Linalool combats Saprolegnia parasitica infections through direct killing of microbes and modulation of host immune system
Figures
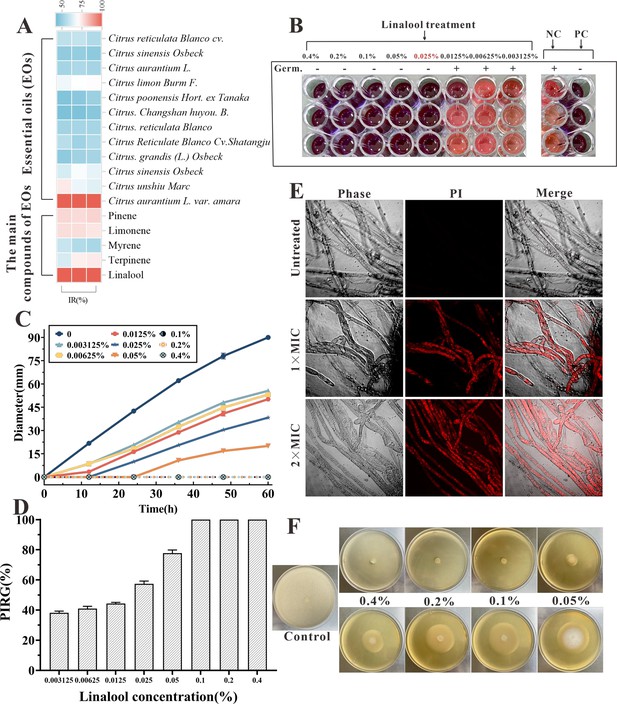
The in vitro anti-oomycetes activity of linalool against S. parasitica CQT2.
(A) Inhibition rate (IR) of essential oils (EOs) and their major components against S. parasitica CQT2. (B) Determination of minimum inhibitory concentration (MIC) of linalool against S. parasitica spores, employing malachite green (2.5 mg/ml) as a positive control (PC) and Tween 20 as a negative control (NC). (C) Effects of linalool on mycelium growth inhibition over a 60-hr period at varying concentrations. (D) Examination of the percentage inhibition of radial growth (PIRG %) in mycelium treated with linalool after 60 hr. (E) Effects of linalool on the viability of S. parasitica CQT2 through a prodium iodide (PI) staining assay was conducted under three conditions: Control (no linalool), 1× MIC (0.05%) linalool treatment, and 2× MIC (0.1%) linalool treatment. (F) Effects of different concentrations of linalool on mycelium growth on potato dextrose agar (PDA) plates.
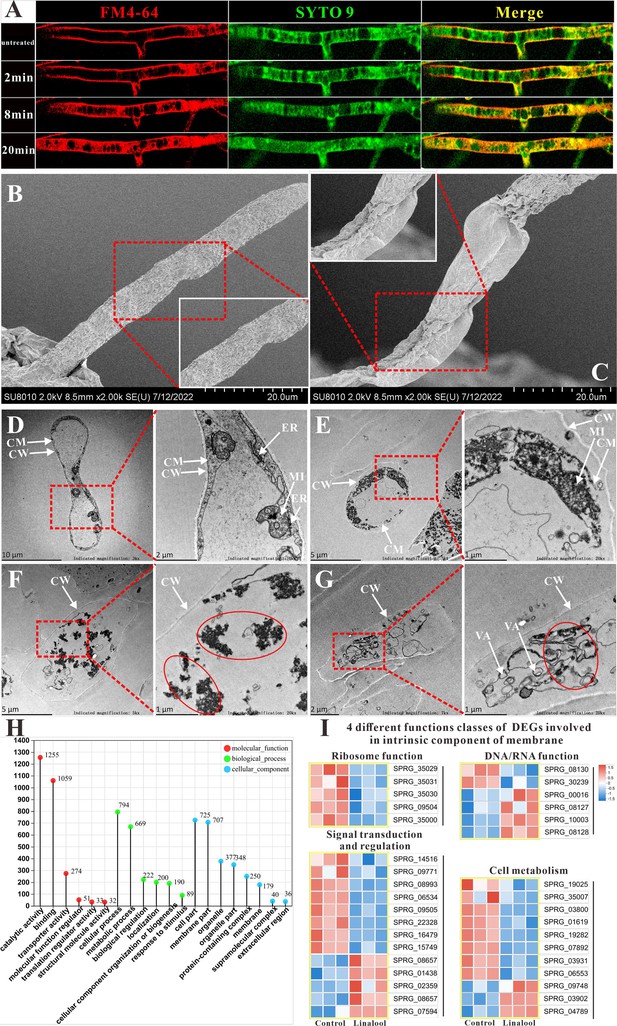

Effects of linalool on the cell membrane integrity of S. parasitica CQT2.
(A) Observation of the effect of linalool at 6× minimum inhibitory concentration (MIC) (0.3%) on the cell membrane of S. parasitica CQT2 using the FM4-64 and SYTO 9 staining assay. Scanning electron microscopy (SEM) images of S. parasitica CQT2 mycelium without (no linalool, B) and with 1× MIC (0.05%, C) linalool treatment. Transmission electron microscopy (TEM) images of S. parasitica CQT2 mycelium without (no linalool, D) and with 1× MIC (0.05%, E–G) linalool treatment. CM: cell membrane; CW: cell wall; ER: rough endoplasmic reticulum; MI: mitochondria; VA: vacuoles. (H) The Gene Ontology (GO) classification of differentially expressed genes (DEGs). (I) Four different functions classes (linked to ribosome function, involved in DNA/RNA function, connected to signal transduction and regulation, and pertaining to cell metabolism) of DEGs involved in intrinsic component of membrane.
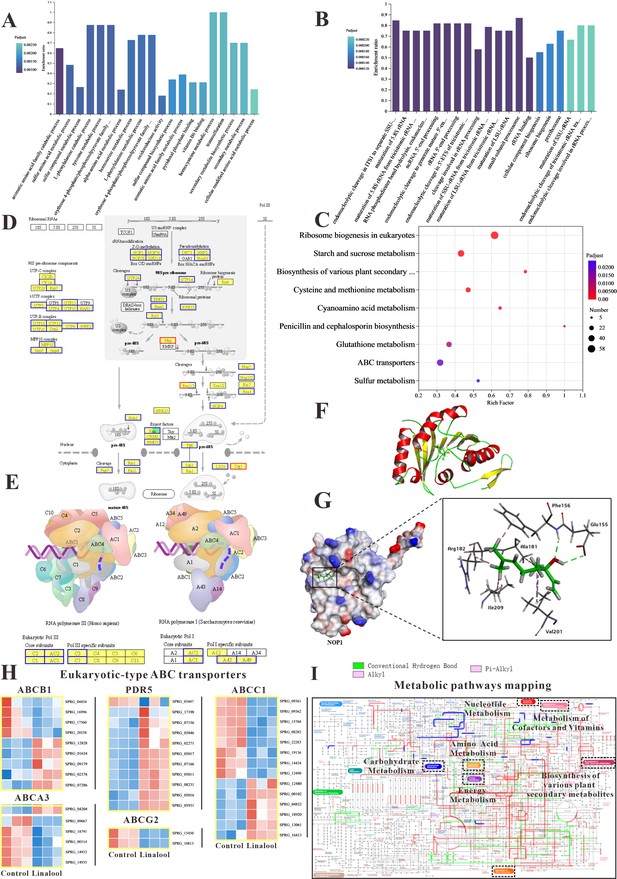
The in vitro anti-oomycete mechanisms revealed by transcriptome analysis.
(A, B) The Gene Ontology (GO) enrichment of up- and downregulated differentially expressed genes (DEGs). (C) The Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment of DEGs. (D) Comparison of ribosome biogenesis in eukaryotes pathway between the linalool treated mycelium and the control group. (E) Comparison of RNA polymerase pathway between the linalool treated mycelium and the control group. (F) The tertiary structure of NOP1 was displayed using a cartoon representation. (G) Molecular docking of linalool with NOP1 was performed by enlarging the regions binding to the NOP1 activation pocket to showcase the detailed amino acid structures, which were presented using a surface model, while the small molecule was displayed with a ball-and-stick representation. (H) Comparison of ABC transporters pathway between the linalool treated mycelium and the control group. The red squares represented upregulated genes, and the blue squares represented downregulated genes. (I) Visual analysis of metabolic pathways with iPath3.0. The figure represented gene set annotated pathways, red and green represented pathways annotated by genes in different gene sets, respectively, and blue represented pathways co-annotated by genes in two gene sets.
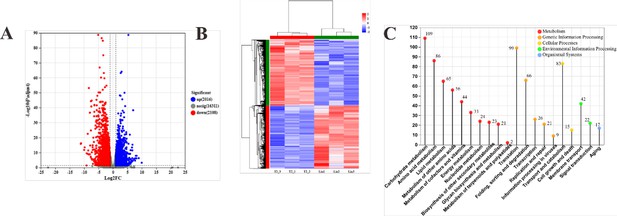
Global transcriptomic analysis between the treated (mycelium treated with linalool) and control group (untreated with linalool).
(A) The number of differentially expressed genes (DEGs). (B) Heat map of DEGs. The red cluster represented upregulated genes, whereas the blue cluster represented downregulated genes. (C) The Kyoto Encyclopedia of Genes and Genomes (KEGG) classification of DEGs.
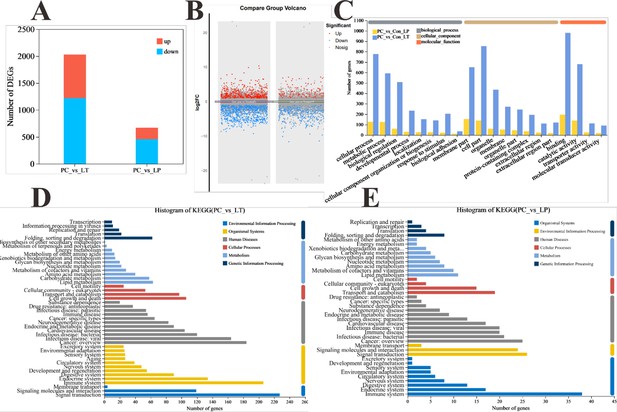
Global transcriptomic analysis after S. parasitica infection and linalool treatment in grass carp.
(A) The number of differentially expressed genes (DEGs). (B) The scatter diagram of DEGs. (C) Gene Ontology (GO) classification of DEGs in each group. (D) Kyoto Encyclopedia of Genes and Genomes (KEGG) classification of DEGs in group LT. (E) KEGG classification of DEGs in group linalool prophylactic (LP).
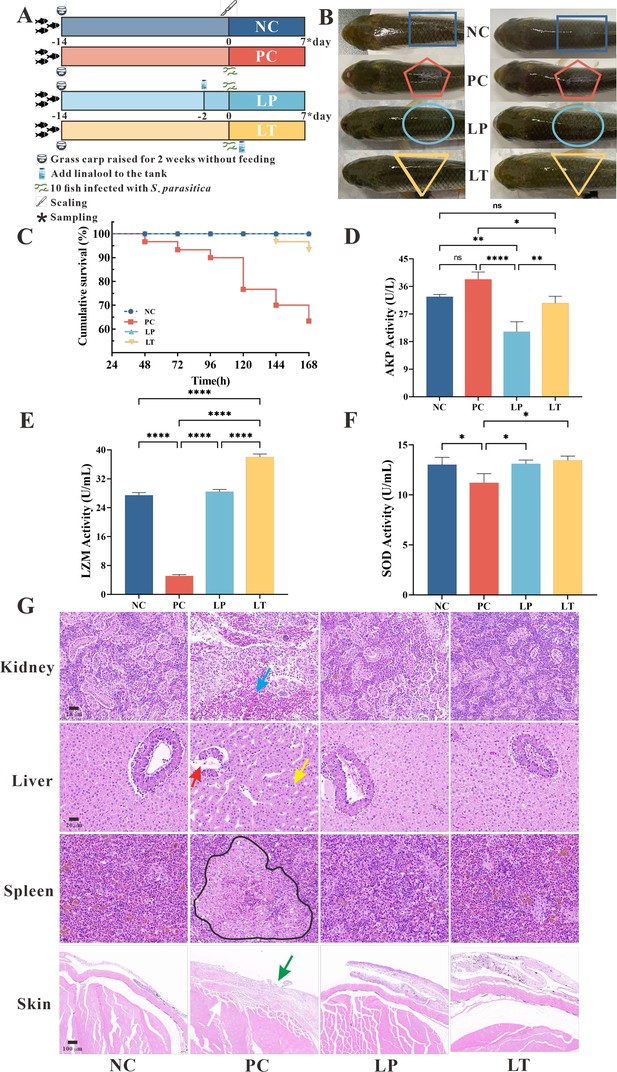
Protective effect of linalool on grass carp infected with S. parasitica.
(A) The experimental design. Grass carp were raised for 2 weeks without feeding, fish without infection and linalool (negative control group, NC), fish infected with S. parasitica (positive control group, PC), and 10 fish uninfected soaked water containing linalool for 2 days and then 1 × 106 spores/ml secondary zoospores were added (linalool prophylactic group, LP), and fish infected with S. parasitica soaked for 7 days in water containing linalool (linalool therapeutic group, LT). (B) The symptoms of S. parasitica infection in grass carps of different groups. (C) The survival rates of grass carp infected with S. parasitica of different groups. (D–F) Alkaline phosphatase (AKP), lysozyme (LYZ), and superoxide dismutase (SOD) activities in serum of grass carp of different groups. The results were presented as the means ± SD and were analyzed using one-way ANOVA followed by post hoc tests (*p < 0.05, **p < 0.01, ****p < 0.0001). (G) Histopathological analysis grass carp tissues in different groups. The arrows of different colors indicated: inflammatory cell infiltration in the kidney (blue arrow), cytoplasmic pyknosis (red arrow), nuclei displaced toward one side (yellow arrow), the red and white pulp was poorly demarcated, and a larger volume of melano-macrophage centers (black area), critical damage to the epithelium (green arrow) and myofiber (white arrow).
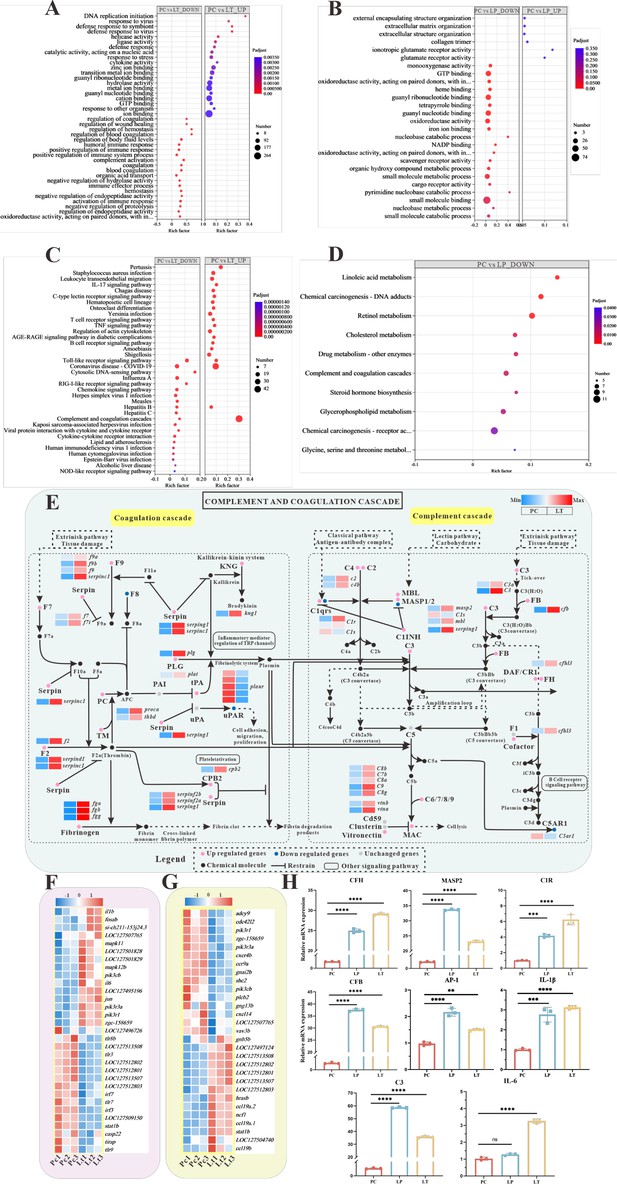
Global transcriptomic analysis after S. parasitica infection and linalool treatment in grass carp and in-depth analysis of crucial Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and differentially expressed genes (DEGs).
(A, B) The Gene Ontology (GO) enrichment of up- and downregulated DEGs in linalool therapeutic (LT) and linalool prophylactic (LP) groups. (C, D) The KEGG enrichment of up- and downregulated DEGs in LT and LP groups. (E) Complement and coagulation cascades pathway between S. parasitica infection and linalool treatment in the spleens of grass carp. (F) Thirty-one DEGs related to Toll-like receptor signaling pathway. (G) Twenty-nine DEGs related to chemokine signaling pathway. The red squares represented upregulated genes, and the blue squares represented downregulated genes. (H) Expression levels of cfh, masp2, c1r, c3, cfb, ap-1, il-1β, and il-6 in the spleen revealed variations among the positive control (PC), LP, and LT groups. For RT-qPCR, the results were presented as the means ± SD and were analyzed using independent t-tests (**p < 0.01, ***p < 0.001, ****p < 0.0001).
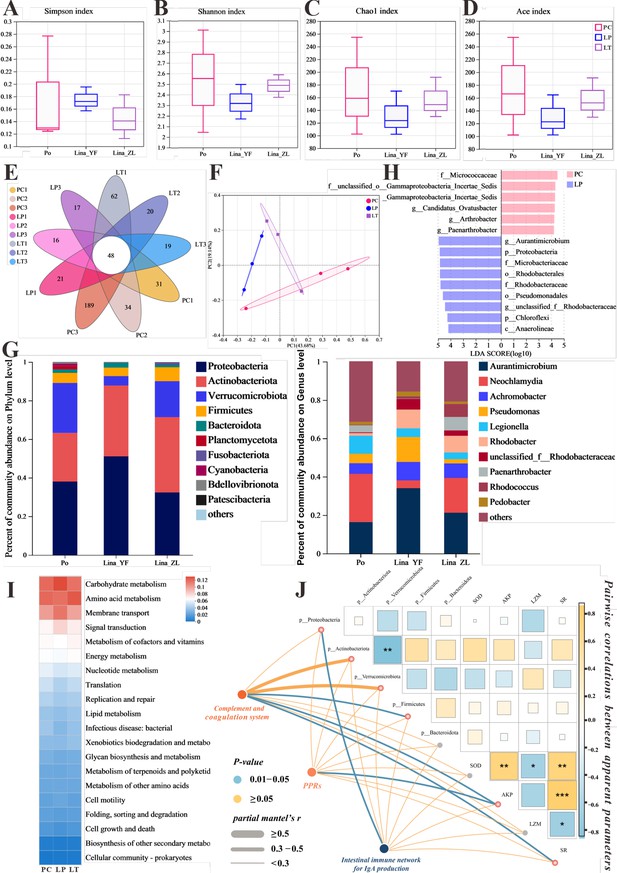
The effect of linalool on regulating gut microbiota of grass carp infected with S. parasitica and correlation analysis.
(A–D) The α diversity index comparison among the different groups. (E) The OTUs petal map of positive control (PC), linalool prophylactic (LP), and linalool therapeutic (LT) groups. (F) Principal coordinate analysis (PCoA) using Bray–Curtis distance revealed variations among the PC, LP, and LT groups (ANOSIM R = 0.3086, p = 0.061). (G) Relative abundance of the top 10 species in the gut from the different groups (phylum and genus levels). (H) Column chart of LDA value distribution. Discriminative biomarkers identified by linear discriminant analysis effect size (LEfSe) with logarithmic LDA score greater than 3.0. (I) Heat map of the differences in predicted functional metabolisms within gut bacterial Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. (J) The correlation network illustrates potential associations among immune genes, microbial species, and physical characteristics. Pairwise Pearson's correlation coefficients are visualized with color gradients and block sizes, where *|r| ≥ 0.3 with p < 0.05, **|r| ≥ 0.5 with p < 0.01, and ***|r| ≥ 0.7 with p < 0.001.
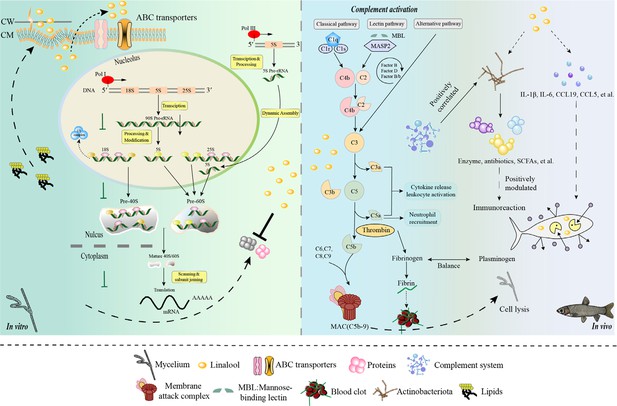
Model diagram of the mode of action of linalool on S. parasitica and grass carp.
In vitro, (1) Linalool influenced DNA transcription, tRNA transport, rRNA processing and maturation (5.8S, 18S, and 25S), and the biogenesis and assembly of ribosome subunits (40S and 60S) in the cell, which might lead to the reduction of S. parasitica growth; (2) Linalool disrupted the cell membrane, and the upregulation of glycerophospholipid metabolism likely represents the cell’s response to cope with this damage; (3) ABC transporters contributed to metabolic resistance by pumping linalool out of the cell. In vivo, (1) Linalool enhanced the complement and coagulation system which in turn activated host immune defense and lysate S. parasitica cells; (2) Linalool promoted wound healing, tissue repair, and phagocytosis to cope with S. parasitica infection; (3) Linalool positively modulated the immune response by increasing the abundance of beneficial Actinobacteriota; (4) Linalool stimulated the production of inflammatory cytokines (il-1β and il-6) and chemokines (ccl19 and ccl5) to lyse S. parasitica cells.
Tables
Primers for qPCR were used in this study.
| Primer name | Primer sequence 5′–3′ |
|---|---|
| c3-F | ACAAGCCCATCTACACGCCT |
| c3-R | GAAACCTCAGGAATGGCGTA |
| cfb-F | AGAATGGAGAGGTGACCCCA |
| cfb-R | CCGCTATCCTGACACACTCG |
| c1r-F | GGGTTTCGGAGGGTTTGA |
| c1r-R | CCTGTATTGCCACATCTTTCA |
| cfh-F | ATAGGAGGGAAATGTGGGC |
| cfh-R | TCCCGTCCACGCCATC |
| masp2-F | TGGGCACCAGGTTCATCG |
| masp2-R | GCTGTTTGACTACGCCCATCTT |
| il-1b-F | ATCTCCTCGTCTGCTGGGTG |
| il-1b-R | TTGAGAGTTTCAGTGACCTCCTTC |
| il-6-F | AAAATCTGACCTCAACCCTGG |
| il-6-R | TGTCCACCCTTCCTCTTGCT |
| ap-1-F | TCTCACTCAACAGATGCCAAAC |
| ap-1-R | CTTAGCGGCTGCGAGTTTAT |
| β-ActinF | GCTATGTGGCTCTTGACTTCG |
| β-ActinR | GGGCACCTGAACCTCTCATT |





