Reconstructing the phylogeny and evolutionary history of freshwater fishes (Nemacheilidae) across Eurasia since early Eocene
Figures
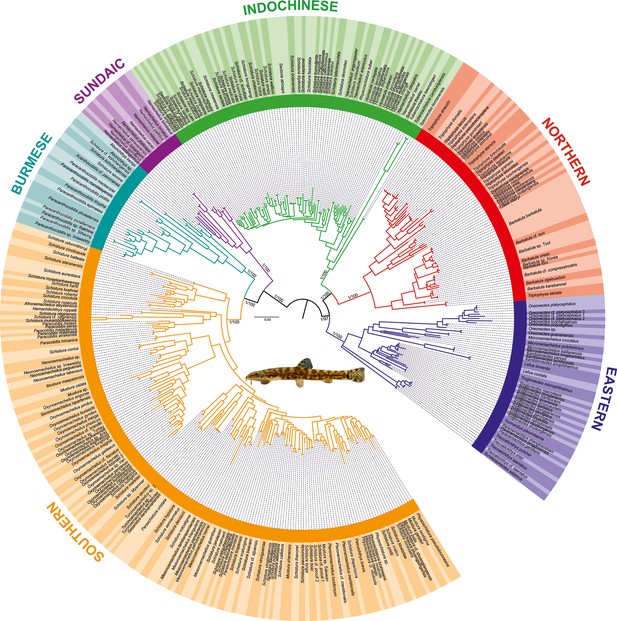
Phylogenetic tree of Nemacheilidae, including all 471 analysed nemacheilid specimens, reconstructed in MrBayes based on six loci.
The topologies of Bayesian inference (BI) and maximum likelihood (ML) trees were congruent. The values at the basal nodes correspond to Bayesian posterior probabilities and ML bootstrap supports, respectively. Only the ingroup is shown. The nemacheilid pictured in the centre is Barbatula sp. Korea.
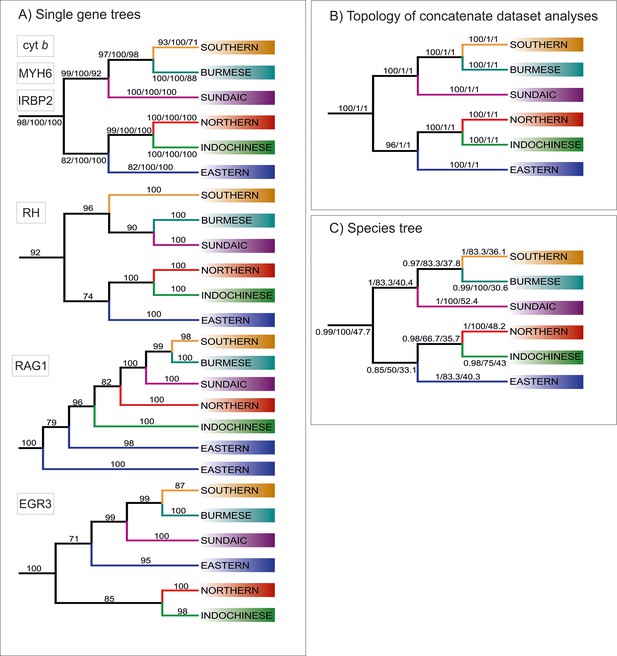
Topologies of main clades derived from various analyses.
(A) Topologies recovered by ML analyses of single gene trees, with values at branches indicating bootstrap supports. Topologies congruent to the concatenated data-derived as well as species tree is revealed by Cyt b, MYH6 and IRBP2. Three slightly different topologies were derived from RH, EGR3 and RAG1 (B) Topology obtained from analyses of concatenate dataset, the values at the branches correspond to ML/MrBayes PP/BEAST PP supports, respectively. (C) Topology inferred by ASTRAL-III using the unrooted ML single gene trees as input. The values at the branches correspond to support values/gCF/sCF.
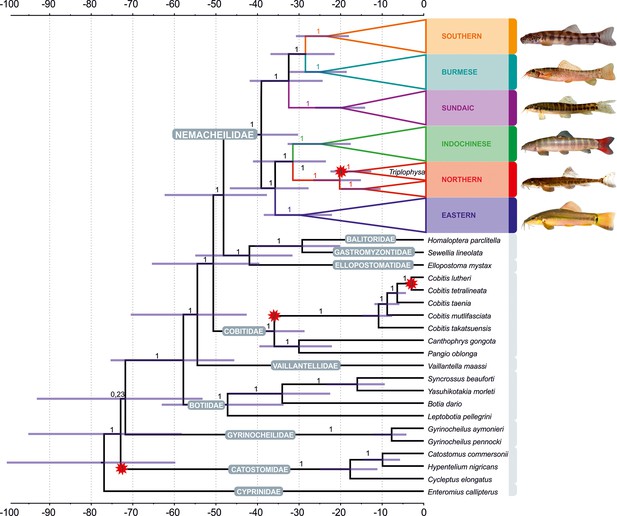
Divergence time estimation resulting from Bayesian divergence time analysis of concatenated dataset in BEAST 2: maximum clade credibility (MCC) tree of the whole dataset, with the Nemacheilidae ingroup collapsed into main clades.
Red stars indicate calibration points, values at branches represent posterior probabilities (PPs lower than 0.90 not shown) and blue horizontal bars denote 95% HPD. The time scale is in millions of years. Pictures on the right show one example species for each clade; from top: Schistura hypsiura (Southern Clade), Paracanthocobitis pictilis (Burmese Clade), Nemacheilus chrysolaimos (Sundaic Clade), Schistura yersini (Indochinese Clade), Triplophysa stenura (Northern Clade), and Traccatichthys taeniatus (Eastern Clade).
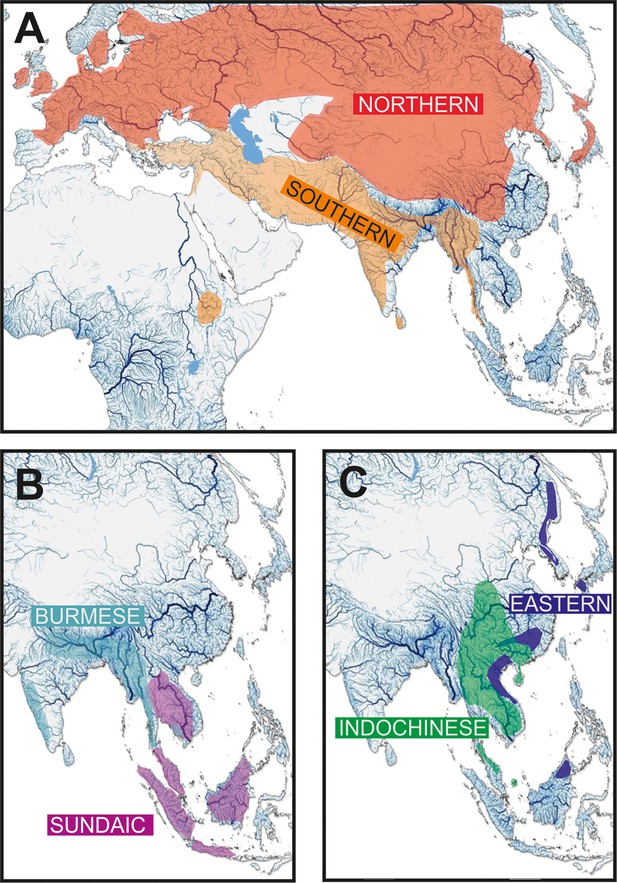
Geographic distribution of the six major clades of Nemacheilidae.
(A) The Northernand Southern Clades, (B) The Burmese and Sundaic Clades, (C) The Eastern and Indochinese Clades.
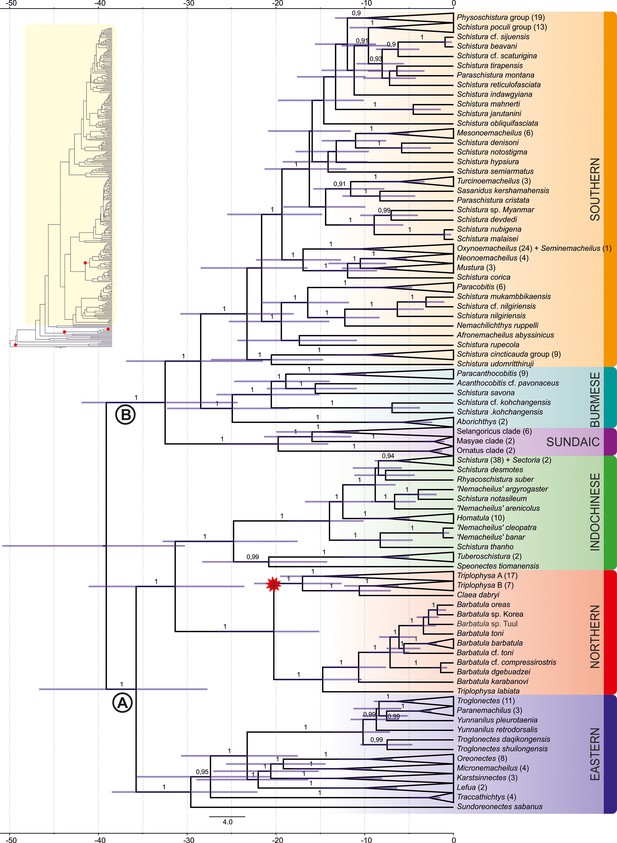
Divergence time estimation: ingroup of the maximum clade credibility (MCC) tree resulting from Bayesian divergence time analysis of concatenated dataset in BEAST 2.
The red star indicates internal calibration point, the values at the nodes represent posterior probabilities (PPs lower than 0.90 not shown), and the blue bars relevant 95% HPD. The time scale is in millions of years. Inset in the top-left corner shows complete tree: Nemacheilidae is highlighted in yellow, while the outgroup is not highlighted; all calibration points are indicated by red stars.
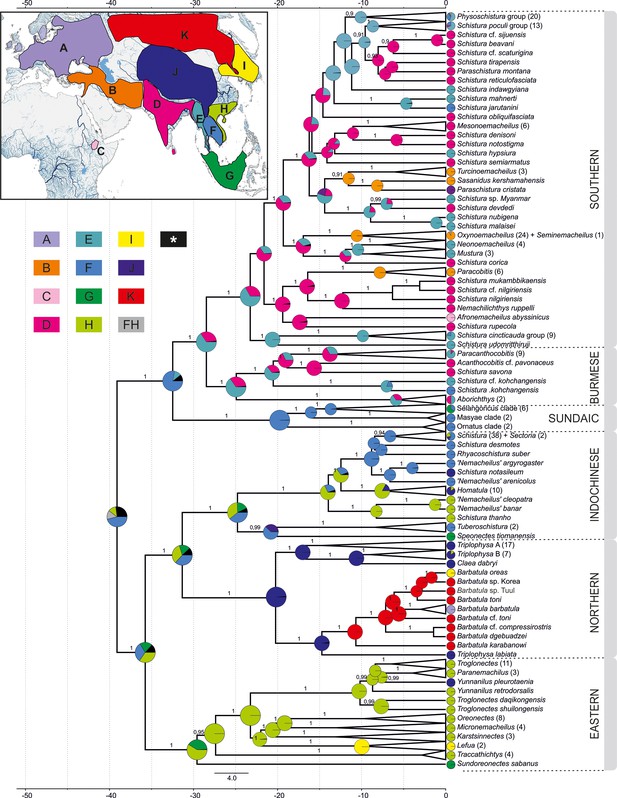
Reconstruction of the biogeographical history of Nemacheilidae.
The inset map in the top-left corner shows the defined biogeographical regions of Eurasia recognised in the analysis; below the map the colour – region code for the main tree. Grey colour indicates combination of two regions as ancestral (FH); black colour indicates ancestral ranges identified with likelihood <10%. Main tree: ancestral range reconstruction with use of DEC+J model based on the phylogeny derived from BEAST2. Pie charts at the nodes indicate relative likelihood of ancestral ranges of most recent common ancestors (MRCA), while the current distribution of taxa is indicated at the tips.
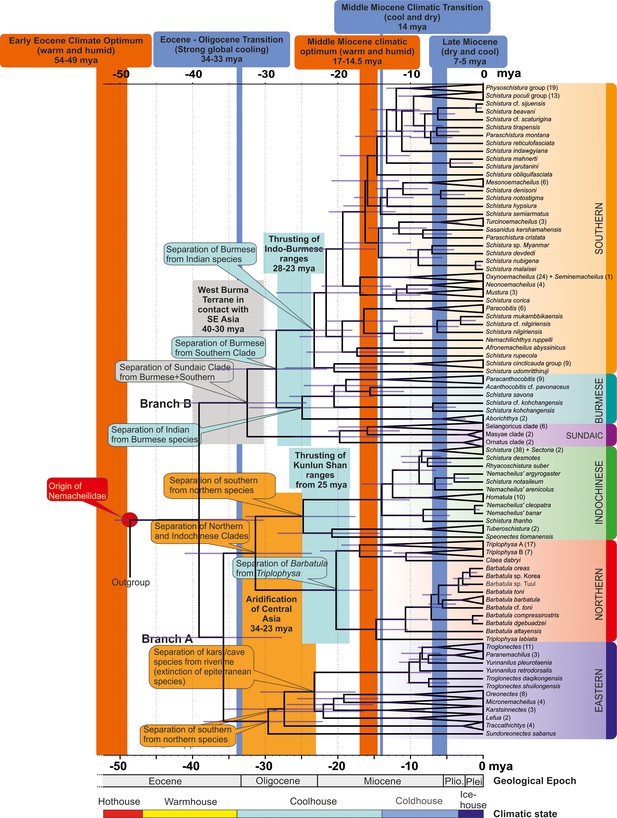
Time tree of the evolutionary history of Nemacheilidae: the figure presents the evolutionary history of Nemacheildae, highlighting the dating of key geological or climatic key paleo-events that might have significantly impacted their evolution.
Orange bars represent exceptionally warm periods, blue bars denote cold periods. Boxes indicate specific geological or climatic event. For detailed explanations of these events and their impact on nemacheilid evolution, refer to text. Climatic states from Westerhold et al., 2020.
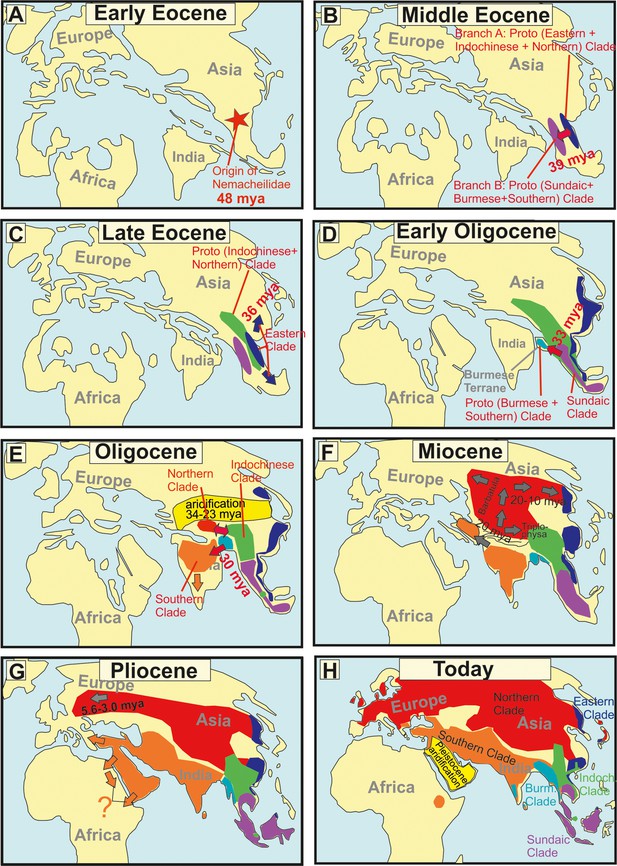
Paleomaps visualising the geographical and chronological aspects of the main events during the evolutionary history of the freshwater fish family Nemacheilidae across Eurasia.
(A) Early Eocene, (B) Middle Eocene, (C) Late Eocene, (F) Miocene, (G) Pliocene, (H) Today A red star marks the point of origin, arrows indicate dispersal events. Background maps were drawn according to information given in Blakey, 2023, Atlas, 2023, Wang, 2004, Westerweel et al., 2020.
Tables
Overview of the major evolutionary events in the six main clades within Nemacheilidae from its origin 48 mya until today.
| Period | Eastern | Indochinese | Northern | Sundaic | Burmese | Southern |
|---|---|---|---|---|---|---|
| Early Eocene 48 mya | Origin of Nemacheilidae in the area of present-day Indochina; probably related to a strong diversification period during the Early Eocene Climatic Optimum (54–49 mya). | |||||
| Middle Eocene 39 mya | Split of Nemacheilidae into two branches (Proto Eastern + Indochinese + Northern branch and Proto Sundaic +Burmese + Southern branch). The ultimate cause for this split is unknown. | |||||
| Late Eocene 36 mya | Formation of Eastern Clade; spreading along the eastern margin of Asia. | The Proto Indochinese + Northern Clade starts expanding northwards and southwards (at least to present-day southern Malay Peninsula). | ||||
| Early Oligocene 33 mya | Formation of the Sundaic Clade by separation of the Proto Burmese +Southern Clade. | The Proto Burmese + Southern Clade forms by colonising the Burmese Terrane, a tectonic microplate that travels northwards with the Indian plate. | ||||
| Oligocene 34–23 mya | The distribution of the Eastern Clade becomes fragmented by strong aridification of Central Asia. | Proto Indochinese + Northern Clade in two refuges during strong aridification of Central Asia. The Proto- Northern Clade endures the period on the northern part of the young Tibetan Plateau, the Proto-Indochinese Clade east of the Tibetan Plateau. After this long time of separation they form the Northern and Indochinese Clades. | The Proto Burmese +Southern Clade colonises the Indian Plate (30 mya). Burmese Terrane and India collide with Asian mainland, causing the rise of the Indo-Burmese mountain ridge (from 28 mya), separating the Southern Clade from the Burmese Clade. | |||
| Early and Middle Miocene 23–10 mya | The genus Barbatula spreads northwards as the Kunlun, Pamir and Altai mountains form (from 25 mya), then eastwards to Korean Peninsula; the genus Triplophysa spreads eastwards into southern China. | The Southern Clade colonises the freshly forming Near East in several waves. The Proto-Afronemacheilus separates from Indian species. | ||||
| Miocene 23–5.3 mya | Period of strong diversification, especially around the time of the Middle Miocene Climatic optimum (17–14.5 mya), Number of lineages in Nemacheilidae rises from 16 to 177. | |||||
| Pliocene 5–2.5 mya | Barbatula spreads westwards due to ice formation in northern Siberia and colonises Europe. In the east it colonises the Japanese Archipelago. | |||||
| Pleistocene from 2.6 mya | Extinctions from Arab Peninsula and the lower Nile valley due to strong aridification. | |||||
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (Nemacheilidae) | DNA sequences | GenBank | 104 different species from the fish family Nemacheilidae (details in Supplementary file 1) | |
| Biological sample (Nemacheilidae) | Tissue samples | This paper | 175 different species from the fish family Nemacheilidae (details in Supplementary file 1) | |
| Sequence-based reagent | PCR primers | See Supplementary file 2 | ||
| Commercial assay or kit | DNeasy Blood and Tissue kit | QIAGEN | DNA isolation kit | |
| Software, algorithm | DNA Star | LASERGENE | ||
| Software, algorithm | BioEdit | Hall, 1999 | ||
| Software, algorithm | PhyloSuite v1.2.2 | Zhang et al., 2020 | ||
| Software, algorithm | MrBayes 3.2.7a | Ronquist and Huelsenbeck, 2003 | ||
| Software, algorithm | ASTRAL III | Zhang et al., 2018 | ||
| Software, algorithm | BEAST 2.6.4 | Bouckaert et al., 2014 |
Additional files
-
Supplementary file 1
List of analysed samples, their identification, geographical origin, voucher number, and GenBank accession numbers for their sequences.
Voucher numbers starting with ‘A’ refer to the collection of IAPG, Liběchov, Czech Republic; ‘CMK’ numbers refer to the collection of Maurice Kottelat; ‘GenBank’ refers to sequences from GenBank; ‘ZRC’ to samples housed in the Lee Kong Chian Natural History Museum, National University of Singapore, Singapore.
- https://cdn.elifesciences.org/articles/101080/elife-101080-supp1-v1.docx
-
Supplementary file 2
List of primers used in the present study for amplification and/or sequencing.
- https://cdn.elifesciences.org/articles/101080/elife-101080-supp2-v1.docx
-
Supplementary file 3
Alignment attributes and best-fit models.
Lengths of alignments, numbers of variable (VP), and parsimony informative (PI) positions and models estimated for all partitions. BEAST and MrBayes models were calculated in Partition Finder 2 (PF2, Lanfear et al., 2016) implemented in PhyloSuite v1.2.2 (Zhang et al., 2020) under AICc criterion, with greedy algorithm (Lanfear et al., 2016) and branch lengths linked. For ML trees, the models and partitioning schemes were estimated under BIC with ModelFinder (Kalyaanamoorthy et al., 2017) implemented in IQ tree. The values and models were calculated for both (A) full as well as (B) reduced dataset. Table (C) provides an overview of data attributes for the ingroup dataset only.
- https://cdn.elifesciences.org/articles/101080/elife-101080-supp3-v1.docx
-
Supplementary file 4
Comparison of biogeographic models in RASP.
The last column shows the p-values of the likelihood ratio test. Based on AICc weight (AICc wt), the DEC+J model is recommended as the best fit for our dataset, supported by low p-values indicating the significant influence of the J factor on model likelihood.
- https://cdn.elifesciences.org/articles/101080/elife-101080-supp4-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/101080/elife-101080-mdarchecklist1-v1.docx





