene_223516 | sequence and plasmid available via Addgene |
| Recombinant DNA reagent | pDUP2 (plasmid) | this paper | RRID:Addgene_223517 | sequence and plasmid available via Addgene |
| Commercial assay or kit | Illustra TempliPhi DNA Amplification Kit | GE Healthcare | GE Healthcare #25640010 | |
| Commercial assay or kit | SMRTbell Barcoded Adapter Complete Prep Kit 96 | Pacific Biosciences | PacBio # 100-514-900 | |
| Commercial assay or kit | The SMRTbellTM Damage Repair Kit | Pacific Biosciences | PacBio # 100-486-900 | |
| Commercial assay or kit | AMPure PB Kit | Pacific Biosciences | PacBio # 100-265-900 | |
| Commercial assay or kit | NEBuilder HiFi DNA Assembly Master Mix | New England Biolabs | NEB #E2621 | |
| Commercial assay or kit | EcoRI | New England Biolabs | NEB #R0101S | |
| Commercial assay or kit | NotI | New England Biolabs | NEB #R3189S | |
| Commercial assay or kit | SacI | New England Biolabs | NEB #R3156S | |
| Commercial assay or kit | KpnI | New England Biolabs | NEB #R3142S | |
| Commercial assay or kit | NdeI | New England Biolabs | NEB #R0111S | |
| Commercial assay or kit | NcoI | New England Biolabs | NEB #R0193S | |
| commercial assay or kit | DpnI | New England Biolabs | NEB #R0176S | |
| Chemical compound, drug | MnCl2 | Sigma-Aldrich | Sigma #M3634 |
Values of structural stability difference DDG between homology models of coGFP structures and the mutated structures, calculated with Buildmodel FoldX function.
The results pointing to destabilizing effects are colored orange and stabilizing or synergistic are colored green.
| Mutation | DDG by FoldX (kcal/mol) | å of individual single mutant DDG (kcal/mol) | Difference |
|---|---|---|---|
| L98M | –0.25 | –0.25 | 0 |
| G147S | –0.57 | –0.57 | 0 |
| V162D | +3.46 | +3.46 | 0 |
| G147S_V162D | +3.33 | +2.86 | +0.47 |
| L98M_G147S_V162D | +1.95 | +2.64 | –0.69 |
| L98M_V162D | +3.28 | +3.21 | 0.08 |
| L98M_G147S | –1.05 | –0.82 | –0.23 |
Detailed statistics for data reported in Figure 6.
The test is based on a generalized linear model (binomial model, mutation counts vs library type, likelihood ratio test). Beta means enrichment, p-values are bonferroni corrected. Negative beta means enriched in single-copy populations compared to double-copy populations. Positive beta means enriched in double-copy populations compared to single-copy populations.
| Mutation | Generation | Median frequency in single-copy populations | Median frequency in double-copy populations | p-value | beta |
|---|---|---|---|---|---|
| G147S | 1 | 0.38 | 0.52 | 1.00E+00 | 0.38 |
| G147S | 2 | 0.51 | 2.01 | 2.21E-112 | 1.51 |
| G147S | 3 | 0.07 | 5.55 | 0.00E+00 | 3.95 |
| G147S | 4 | 7.89 | 1.26 | 1.00E+00 | –0.02 |
| G147S | 5 | 5.58 | 12.98 | 0.00E+00 | –1.81 |
| V162D | 1 | 0.11 | 0.30 | 1.49E-02 | 1.31 |
| V162D | 2 | 1.98 | 5.85 | 4.99E-244 | 1.06 |
| V162D | 3 | 92.13 | 89.02 | 1.26E-01 | 0.06 |
| V162D | 4 | 99.41 | 98.74 | 6.21E-12 | –1.20 |
| V162D | 5 | 98.99 | 99.23 | 4.25E-03 | –0.41 |
| L98M | 1 | 0.68 | 0.93 | 5.23E-01 | 0.53 |
| L98M | 2 | 4.90 | 4.50 | 1.00E+00 | –0.02 |
| L98M | 3 | 4.27 | 8.29 | 2.27E-122 | 0.58 |
| L98M | 4 | 5.44 | 20.00 | 4.28E-210 | 2.11 |
| L98M | 5 | 10.00 | 11.13 | 7.96E-138 | 0.59 |
| L98M+G147 S | 1 | 0.00 | 0.00 | 1.00E+00 | 1.04 |
| L98M+G147 S | 2 | 0.01 | 0.00 | 4.79E-02 | –1.65 |
| L98M+G147 S | 3 | 0.00 | 0.00 | 9.42E-141 | 4.11 |
| L98M+G147 S | 4 | 0.00 | 0.69 | 5.29E-08 | 2.51 |
| L98M+G147 S | 5 | 0.64 | 4.24 | 0.00E+00 | –2.28 |
| L98M+V162D | 1 | 0.00 | 0.00 | 1.00E+00 | 1.04 |
| L98M+V162D | 2 | 0.00 | 0.00 | 1.00E+00 | –0.67 |
| L98M+V162D | 3 | 0.06 | 4.02 | 0.00E+00 | 3.22 |
| L98M+V162D | 4 | 5.08 | 19.43 | 2.91E-186 | 2.09 |
| L98M+V162D | 5 | 9.67 | 10.95 | 1.11E-114 | 0.54 |
| G147S+V162D | 1 | 0.00 | 0.00 | 1.00E+00 | 1.32 |
| G147S+V162D | 2 | 0.00 | 0.00 | 1.00E+00 | 0.25 |
| G147S+V162D | 3 | 0.00 | 0.00 | 1.00E+00 | –0.34 |
| G147S+V162D | 4 | 7.89 | 0.69 | 1.00E+00 | –0.16 |
| G147S+V162D | 5 | 5.58 | 12.91 | 0.00E+00 | –1.82 |
| L98M+G147S+V162D | 1 | 0.00 | 0.00 | 1.00E+00 | 1.04 |
| L98M+G147S+V162D | 2 | 0.00 | 0.00 | 1.00E+00 | 0.25 |
| L98M+G147S+V162D | 3 | 0.00 | 0.00 | 1.00E+00 | 0.17 |
| L98M+G147S+V162D | 4 | 0.00 | 0.57 | 8.99E-08 | 2.49 |
| L98M+G147S+V162D | 5 | 0.64 | 4.24 | 0.00E+00 | –2.28 |
Primers used in this study.
| Oligonucleotide | Oligonucleotide sequence (5 → 3) |
|---|---|
| Seq_0_f | GAGTTGTAAAACGACGGCCAG |
| Seq_2_r | GAAAGCTGGTCCAAGCGATTG |
| Seq_3_f | CTCATTCGCTAATCGCCAC |
| pBAD_f | GCCGTCACTGCGTCTTTTAC |
| LJM01_f | GTG ATG ATG GTG ATG ATG GCC CAT ATG TAT ATC TCC |
| LJM01_r | GTG ATG ATG GTG ATG ATG GCC CAT GGT ATA TCT CCT |
| LJM02 | GAT ATA CAT ATG GGC CAT CAT CAC CAT CAT CAC AGC ATT CCG GAA AAT |
| LJM03_r | GTTACCAAACTGGAACCGGCGAGCGAAAGCATGTATGTTAG |
| LJM03_f | CTAACATACATGCTTTCGCTCGCCGGTTCCAGTTTGGTAAC |
| LJM04_f | GTTACCAAACTGGAACCGGGCAGCGAAAGCATGTATGTTAG |
| LJM04_r | CTAACATACATGCTTTCGCTGCCCGGTTCCAGTTTGGTAAC |
| LJM05 | GGC CAT CAT CAC CAT CAT CAC |
| LJM06_f | CTTATTCGGCCTTGAATTGATTATATGCGGATTAGAAAAACAACT |
| LJM06_r | AGTTGTTTTTCTAATCCGCATATAATCAATTCAAGGCCGAATAAG |
| LJM07_r | CAACTCGAATTCTTCCACCGTACGTCGAGCGGGAG |
| LJM08_f | GATATAGCGGCCGCAATGGCGGCGCGCCATCGAATG |
| LJM09_f | GTCATGGAATTCGAGTTGTAAAACGACG |
| LJM09_r | GATTATGCGGCCGCGCCGTCACTGCGTCTTTTAC |
| LJM10_f | GCTAGC CCATGG GCCATCATCATCACCATCATAG |
| LJM10_r | CTCTAC GGTACC TTATTACGGTTTGGCAATTGCGGTTTC |
List of site-directed mutagenesis primers.
| Mutation | Forward primer | Reverse primer |
|---|---|---|
| Q74A, Y75S, G76A | GATATTCTGAGCGTTGCATTT GCC AGC GCG AATCGTACCTATACCAGCTATC | GATAGCTGGTATAGGTACGATT CGC GCT GGC AAATGCAACGCTCAGAATATC |
| V169D | GGTGAAGATGTTCTGAGCTATAAAACCCAGAGCACCCATT | CAGAACATCTTCACCAACCAGGGTGCCATCACTAACATAC |
| P142L | GATGGTCTGGTTATGAAAAAAGAAGTTACCAAACTGGAAC | CATAACCAGACCATCTTCCGGGAAACCTTCACCGTTATAT |
| Y173F | CTGAGCTTTAAAACCCAGAGCACCCATTATACCTGTCACA | GGTTTTAAAGCTCAGAACAACTTCACCAACCAGGGTGCCA |
| S9R | CATCACCGCATTCCGGAAAATAGCGGTCTGACCGAAGAAA | CGGAATGCGGAATGCTGTGATGATGGTGATGATGGCCCAT |
| S9I | CATCACATCATTCCGGAAAATAGCGGTCTGACCGAAGAAA | CGGAATGATGAATGCTGTGATGATGGTGATGATGGCCCAT |
| S9C | CATCACTGCATTCCGGAAAATAGCGGTCTGACCGAAGAAA | CGGAATGCAGAATGCTGTGATGATGGTGATGATGGCCCAT |
| L105M | CGTACCATGAGCTTTGAAGATGGTGCCATTGTTAAAGTGG | AAAGCTCATGGTACGTTCAAAGGTAAAACCTTCCGGAAAG |
| G154A | GAACCGGCCAGCGAAAGCATGTATGTTAGTGATGGCACCC | TTCGCTGGCCGGTTCCAGTTTGGTAACTTCTTTTTTCATA |
| G154C | GAACCGTGCAGCGAAAGCATGTATGTTAGTGATGGCACCC | TTCGCTGCACGGTTCCAGTTTGGTAACTTCTTTTTTCATA |
| G154S | GAACCGAGCAGCGAAAGCATGTATGTTAGTGATGGCACCC | TTCGCTGCTCGGTTCCAGTTTGGTAACTTCTTTTTTCATA |
| G154D | GAACCGGACAGCGAAAGCATGTATGTTAGTGATGGCACCC | TTCGCTGTCCGGTTCCAGTTTGGTAACTTCTTTTTTCATA |
| G48S | CTGACCAGTATTCAGAAACTGGATATTCGTGTTATTGAAG | CTGAATACTGGTCAGAATATTACCACCACCAATACCTTCC |
| T79N | AATCGTAACTATACCAGCTATCCGGCAAAAATCCCGGATT | GGTATAGTTACGATTGCCATACTGAAATGCAACGCTCAGA |
| V127L | AAATTTCTGGGCAAAATCAAATATAACGGTGAAGGTTTCC | TTTGCCCAGAAATTTACCATCCTCGATGCTAATATCGCTT |
| G76D | CAGTATGACAATCGTACCTATACCAGCTATCCGGCAAAAA | ACGATTGTCATACTGAAATGCAACGCTCAGAATATCAAAG |
| G42A | ATTGGTGCTGGTAATATTCTGACCGGTATTCAGAAACTGG | ATTACCAGCACCAATACCTTCCATGCTAAAGGCATGACCA |
| H183R | ACCTGTCGCATGAAAACCATTTATCGCAGCAAAAAACCGG | TTTCATGCGACAGGTATAATGGGTGCTCTGGGTTTTATAG |
| K129R | GTGGGCAGAATCAAATATAACGGTGAAGGTTTCCCGGAAG | TTTGATTCTGCCCACAAATTTACCATCCTCGATGCTAATA |
| S155I | CCGGGCATCGAAAGCATGTATGTTAGTGATGGCACCCTGG | GCTTTCGATGCCCGGTTCCAGTTTGGTAACTTCTTTTTTC |
| E168D | GTTGGTGATGTTGTTCTGAGCTATAAAACCCAGAGCACCC | AACAACATCACCAACCAGGGTGCCATCACTAACATACATG |
| G163D | AGTGATGACACCCTGGTTGGTGAAGTTGTTCTGAGCTATA | CAGGGTGTCATCACTAACATACATGCTTTCGCTGCCCGGT |
| L197M | GAAAACATGCCGAAATTTCATTATGTTCATCACCGCCTGG | TTTCGGCATGTTTTCAACCGGTTTTTTGCTGCGATAAATG |
| O229Y | AAACCGTATTAAGAGCTCCAATCGCTTGGACCAGCTTTCC | CTCTTAATACGGTTTGGCAATTGCGGTTTCATGCTGCTCG |
| R206L | CATCACCTCCTGGAAAAAAAAATTGTGGAAGAGGGCTATT | TTCCAGGAGGTGATGAACATAATGAAATTTCGGCAGGTTT |
| S106N | ACCCTGAACTTTGAAGATGGTGCCATTGTTAAAGTGGAAA | TTCAAAGTTCAGGGTACGTTCAAAGGTAAAACCTTCCGGA |
| S177N | ACCCAGAACACCCATTATACCTGTCACATGAAAACCATTT | ATGGGTGTTCTGGGTTTTATAGCTCAGAACAACTTCACCA |
| S155N | CCGGGCAACGAAAGCATGTATGTTAGTGATGGCACCCTGG | GCTTTCGTTGCCCGGTTCCAGTTTGGTAACTTCTTTTTTC |
| S172R | GTTCTGAGATATAAAACCCAGAGCACCCATTATACCTGTC | TTTATATCTCAGAACAACTTCACCAACCAGGGTGCCATCA |
| V148I | AAAGAAATTACCAAACTGGAACCGGGCAGCGAAAGCATGT | TTTGGTAATTTCTTTTTTCATAACCGGACCATCTTCCGGG |
| R206H | CATCACCACCTGGAAAAAAAAATTGTGGAAGAGGGCTATT | TTCCAGGTGGTGATGAACATAATGAAATTTCGGCAGGTTT |
| H183R | ACCTGTCGCATGAAAACCATTTATCGCAGCAAAAAACCGG | TTTCATGCGACAGGTATAATGGGTGCTCTGGGTTTTATAG |
Plasmids used in this study.
| Plasmid name | Description | Source | Addgene number |
|---|---|---|---|
| pAND | Source of backbone | Addgene #49377 | #49377 |
| pAND-MCS | MCS added, NdeI site removed from TetR | This study | #223514 |
| pDUP | cogfp under Ptac | This study | #223515 |
| pDUP1 | cogfp inactive under Ptet, cogfp under Ptac | This study | #223516 |
| pDUP2 | cogfp under Ptet, cogfp under Ptac | This study | #223517 |
Additional files
-
MDAR checklist
- https://cdn.elifesciences.org/articles/97216/elife-97216-mdarchecklist1-v1.docx
-
Source data 1
Summary of single-molecule real-time (SMRT) sequencing results.
Number of reads sequenced by SMRT sequencing and mean number of amino-acid changes per cogfp gene. X, Y, Z: replicate populations. 1–5: generations of evolution.
- https://cdn.elifesciences.org/articles/97216/elife-97216-data1-v1.xlsx