CXXC-finger protein 1 associates with FOXP3 to stabilize homeostasis and suppressive functions of regulatory T cells
Figures
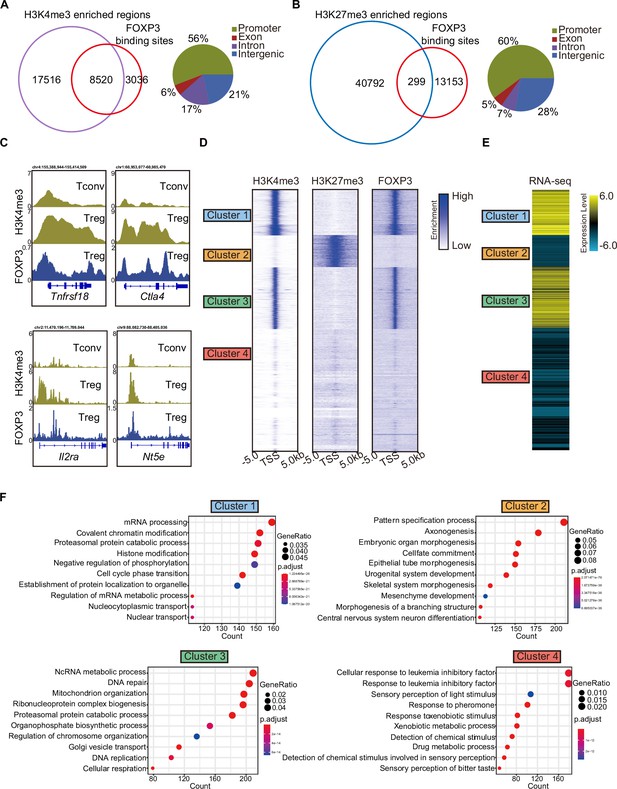
H3K4me3 is required for FOXP3-dependent gene activation in Treg cells.
(A) Venn diagram showing overlap of H3K4me3-enriched regions (this study) and FOXP3-binding sites Konopacki et al., 2019 in sorted CD4+YFP+Treg cells (left). Genomic distribution of overlapped peaks (right). Note that the overlapped peaks are predominantly enriched at promoters. (B) Venn diagram showing overlap of H3K27me3-enriched regions (Wei et al., 2009) and FOXP3-binding sites in Treg cells (left). Genomic distribution of overlapped peaks (right). Note that the overlapped peaks are predominantly enriched at promoters. (C) Representative genome browser view showing the enrichments of H3K4me3 and FOXP3 in Tconv or Treg cells. (D) Heatmap showing enrichment of H3K27me3, H3K4me3, and FOXP3 surrounding the transcription start site (TSS). Unsupervised k-means clustering was conducted on H3K27me3 and H3K4me3 signals. (E) Heatmap showing gene expression levels in Treg cells (RNA-sequencing [RNA-seq] data was obtained from Oh et al., 2017). The clusters were consistent as in C. (F) Gene Ontology (GO) pathway analysis of different clusters.
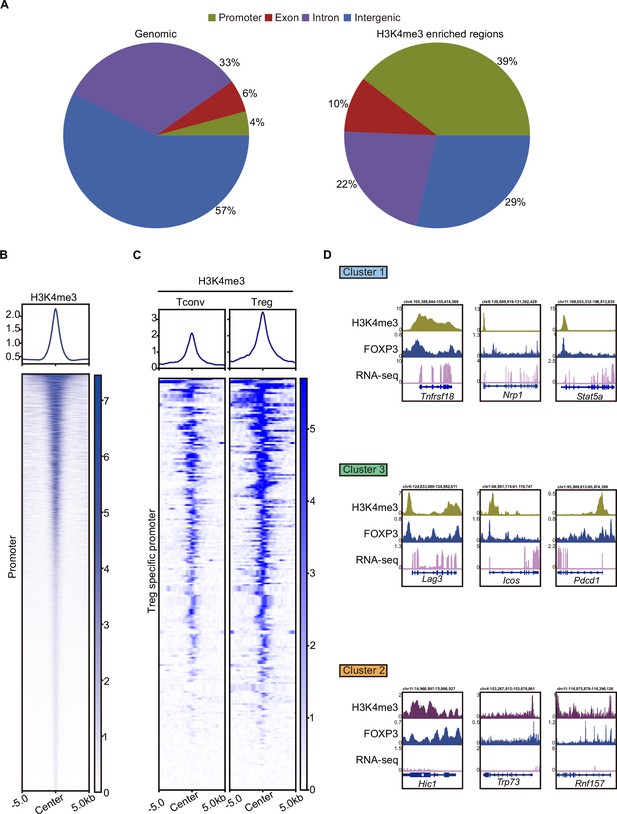
H3K4me3 is required for FOXP3-dependent gene activation in Treg cells.
(A) Genomic distribution of random and H3k4me3-enriched peaks in Treg cells. (B) Heatmap showing enrichment of H3K4me3 surrounding the transcription start site (TSS). The top panel of the profile plots shows the average CUT&Tag signals. (C) Heatmap showing H3K4me3 enrichment at Treg-specific gene promoter in Tconv and Treg cells. The top panel of the profile plots shows the average CUT&Tag signals. (D) Representative Genome Browser view showing RNA-sequencing (RNA-seq), FOXP3 binding, and H3K4me3/H3K27me3 enrichment in Treg cells.
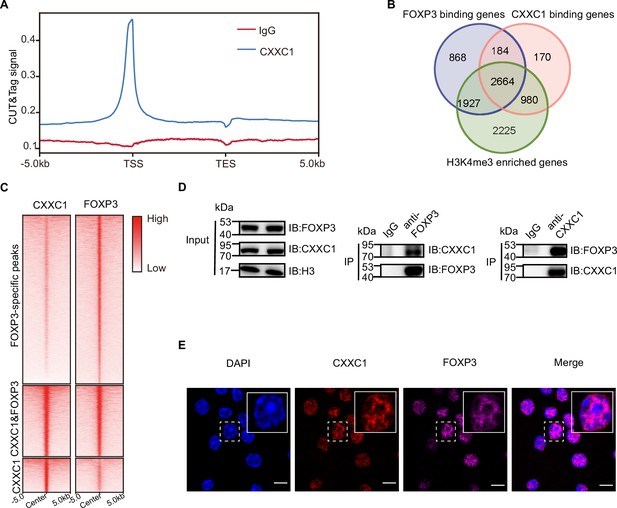
CXXC1 interacts with FOXP3 in Treg cell.
(A) Average CXXC1 CUT&Tag signals around genes in Treg cells. IgG was used as the control. (B) Venn diagrams showing the overlap of FOXP3-binding genes, CXXC1-binding genes, and H3K4me3-enriched genes in Treg cells. Genes covered by FOXP3-binding sites, CXXC1-binding sites, or exhibited high H3K4me3 levels at promoters were defined as FOXP3-bound genes, CXXC1-bound genes, or H3K4me3-enriched genes. (C) Heatmaps showing FOXP3 ChIP-seq and CXXC1 CUT&Tag signals at indicated regions. (D) Interaction between FOXP3 and CXXC1 was assessed by co-IP (forward and reverse) using Treg cell lysates. (E) Immunofluorescence for FOXP3 and CXXC1 colocalization in Treg cells. Scale bars, 2 μm.
-
Figure 2—source data 1
File containing labeled original western blots for Figure 2.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig2-data1-v1.zip
-
Figure 2—source data 2
Original gel image files for western blot analysis displayed in Figure 2.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig2-data2-v1.zip
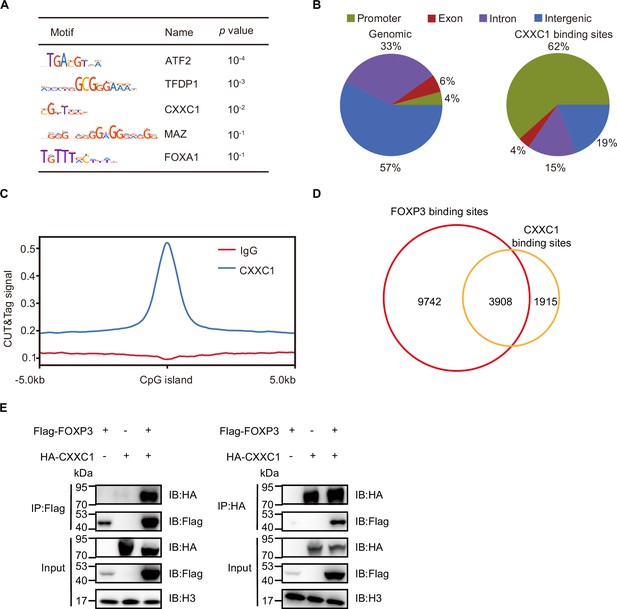
FOXP3 interacts with CXXC1.
(A) Motif enrichment analysis of overlapped peaks in H3K4me3-enriched regions and FOXP3-binding sites. (B) Genomic distribution of control and CXXC1-binding sites in Treg cells. (C) Average CXXC1 CUT&Tag signals around CpG island in Treg cells. IgG was used as the control. (D) Venn diagrams showing the overlap of FOXP3- and CXXC1-binding sites in Treg cells. (E) Physical interaction between CXXC1 and FOXP3 in HEK 293T cells transiently transfected with expression constructs. IP, immunoprecipitation.
-
Figure 2—figure supplement 1—source data 1
File containing labeled original western blots for Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig2-figsupp1-data1-v1.zip
-
Figure 2—figure supplement 1—source data 2
Original gel image files for western blot analysis displayed in Figure 2—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig2-figsupp1-data2-v1.zip
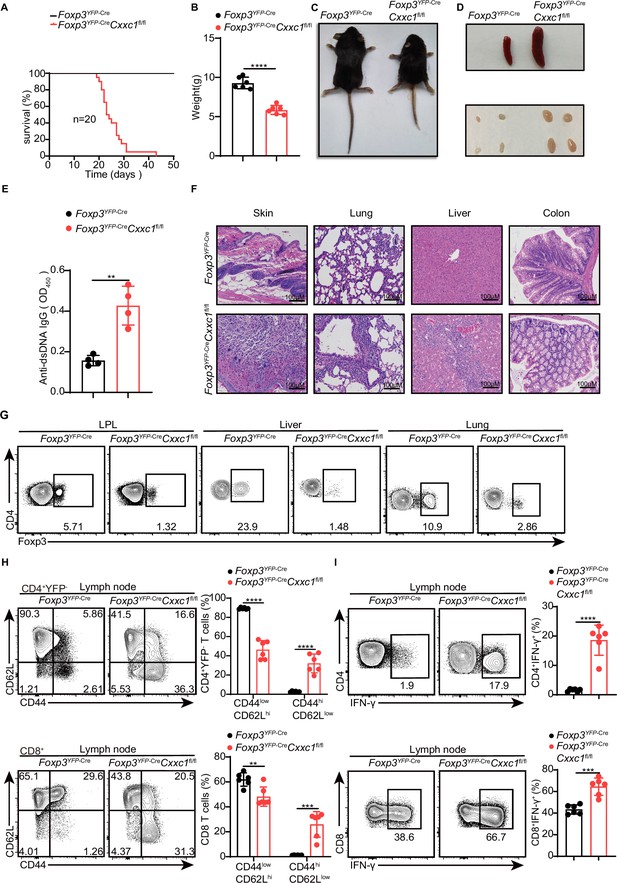
Foxp3YFP-CreCxxc1fl/fl mice spontaneously develop a fatal early-onset inflammatory disorder.
(A) Survival curves of Foxp3YFP-Cre (black line) and Foxp3YFP-Cre Cxxc1fl/fl (red line) mice (n = 20). (B) Gross body weight of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). (C) A representative image of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (D) Representative images showing the spleen and peripheral lymph nodes from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (E) ELISA quantification of anti-dsDNA IgG in the serum of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 4). (F) Hematoxylin and eosin staining of the skin, lung, liver, and colon from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (scale bar, 100 μm). (G) Representative flow cytometry plots of CD4+ Foxp3+ Treg cells isolated from the small intestinal lamina propria (LPL), liver, and lung of Foxp3YFP-Cre and Foxp3YFP-Cre Cxxc1fl/fl mice. (H) Flow cytometry analysis of CD62L and CD44 expression on peripheral lymph node CD4+YFP− and CD8+T cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (left). Right, frequency of CD44lowCD62Lhi and CD44hiCD62Llow population in CD4+YFP− or CD8+ T cells (n = 6). (I) Lymph node cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice were stimulated ex vivo with PMA + ionomycin for 4 hr and analyzed for IFN-γ expressing in CD4+ YFP− or CD8+ T cells using flow cytometry (left). Right, percentages of IFN-γ+CD4+ YFP− or IFN-γ+CD8+ T cells in the lymph nodes of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). All mice analyzed were 18–20 days old unless otherwise specified. Error bars show mean ± SD. The log-rank survival curve was used for survival analysis in A, and unpaired t-test or multiple unpaired t-test were used for statistical analyses in B, E, G– I (**p < 0.01, ***p < 0.001, ****p < 0.0001). The flow cytometry results are representative of three independent experiments.
-
Figure 3—source data 1
Original source data for graphs displayed in Figure 3.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig3-data1-v1.xlsx
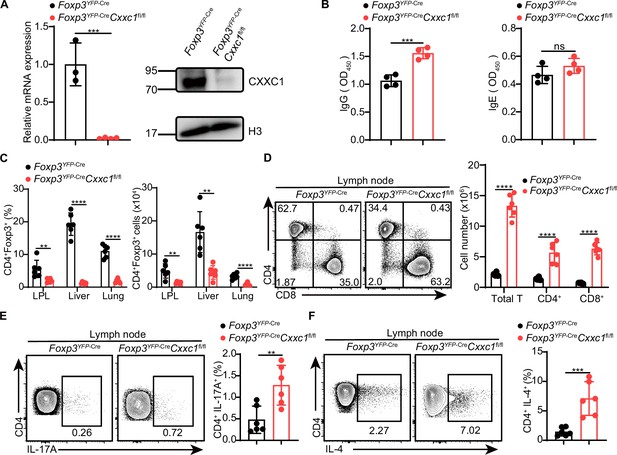
Disrupted immune homeostasis in Foxp3YFP-CreCxxc1fl/fl mice.
(A) Analysis of Cxxc1 mRNA (left; Foxp3YFP-Cre, n = 3, Foxp3YFP-CreCxxc1fl/fl, n = 4) and protein (right) expression in CD4+YFP+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (B) Quantification of serum IgG and IgE from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 4). (C) Quantification of the frequencies and absolute numbers of CD4+ Foxp3+ Treg cells in the small intestinal lamina propria (LPL), liver, and lung of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). (D) Flow cytometry analysis of CD4 and CD8 expression in peripheral lymph nodes from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. Absolute cell counts of Total T, CD4+, and CD8+ T cell subsets are shown (n = 6). (E, F) Lymph node cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice were stimulated ex vivo with PMA + ionomycin for 4 hr and analyzed for IL-17A+ and IL-4+ CD4+YFP− T cells by flow cytometry. Percentages of IL-17A+ and IL-4+ CD4+ T cells in lymph nodes from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). Error bars show mean ± SD. p values are determined by an unpaired t-test or multiple unpaired t-test (A–F) (**p < 0.01, ***p < 0.001, ****p < 0.0001). The flow cytometry and western blot results represent at least three independent experiments.
-
Figure 3—figure supplement 1—source data 1
File containing labeled original western blots for Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig3-figsupp1-data1-v1.zip
-
Figure 3—figure supplement 1—source data 2
Original gel image files for western blot analysis displayed in Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig3-figsupp1-data2-v1.zip
-
Figure 3—figure supplement 1—source data 3
Original source data for graphs displayed in Figure 3—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig3-figsupp1-data3-v1.xlsx
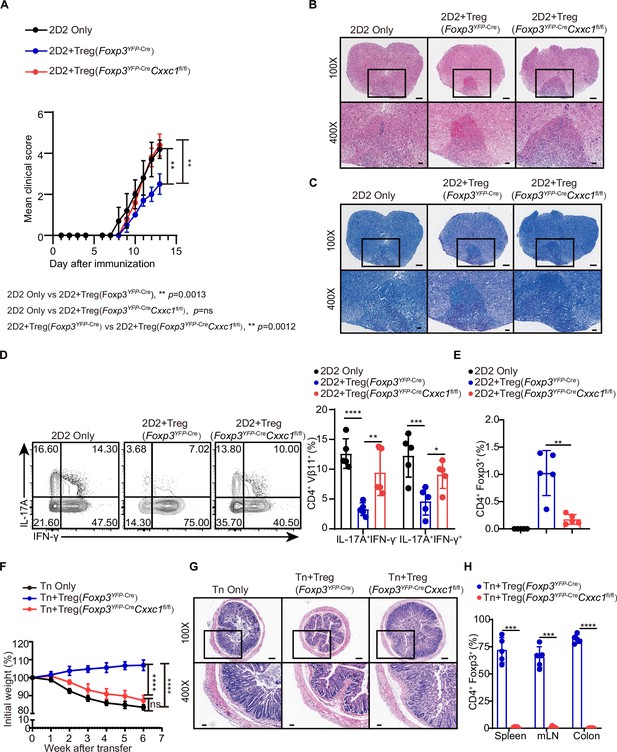
CXXC1 is essential for Treg cells to suppress T cell-mediated experimental autoimmune encephalomyelitis (EAE) and colitis.
(A) Mean clinical scores for EAE in Rag1−/− recipients of 2D2 CD4+ T cells, either alone or in combination with Foxp3YFP-Cre or Foxp3YFP-CreCxxc1fl/fl mice after immunization with MOG35–55, complete Freund’s adjuvant (CFA), and pertussis toxin (n = 5). (B, C) Representative histology of the spinal cord of Rag1−/− mice after EAE induction. Hematoxylin and eosin (H&E) staining (upper), Luxol fast blue (F&B) staining (lower). Scale bars, 50 μm (×400) and 200 μm (×100). (D) Representative flow cytometry plots and quantification of the percentages of IFNγ+ or IL-17A+ CD4+Vβ11+ T cells (n = 5). (E) Statistical analysis of the percentage CD4+ FOXP3+ Treg cell in central nervous system (CNS) tissues 14 days after EAE induction (n = 5). (F) Changes in body weight of Rag1−/− mice after colitis induction (n = 6). (G) H&E staining of colons from T cell-induced colitis mice 6 weeks after T cell transfer. Scale bars, 50 μm (×400) and 200 μm (×100). (H) Statistical analysis of the percentage CD4+ FOXP3+ Treg cell in the spleen, mesenteric lymph nodes, and colon 6 weeks after colitis induction (n = 5). Error bars show mean ± SD. p values are determined by a unpaired t-test or two-way ANOVA and Holm–Sidak post hoc test (A, D, E, F, H) (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
-
Figure 4—source data 1
Original source data for graphs displayed in Figure 4.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig4-data1-v1.xlsx
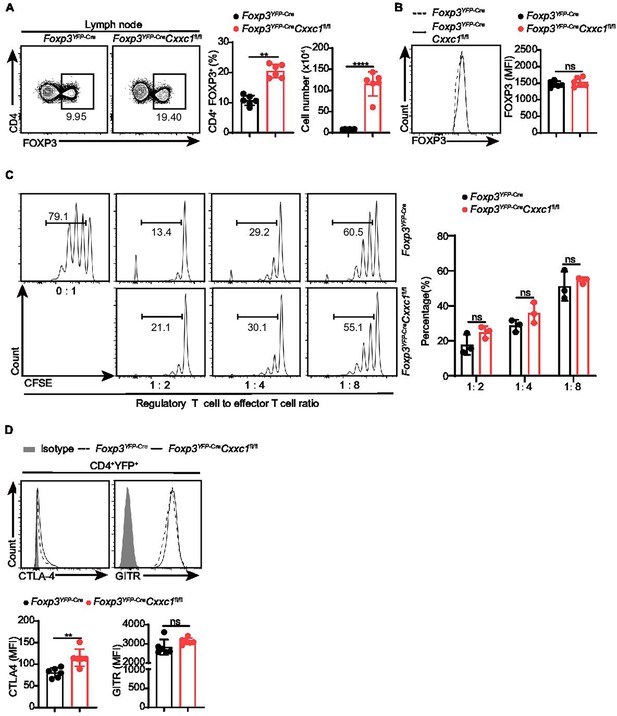
The number of Treg cells deficient in CXXC1 did not decrease.
(A) Flow cytometry analysis of CD4+FOXP3+ Treg cells in lymph nodes of Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. On the right, the proportion and number of Treg cells are shown (n = 6). (B) Representative figure showing FOXP3 protein expression in CD4+ T cells from the lymph nodes (n = 6). (C) Suppression of CFSE-labeled Tn cell proliferation by different ratios of CD4+YFP+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. On the right, the percentage of proliferated responding T cells is presented (n = 3). (D) Expression of Treg signature molecules in Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). Error bars show mean ± SD. p values are determined by an unpaired t-test or multiple unpaired t-tests (A–D) (ns, not significant. **p < 0.01, ****p < 0.0001). The flow cytometry results are representative of three independent experiments.
-
Figure 4—figure supplement 1—source data 1
Original source data for graphs displayed in Figure 4—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig4-figsupp1-data1-v1.xlsx
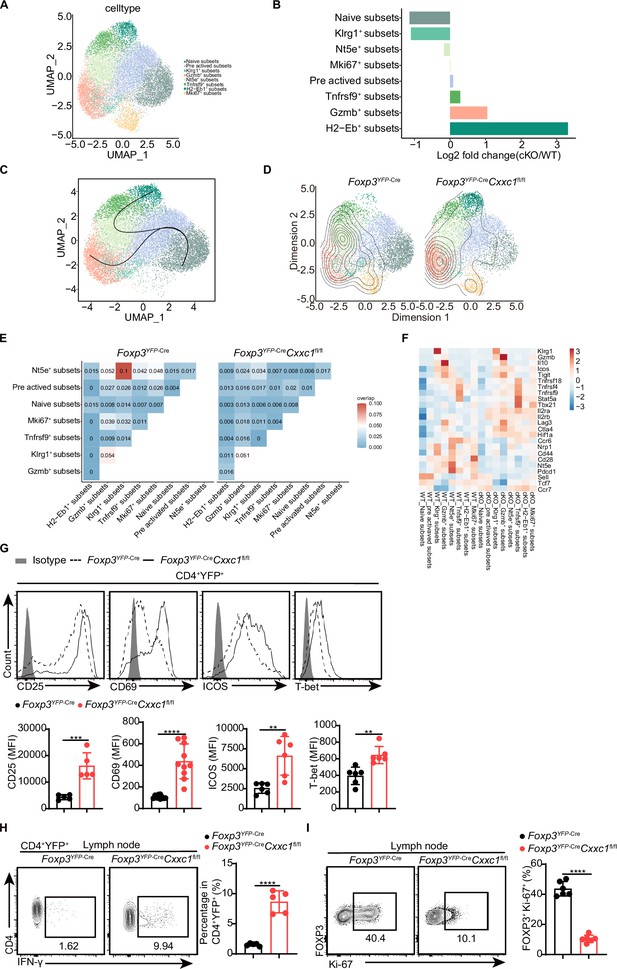
Single-cell transcriptomics reveals distinct Treg cell populations.
(A) Uniform manifold approximation and projection (UMAP) plot showing clusters identified based on variable gene expression of sorted CD4+YFP+ Treg cells. Each dot represents a cell, and each color corresponds to a different population of cell types. Clustering analysis revealed eight distinct Treg cell populations. (B) Mean fold changes in cluster abundance between Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (C) Pseudotime trajectories of Treg cells based on Slingshot, color-coded by Treg cell subpopulations. (D) Visualization of density and clonotype richness across Treg clusters from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (E) T cell receptor (TCR) sharing of expanded clonotypes across all possible combinations of Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (F) Heatmap showing Z scores for the average expression of Treg-specific genes in each cluster between Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. Representative flow cytometry plots and quantification of (G) expression of CD25, CD69, ICOS, T-bet, and (H) IFN-γ in CD4+YFP+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 5 CD25, n = 10 CD69, n = 5 ICOS, n = 6 T-bet, n = 5 IFN-γ). (I) Ki-67 expression (left) and frequency (right) in CD4+FOXP3+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6). Error bars show mean ± SD. p values are determined by a unpaired t-test (G–I) (**p < 0.01, ***p < 0.001, ****p < 0.0001). The flow cytometry results are representative of three independent experiments.
-
Figure 5—source data 1
Original source data for graphs displayed in Figure 5.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig5-data1-v1.xlsx
-
Figure 5—source data 2
Single-cell T cell receptor (TCR) V(D)J repertoire profiling of CD4+YFP + Treg cells in Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig5-data2-v1.csv
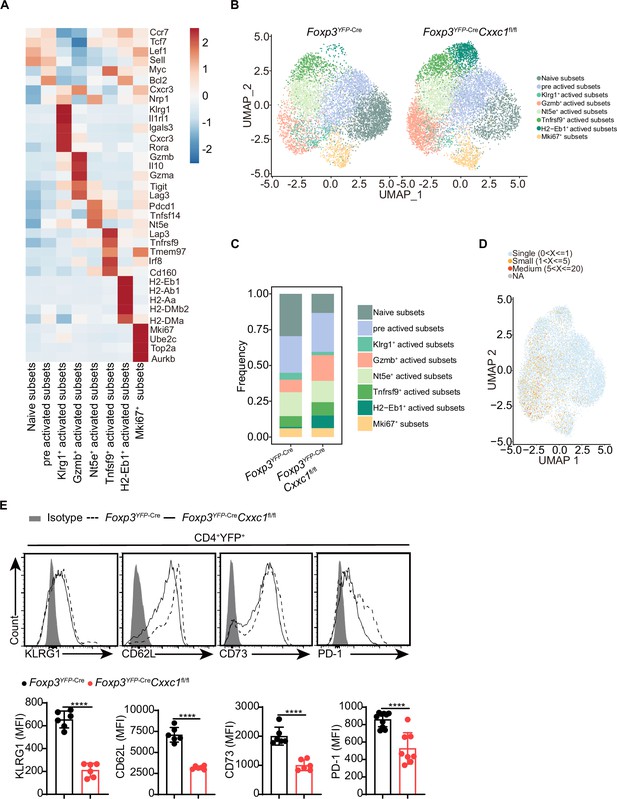
Single-cell transcriptomics reveals distinct Treg cell populations.
(A) Heatmap displaying the Z scores for the average expression of Treg-specific genes across each cluster. (B) Uniform manifold approximation and projection (UMAP) plot showing clusters identified by variable gene expression in sorted CD4+YFP+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice. (C) Relative proportions of Treg cell subpopulations in Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl, revealing heterogeneity. (D) Density and clonotype richness across Treg cell clusters, with colors indicating clone size. (E) Representative flow plots and quantified expression of KLRG1, CD62L, CD73, and PD-1 in CD4+YFP+ Treg cells from Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl mice (n = 6 KLRG1, n = 6 CD62L, n = 6 CD73, n = 8 PD-1). Error bars show mean ± SD. p values are determined by an unpaired t-test (E) (****p < 0.0001).
-
Figure 5—figure supplement 1—source data 1
Original source data for graphs displayed in Figure 5—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig5-figsupp1-data1-v1.xlsx
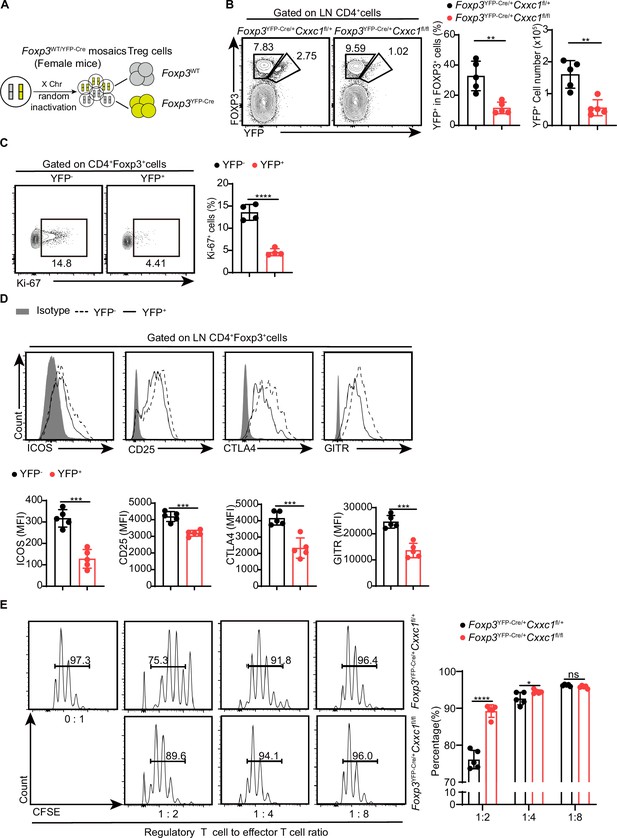
Cxxc1-deficient Treg cells exhibit functional impairment and disrupted homeostasis in steady-state conditions.
(A) Schematic representation of wild-type and Cre-positive Treg cells in female Foxp3YFP-Cre/+ mice.(B) Flow cytometry analysis of the YFP-FOXP3+ (WT) and YFP+FOXP3+ (KO) Treg cells in Foxp3YFP-Cre/+ Cxxc1fl/+ (het-WT) and Foxp3YFP-Cre /+ Cxxc1fl/fl (het-KO) female mice (left), along with the frequency and absolute numbers of YFP+ cells within the total Treg population (right) (n = 5). (C) Flow cytometry analysis of Ki-67expression (left) and MFI (right) in YFP− and YFP+ cells within the CD4+FOXP3+ Treg cells from 6- to 8-week-old het-KO female mice (n = 4). (D) Representative flow cytometry plots and quantification of ICOS, CD25, CTLA4, and GITR expression in YFP− and YFP+ cells within CD4+FOXP3+ Treg cells from het-KO female mice (n = 5). (E) Suppression of CFSE-labeled Tn cell proliferation by different ratios of CD4+YFP+ Treg cells from Foxp3YFP-Cre/+ Cxxc1fl/+ and Foxp3YFP-Cre/+ Cxxc1fl/fl female mice. On the right, the percentage of proliferated responding T cells is presented (n = 5). Error bars show mean ± SD. p values are determined by a unpaired t-test or multiple unpaired t-test (B–E) (ns, not significant. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001). The flow cytometry results are representative of three independent experiments.
-
Figure 6—source data 1
Original source data for graphs displayed in Figure 6.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig6-data1-v1.xlsx
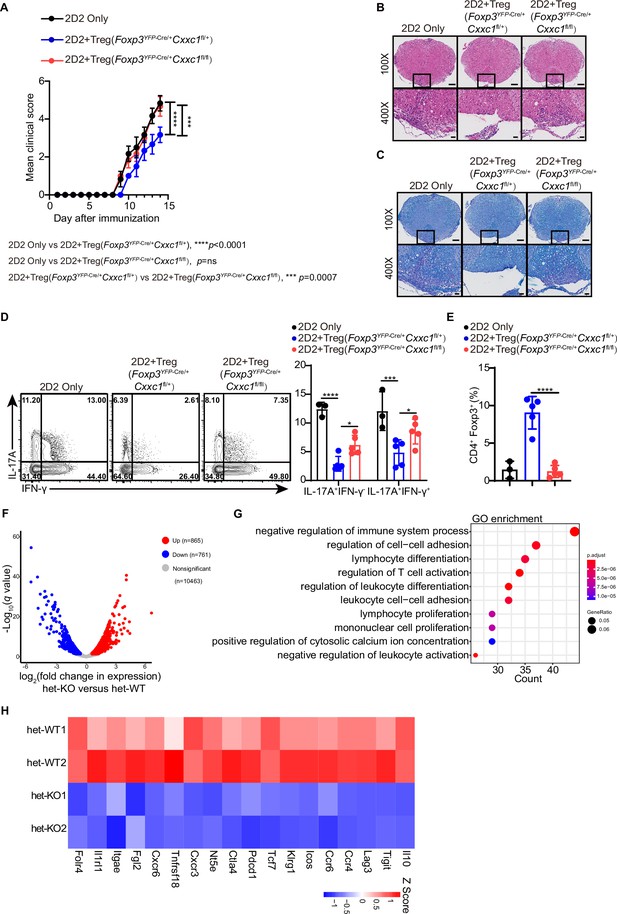
Cxxc1 deficiency impairs Treg cell suppressive function and aggravates experimental autoimmune encephalomyelitis (EAE) severity.
(A) Mean clinical scores for EAE in Rag1−/− recipients of 2D2 CD4+ T cells, either alone or in combination with Foxp3YFP-Cre/+ Cxxc1fl/+ or Foxp3YFP-Cre/+ Cxxc1fl/fl female mice after immunization with MOG35–55, complete Freund’s adjuvant (CFA), and pertussis toxin (n = 6). (B, C) Representative histology of the spinal cord of Rag1−/− mice after EAE induction. Hematoxylin and eosin (H&E) staining (upper), Luxol fast blue (F&B) staining (lower). Scale bars, 50 μm (×400) and 200 μm (×100). (D) Representative flow cytometry plots and quantification of the percentages of IFNγ+ or IL-17A+ CD4+Vβ11+ T cells. (E) Statistical analysis of the percentage CD4+ FOXP3+ Treg cell in central nervous system (CNS) tissues 14 days after EAE induction (2D2 only n = 3, 2D2+het-WT Treg n = 5, 2D2+het-KO Treg n = 5). (F) Volcano plot showing the expression of genes in CD4+YFP+ Treg cells from het-KO versus het-WT. The significantly up- and downregulated genes were identified as genes with p-value <0.05, fold change greater than 1.5 and less than 0.66, respectively. (G) Gene Ontology (GO) pathway analysis of downregulated genes in het-KO CD4+YFP+ Treg cells. (H) Heatmap showing the expression levels of Treg cell signature genes implicated in suppressive function. Error bars show mean ± SD. p values are determined by an unpaired t-test or two-way ANOVA and Holm–Sidak post hoc test (A, D, E) (*p < 0.05, ***p < 0.001, ****p < 0.0001).
-
Figure 6—figure supplement 1—source data 1
Original source data for graphs displayed in Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig6-figsupp1-data1-v1.xlsx
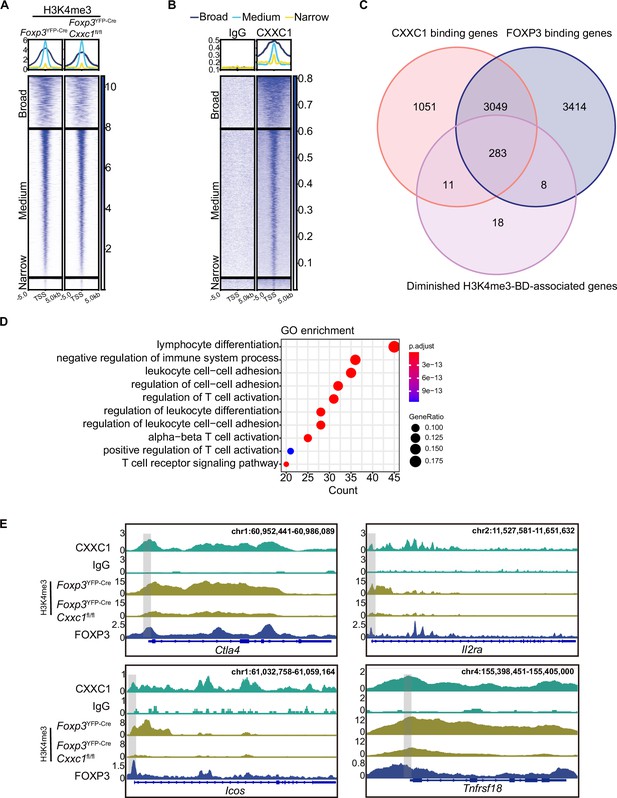
CXXC1 regulates FOXP3-dependent molecule H3K4 trimethylation in Treg cells.
Heatmaps showing H3K4me3 (A) and CXXC1 (B) signals centered on narrow, medium, and broad domains. The top panel shows the average CUT&Tag signals around indicated domains. (C) Venn diagrams showing the overlap of FOXP3-binding genes, CXXC1-binding genes, and H3K4me3-BD-associated genes with decreased H3K4me3 levels after Cxxc1 depletion in Treg cells. (D) Gene Ontology (GO) pathway analysis of the overlapped genes in C. (E) Representative genome browser view showing the enrichments of FOXP3, CXXC1, and H3K4me3 in Treg cells.
-
Figure 7—source data 1
Table: list of overlapped genes in Figure 7C.
- https://cdn.elifesciences.org/articles/103417/elife-103417-fig7-data1-v1.txt
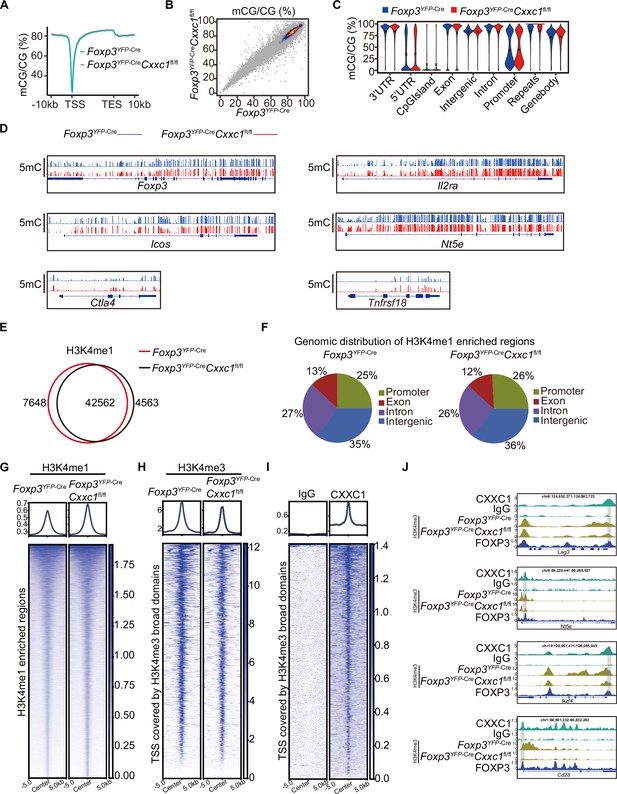
The epigenetic program of Treg cells.
(A) Average CpG methylation in CD4+YFP+ cells from 3-week-old Foxp3YFP-Cre and Foxp3YFP-CreCxxc1fl/fl surrounding the transcription start site (TSS) and transcriptional terminal site (TES). (B) Scatter plot showing methylation levels at 5-kb bins in wild-type (WT) and Cxxc1-deficient Treg cells. (C) A violin plot showing the methylation levels at different genomic regions in WT and Cxxc1-deficient Treg cells. (D) Genome browser view showing DNA methylation levels in WT and Cxxc1-deficient Treg cells. (E) Venn diagrams showing the overlap of H3K4me1 peaks in WT and Cxxc1-null Treg cells. (F) Genomic distribution of H3K4me1 peaks in WT and Cxxc1-null Treg cells. (G) Heatmap showing H3K4me1 enrichment in WT and Cxxc1-null Treg cells. The top panel of the profile plots shows the average CUT&Tag signals. Heatmaps showing H3K4me3 (H) and CXXC1 (I) signals centered on TSS covered by H3K4me3 broad domains. The top panel shows the average CUT&Tag signals around indicated domains. (J) Representative genome browser view showing the enrichments of FOXP3, CXXC1, and H3K4me3 in Treg cells.
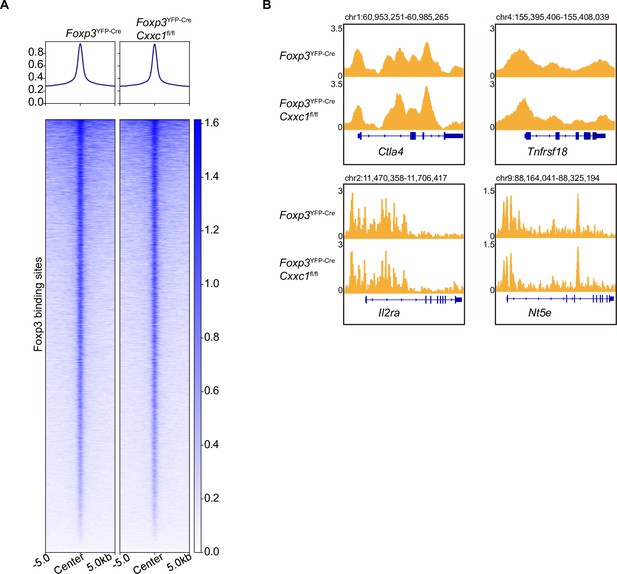
FOXP3-binding profile and enrichment in WT and Cxxc1-Null Treg cells.
(A) Heatmap showing FOXP3 CUT&Tag signals in WT and Cxxc1-null Treg cells. The top panel of the profile plots shows the average CUT&Tag signals. (B) Representative genome browser views showing enrichment of FOXP3 in WT and Cxxc1-null Treg cells.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mus musculus) | Foxp3YFP-Cre mice | Gifted from Prof. Bin Li | Shanghai Jiao Tong University | N/A |
| Strain, strain background (Mus musculus) | Cxxc1fl/fl mice | The Shanghai Research Center for Model Organisms | N/A | N/A |
| Strain, strain background (Mus musculus) | C57BL/6JGpt-Rag1em1Cd3259/Gpt, Rag1−/− mice | GemPharmatech | Cat# T004753; RRID:IMSR_GPT:T004753 | N/A |
| Strain, strain background (Mus musculus) | 2D2 mice | Gifted from Prof. Linrong Lu | Zhejiang University | N/A |
| Biological sample (Mus musculus) | Thymus, lymph node, lung, liver, small intestine lamina propria lymphocytes | This paper | N/A | Freshly isolated tissue |
| Cell line (Mus musculus, mouse) | HEK 293T | ATCC | Cat# ACS-4500; RRID:CVCL_4V93 | N/A |
| Cell line (Mus musculus, mouse) | Plat E | Gifted from Prof. Xiaolong Liu | Shanghai Institutes for Biological Sciences | N/A |
| Antibody | anti-mouse CD16/32 (rat monoclonal) | BioLegend | Cat# 101320; RRID:AB_1574975 | (1:200) |
| Antibody | PE/Cyanine7 anti-TCR-β (hamster monoclonal) | BioLegend | Cat# 109222; RRID:AB_893625 | (1:400) |
| Antibody | PE/Cyanine7 anti-KLRG1(hamster monoclonal) | BioLegend | Cat# 138416; RRID:AB_2561736 | (1:100) |
| Antibody | APC-eFluo 780 anti-CD4 (rat monoclonal) | Invitrogen | Cat# 47-0042-82; RRID:AB_1272183 | (1:400) |
| Antibody | PE anti-CD152 (CTLA4) (hamster monoclonal) | BioLegend | Cat# 106306; RRID:AB_313255 | (1:100) |
| Antibody | PE anti-CD278 (ICOS) (hamster monoclonal) | BioLegend | Cat# 107706; RRID:AB_313335 | (1:100) |
| Antibody | PE anti-CD357 (GITR) (rat monoclonal) | BioLegend | Cat# 126310; RRID:AB_1089132 | (1:100) |
| Antibody | PE anti-CD69 (hamster monoclonal) | Invitrogen | Cat# 12-0691-83; RRID:AB_ 465733 | (1:400) |
| Antibody | PE anti-CD25 (rat monoclonal) | BioLegend | Cat# 102008; RRID:AB_312856 | (1:400) |
| Antibody | Brilliant Violet 650 anti-mouse CD8a (rat monoclonal) | BioLegend | Cat# 100742; RRID:AB_2563056 | (1:400) |
| Antibody | APC anti-TCR V beta 11 (rat monoclonal) | Invitrogen | Cat# 17-5827-82; RRID:AB_2573226 | (1:400) |
| Antibody | APC/Cyanine7 anti-CD44 (rat monoclonal) | BioLegend | Cat# 103028; RRID:AB_830785 | (1:400) |
| Antibody | APC anti-CD62L (rat monoclonal) | BioLegend | Cat# 104412; RRID:AB_313099 | (1:400) |
| Antibody | APC anti-Foxp3 (rat monoclonal) | Invitrogen | Cat# 17-5773-82; RRID:AB_469457 | (1:100) |
| Antibody | Pacific Blue anti-IFN-γ (rat monoclonal) | BioLegend | Cat# 505818; RRID:AB_893526 | (1:100) |
| Antibody | PE anti-IL-4 (rat monoclonal) | Invitrogen | Cat# 12-7041-82; RRID:AB_466156 | (1:100) |
| Antibody | PE-Cyanine7 anti-IL-17A (rat monoclonal) | Invitrogen | Cat# 25-7177-82; RRID:AB_10732356 | (1:100) |
| Antibody | PE anti- T-bet (mouse monoclonal) | Invitrogen | Cat# 12-5825-82; RRID:AB_925761 | (1:100) |
| Antibody | BV421 anti-PD-1 (hamster monoclonal) | BD | Cat# 562584; RRID:AB_2737668 | (1:100) |
| Antibody | APC anti-CD45RB (rat monoclonal) | BioLegend | Cat# 103319; RRID:AB_2565228 | (1:100) |
| Antibody | PE anti-CD73 (rat monoclonal) | BioLegend | Cat# 127205; RRID:AB_ 2154094 | (1:100) |
| Antibody | PerCPCy5.5 anti-Ki-67(mouse monoclonal) | BD | Cat# 561284; RRID:AB_10611574 | (1:200) |
| Antibody | Anti-Cxxc1 (Rabbit monoclonal) | Abcam | Cat# ab198977 RRID:AB_3101764 | WB (1:1000), IF (1:100) |
| Antibody | Anti-Foxp3 (mouse monoclonal) | Invitrogen | Cat# 14-4774-82; RRID:AB_467552 | WB (1:1000) |
| Antibody | Anti-Flag (Rabbit monoclonal) | Cell Signaling | Cat# 14793; RRID:AB_2572291 | WB (1:1000) |
| Antibody | Anti-HA (Rabbit monoclonal) | Cell Signaling | Cat# 3724S RRID:AB_1549585 | WB (1:1000) |
| Antibody | anti-H3K4me3 (Rabbit Polyclonal) | Active Motif | Cat# 39016 RRID:AB_2687512 | CUT&Tag (1:50) |
| Antibody | anti-FOXP3 (Rabbit Polyclonal) | Abcam | Cat# ab150743 | CUT&Tag (1:50) |
| Antibody | Normal Rabbit IgG (Rabbit Polyclonal) | Cell Signaling | Cat# 2729 RRID:AB_1031062 | CUT&Tag (1:50) |
| Antibody | HRP-conjugated anti-mouse IgG | SouthernBiotech | Cat# 1033-05 RRID:AB_2737432 | (1:2000) |
| Antibody | HRP-conjugated anti-mouse IgE | SouthernBiotech | Cat# 1110-05 RRID:AB_2794604 | (1:2000) |
| Antibody | Anti-Mo CD3e (hamster monoclonal) | Invitrogen | Cat# 16-0031-85; RRID:AB_468848 | 2 μg/ml |
| Antibody | Anti-Mo CD28 (hamster monoclonal) | Invitrogen | Cat# 16-0281-85; RRID:AB_468922 | 3 μg/ml |
| Antibody | Anti-mouse IFN-γ (rat monoclonal) | BioLegend | Cat# 505847; RRID:AB_2616675 | 10 μg/ml |
| Antibody | Anti-mouse IL-12 (rat monoclonal) | BioLegend | Cat# 505309; RRID:AB_2783330 | 10 μg/ml |
| Antibody | Anti-mouse IL-4 (rat monoclonal) | BioLegend | Cat# 504135; RRID:AB_2750404 | 10 μg/ml |
| Sequence-based reagent | Cxxc1 genotyping Forward | This paper | Genotyping PCR primer | CGAGAGATGAAGAGGAGCCA |
| Sequence-based reagent | Cxxc1 genotyping Reverse | This paper | Genotyping PCR primer | CACAAAGATAGGCTCCATCC |
| Sequence-based reagent | Foxp3YFP-Cre WT genotyping Forward | This paper | Genotyping PCR primer | CTATGGAAACCGGGCGATGA |
| Sequence-based reagent | Foxp3YFP-Cre WT genotyping Reverse | This paper | Genotyping PCR primer | AGTGGCAAGTGAGACGTGGG |
| Sequence-based reagent | Foxp3YFP-Cre genotyping Forward | This paper | Genotyping PCR primer | AGGATGTGAGGGACTACCTCCTGTA |
| Sequence-based reagent | Foxp3YFP-Cre genotyping Reverse | This paper | Genotyping PCR primer | TCCTTCACTCTGATTCTGGCAATTT |
| Sequence-based reagent | Actb qPCR Forward | This paper | qRT-PCR primer | CTGTCCCTGTATGCCTCTG |
| Sequence-based reagent | Actb qPCR Reverse | This paper | qRT-PCR primer | ATGTCACGCACGATTTCC |
| Sequence-based reagent | Cxxc1 qPCR Forward | This paper | qRT-PCR primer | CTGTGGAGAAGATTTGTGGG |
| Sequence-based reagent | Cxxc1 qPCR Reverse | This paper | qRT-PCR primer | TCTTGTTGTCTAGAGTGGCGATCT |
| Recombinant DNA reagent | pCMV-C-HA plasmid | This paper | N/A | N/A |
| Recombinant DNA reagent | p3×Flag-CMV7.1 plasmid | This paper | N/A | N/A |
| Commercial assay or kit | MojoSort Mouse CD4 T Cell Isolation Kit | BioLegend | Cat# 480005 | N/A |
| Commercial assay or kit | MojoSort Mouse CD4 Naive T Cell Isolation Kit | BioLegend | Cat# 480039 | N/A |
| Commercial assay or kit | RNeasy Plus Mini Kit | QIAGEN | Cat# 74134 | N/A |
| Commercial assay or kit | TruePrep DNA Library Prep Kit V2 for Illumina | Vazyme | Cat# TD501 | N/A |
| Commercial assay or kit | Hyperactive In-Situ ChiP Library Prep Kit for Illumina | Vazyme | Cat# TD901 | N/A |
| Commercial assay or kit | ClonExpress II One Step Cloning Kit | Vazyme | Cat# C112-01 | N/A |
| Commercial assay or kit | E.Z.N.A. Endo-free Plasmid Mini Kit II | Omega | Cat# D6950-02 | N/A |
| Commercial assay or kit | BD Mouse Immune Single-Cell Multiplexing Kit | BD | Cat# 633793 | N/A |
| Commercial assay or kit | BD Mouse Immune Single-Cell Multiplexing Kit | BD | Cat# 633801 | N/A |
| Recombinant protein | IL-2 | Peprotech | Cat# AF-212-12-20ug | 50 U/ml |
| Recombinant protein | TGF-β | Peprotech | Cat# 100-21C-250ug | 5 ng/ml |
| Chemical compound, drug | PMA | Sigma-Aldrich | Cat# P1585 | 50 ng/ml |
| Chemical compound, drug | Ionomycin | Sigma-Aldrich | Cat# I3909 | 1 μg/ml |
| Peptide | MOG35-55 | ChinaPeptides | N/A | 2 mg/ml |
| Software, algorithm | GraphPad Prism v8 | GraphPad | RRID:SCR_002798 | https://www.graphpad.com/ |
| Software, algorithm | FlowJo v10 | TreeStar | RRID:SCR_008520 | https://www.flowjo.com/flowjo/overview |
| Software, algorithm | R version v4.0.2 | R Core | RRID:SCR_001905 | http://www.r-project.org/ |
| Software, algorithm | Adobe Illustrator | Adobe | RRID:SCR_010279 | https://www.adobe.com/products/illustrator.html |
Additional files
-
Supplementary file 1
Quality control metrics for multi-omics datasets: 1a (CUT&Tag), 1b (RNA-seq), and 1c (WGSB).
- https://cdn.elifesciences.org/articles/103417/elife-103417-supp1-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/103417/elife-103417-mdarchecklist1-v1.docx






