BRCA2 BRC missense variants disrupt RAD51-dependent DNA repair
Figures
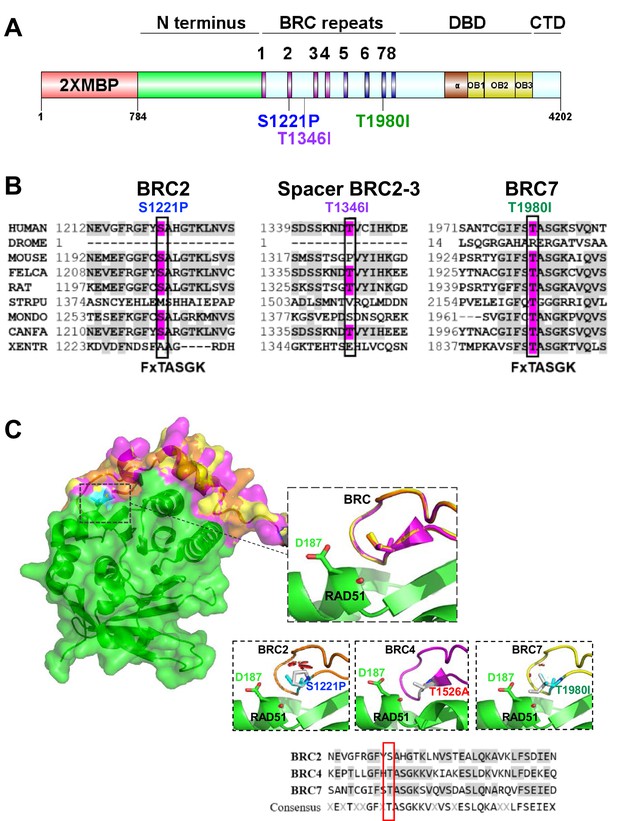
BRCA2 BRC residues S1221, T1346, and T1980 are conserved residues.
S1221P and T1980I are structurally predicted to disrupt BRC folding and RAD51 binding. (A) BRCA2 protein schematic depicting domain organization: 2XMBP tag, N-terminus, BRC repeats, DNA binding domain (DBD), and C-terminal domain (CTD). BRCA2 missense variants used in this study are indicated. (B) Multiple sequence alignment of BRCA2 amino acids flanking each missense variant from different organisms: S1221P (BRC2), T1346I (Spacer BRC2-3), and T1980I (BRC7). Residue corresponding to the missense variant is indicated in pink and conserved residues flanking the missense residue are indicated in light grey. Uniprot and ClustalX (70% threshold for shading) were used for the alignment. FxTASGK is a consensus motif important for RAD51 interactions depicted below BRC2 and BRC7. (C) Structural models based on 1N0W structure. Homology model comparison of BRC2 (orange) and BRC7 (yellow) based on the BRC4 (pink) structure. Top Inset: BRC2 (S1221), BRC4 (T1526), and BRC7 (T1980) overlayed to show conservation. Bottom Insets: predicted disruptions resulting from variants BRC2 (S1221P), BRC4 (T1526A), and BRC7 (T1980I). BRC4 T1526A is a previously characterized missense variant that disrupts RAD51 binding. RAD51 D187 residue is displayed in green. Clashes are indicated in red. Sequence alignment of BRC2, BRC4, and BRC7 repeats. Missense variant residues are boxed in red.
-
Figure 1—source data 1
Contains BRCA2 sequence alignment depicting homology (Figure 1B), SWISS-MODEL homology modeling reports (Figure 1C), close up views of BRC-RAD51 amino acid interactions (Figure 1C).
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig1-data1-v3.zip
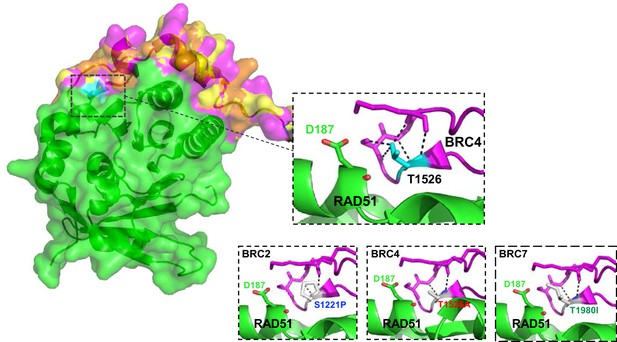
Polar contacts of S1221P, T1526A, and T1980I.
Homology model comparisons of BRC2 and BRC7 based on the BRC4 (1N0W) structure. Top Inset: Polar contacts (black dotted lines) within a BRC4 loop containing residue T1526. Bottom insets: Loss of polar contacts predicted by the S1221P variant within the BRC2 homology model (left), the T1526A variant within BRC4 (middle), and the T1980I variant within the BRC7 homology model (right). RAD51 D187 residue is displayed in green.
-
Figure 1—figure supplement 1—source data 1
T1526 polar contacts.
T1526A polar contacts. T1526I polar contacts. T1526P polar contacts.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig1-figsupp1-data1-v3.zip
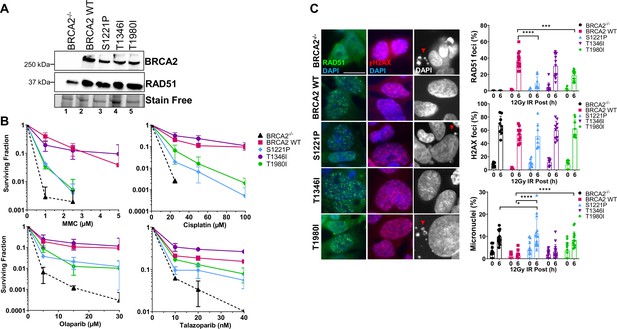
S1221P and T1980I only partially complement BRCA2 knockout cells in response to DNA damage.
T1346I fully rescues DNA damage sensitivity and RAD51 foci formation upon irradiation. (A) Western blot of total cellular lysates from DLD-1 BRCA2-/- cells stably transfected with either empty vector (BRCA2-/-) or BRCA2 Wild Type (WT), BRCA2 S1221P (BRC2), T1346I (Spacer BRC2-3), and T1980I (BRC7) full-length BRCA2 cDNA constructs. BRCA2 was detected with an MBP antibody. 2XMBP-BRCA2 (470 kDa), and RAD51 (37 kDa). (B) Clonogenic survival analyses of stable cell lines treated with mitomycin C (MMC), cisplatin, Olaparib, and Talazoparib. Error bars represent the S.D. for two biological independent experiments. (C) Immunofluorescence images and quantification of RAD51 (green) and gammaH2AX (red) foci, and DAPI staining to visualize nuclei (blue, grey) and micronuclei (grey). Representative images at 6 hr post-IR (12 Gy). Quantification of four biological independent experiments and statistical analysis t-test and one-way ANOVA. Scale bar represents 50 µm. p-value <0.05 p-value <0.0001.
-
Figure 2—source data 1
Contains original gel images depicting cropped regions used for Figure 2A, clonogenic survival data for Figure 2B, foci data for Figure 2C.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig2-data1-v3.zip
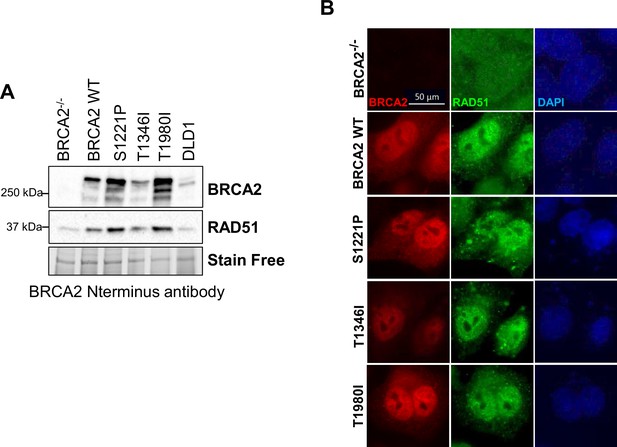
Expression and nuclear localization of S1221P, T1346I, and T1980I BRCA2 proteins.
(A) Western blot of total cellular lysates from DLD1 and DLD-1 BRCA2-/- cells stably transfected with either empty vector (BRCA2-/-) or BRCA2 Wild Type (WT), BRCA2 S1221P (BRC2), T1346I (Spacer BRC2-3), and T1980I (BRC7) full-length BRCA2 cDNA constructs. BRCA2 was detected with rabbit anti-BRCA2 antibody. 2XMBP-BRCA2 (470 kDa), and RAD51 (37 kDa). (B) Localization by immunofluorescence in untreated BRCA2-/- cells stably expressing WT, S1221P, T1346I, and T1980I BRCA2 proteins. Representative images of 2XMBP-BRCA2 (red, anti-MBP), RAD51 (green), and nuclei (blue).
-
Figure 2—figure supplement 1—source data 1
Raw Nt Ct BRCA2 RAD51.
Raw Stain Free. Nt Ct BRCA2 RAD51 highlighedbandsandlabels.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig2-figsupp1-data1-v3.zip
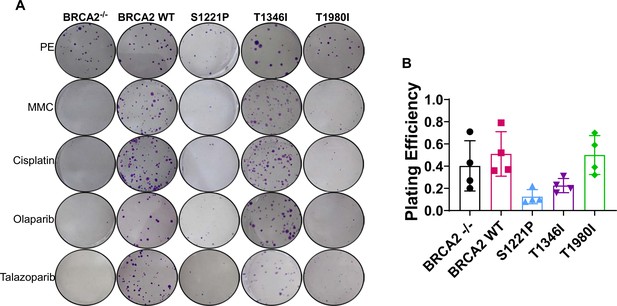
BRCA2-/- cells stably expressing full-length BRCA2 S1221P and T1980I proteins only partially rescue sensitivity to crosslinking agents (MMC or cisplatin) and PARP inhibitors (Olaparib or Talazoparib).
The T1346I protein rescues to the same extent as WT. (A) Representative images of cell colonies stained with crystal violet 14 days after treatment. PE; plating efficiency. Treatments: 2.5 µM MMC 1 hr, 25 µM Cisplatin 1 hr, 15 µM Olaparib 24 hr, 20 nM Talazoparib 24 hr. (B) Plating efficiency of BRCA2-/- cells stably expressing BRCA2.
-
Figure 2—figure supplement 2—source data 1
Revised plating efficiency.xlsx.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig2-figsupp2-data1-v3.xlsx
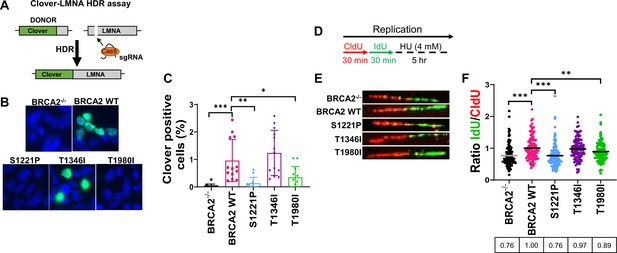
S1221P and T1980I exhibit defects in HDR and fork protection.
(A) Schematic of the Clover-LMNA HDR assay. (B) Immunofluorescence images and quantification of Clover-LMNA HDR (green) and DAPI staining to visualize nuclei (blue) 96 hours post-transfection (threshold 0.25). (C) Quantification of the percentage of cells Clover positive in 4 independent experiments. (D) Schematic of CldU/IdU pulse-labeling followed by a 5 hr hydroxyurea (HU; 4 mM) treatment with representative images (E) of CldU (Red) and IdU (Green) replication tracts after HU treatment. (F) Dot plot of IdU to CldU tract length ratios for individual replication forks in HU-treated cells. All samples are normalized to BRCA2 WT (ratio = 1, i.e., functional fork protection). The median value of 100 or more IdU and CldU tracts per experimental condition is indicated. p-value <0.05, p-value <0.01, p-value <0.001.
-
Figure 3—source data 1
Contains quantification of clover positive cells (Figure 3C), images of DNA fibers (Figure 3E), DNA fiber analysis (Figure 3F).
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig3-data1-v3.zip
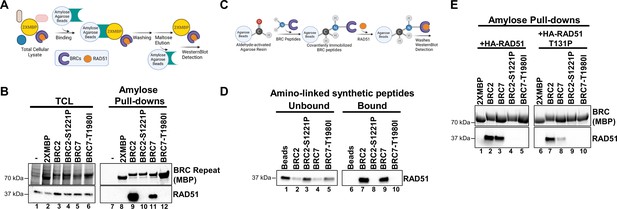
BRC2-S1221P and BRC7-T1980I abolish RAD51 binding.
(A) Schematic of amylose pull-down reaction to detect RAD51 binding. (B) Total cellular lysates (TCL) and amylose pull-downs from HEK 293T cells transiently transfected with 2XMBP, 2XMBP-BRC2, 2XMBP-BRC2 S1221P, 2XMBP-BRC7, and 2XMBP-BRC7 T1980I. Western blot: anti-MBP was used to detect 2XMBP-BRC and anti-RAD51 to detect endogenous RAD51. (C) Schematic of aminolink-conjugated resin reaction to BRC peptides and pull-down reaction to detect purified RAD51 binding. (D) Synthesized peptides BRC2, BRC2 S1221P, BRC7, BRC7 T1980I (35 amino acids) were conjugated to aminolink resin and incubated with purified RAD51. Western blot using RAD51 antibody to detect unbound and bound RAD51 after washes and elution of proteins in laemmli sample buffer. (E) Amylose pull-downs in HEK 293T cells transiently transfected with 2XMBP, 2XMBP-BRC2, 2XMBP-BRC2-S1221P, 2XMBP-BRC7, 2XMBP-BRC7 T1980I, wild type HA-RAD51, and T131P HA-RAD51. Anti-MBP was used to detect 2XMBP-BRC proteins. HA antibody was used to detect recombinant RAD51.
-
Figure 4—source data 1
Contains original (uncropped) gel images for Figure 4B, D, E.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig4-data1-v3.zip
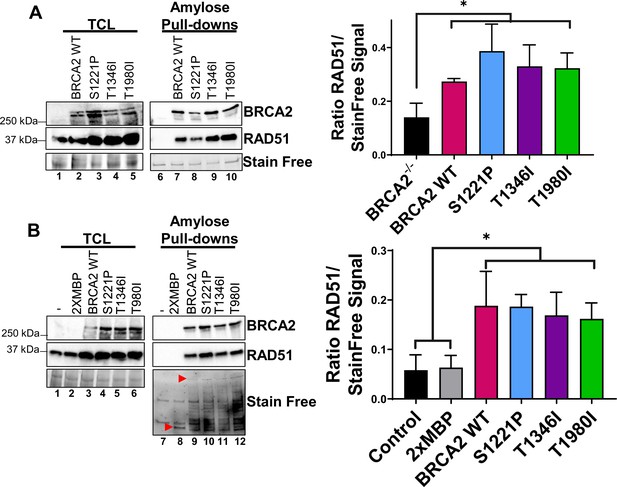
S1221P, T1346I, and T1980I full-length BRCA2 proteins bind RAD51.
(A) Western blots of total cellular lysates (TCL) (left gel) or amylose pull-downs (right gel) from BRCA2-/- cells stably expressing full-length BRCA2 WT, S1221P, T1346I, and T1980I. 2XMBP-BRCA2 (470 kDa) detected using a C-terminal antibody. Depicted RAD51 is the endogenous protein (37 kDa). (B) Western blots of total cellular lysates (TCL) and amylose pull-downs from HEK 293T cells transiently transfected with 2XMBP-BRCA2 WT, S1221P, T1346I, and T1980I full-length constructs. Anti-MBP antibody was used for BRCA2 detection. Depicted RAD51 is the endogenous protein. Lower red arrow in the Stain Free panel indicates 2XMBP (84 kDa) and upper red arrow indicates 2XMBP-BRCA2 (470 kDa) expression. Graphs on the right represent relative RAD51 expression levels using densitometric analysis from the western blots of total cellular lysates. Quantification of four biological independent experiments and statistical analysis t-test of each sample compared to BRCA-/-. p-value < 0.05.
-
Figure 4—figure supplement 1—source data 1
Contains band quantification and original (uncropped) gel images.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig4-figsupp1-data1-v3.zip
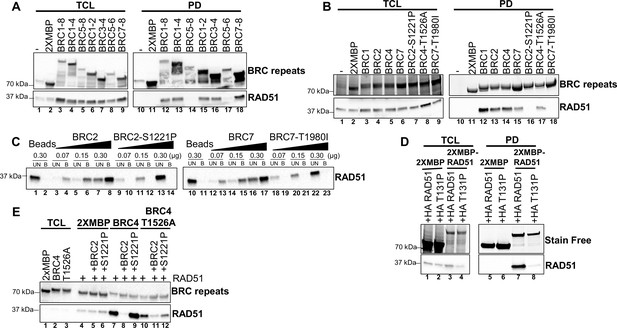
BRC1-2, BRC7-8, BRC2, and BRC7 bind RAD51 whereas BRC2-S1221P and BRC7-T1980I ablate RAD51 binding.
(A) Western blots of total cellular lysates (TCL) and amylose pull-downs from HEK 293T cells transiently transfected with 2XMBP or 2XMBP-BRC repeat constructs: BRC1-8, BRC1-4, BRC5-8, BRC1-2, BRC3-4, BRC 5–6, and BRC 7–8. (B) Western blots of total cellular lysates (TCL) and amylose pull-downs from HEK 293T cells transiently transfected with 2XMBP or 2XMBP-BRC repeat constructs: BRC1, BRC2, BRC4, BRC7, BRC2-S1221P, BRC4-T1526A, and BRC7-T1980I. BRC proteins were detected with an MBP antibody (C) Pull-downs with synthetic peptides: BRC2, BRC2-S1221P, BRC7, BRC7-T1980I fused to aminolink resin. Ten µL of aminolink resin conjugated to each peptide was incubated with increasing concentrations of purified RAD51 protein. Western blots (anti-RAD51) from unbound (UB) and bound (B) elutions from aminolink resin. (D) RAD51 self-interaction pull-down assay. Western blots of total cellular lysates and amylose pulldowns from HEK 293T cells transiently co-transfected with 2XMBP or 2XMBP-RAD51 and HA-RAD51 or HA-RAD51 T131P. RAD51 was detected with an HA antibody. (E) Western blots from peptide competition assay. Purified RAD51 previously incubated with or without BRC2 or BRC2-S1221P peptide was utilized in an amylose pull-down assay with 2XMBP-BRC4 or 2XMBP-BRC4-T1526A recombinantly expressed in HEK 293T cells. BRC repeat proteins detected with anti-MBP. Purified RAD51 eluted from amylose beads bound to 2XMBP-BRC peptide detected with anti-RAD51.
-
Figure 4—figure supplement 2—source data 1
Contains original (uncropped) gel images.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig4-figsupp2-data1-v3.zip
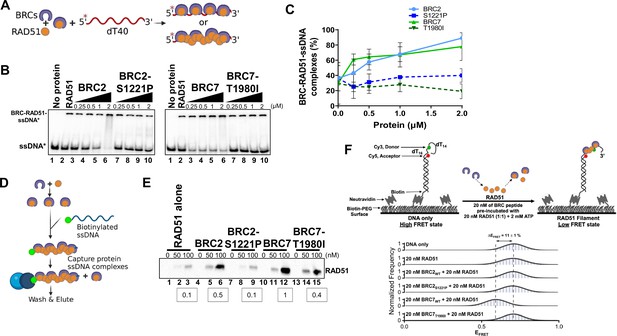
BRC2-S1221P and BRC7-T1980I fail to stimulate RAD51-ssDNA complex formation.
(A) Schematic of reaction to assay BRC repeat stimulation of RAD51-ssDNA complex formation by EMSA. RAD51 was pre-incubated with increasing amounts of BRC protein for 15 min then radiolabeled ssDNA (dT40) was added for 40 min. All reactions were incubated at 37 degrees and visualized on a 6% TAE polyacrylamide gel. (B) Autoradiograms of EMSA gels depicting increasing concentration of BRC proteins: BRC2, BRC2-S1221P, BRC7, BRC7-T1980I incubated with 10 nM RAD51 and 400 pM ssDNA (dT40*). Lane 1 is no protein control. Lane 2 is RAD51 alone. (C) Quantification of BRC-RAD51-ssDNA complexes calculated from gels shown in B. Error bars represent the S.D. for two biological independent experiments. (D) Schematic of biotinylated DNA pull-down assay. Purified BRC proteins were pre-incubated with increasing concentrations of purified RAD51 for 10 min. Biotinylated ssDNA (167-mer) was then added for 10 min to allow nucleoprotein filament formation and captured on magnetic streptavidin beads. The beads were then washed, eluted in sample buffer, analyzed by SDS-PAGE, and bound RAD51 was detected by western blotting using an anti-RAD51 antibody. (E) Western blot depicting RAD51 pulled down and eluted from biotin-DNA-BRC-RAD51 complexes. 0, 50, or 100 nM RAD51 was pre-incubated with 80 nM of BRC peptide, incubated with biotin-ssDNA, washed extensively, and eluted. Densitometric quantitation of RAD51 binding at 100 nM is indicated in the boxes below. (F) Schematic depicting single-molecule FRET (smFRET) assay with 30-nucleotide 3’-tail ssDNA. Addition of BRC7WT results in an increase in RAD51 binding leading to a transition from high FRET (DNA-only) to medium FRET (BRC7WT and RAD51-bound). Histograms display a shift (ΔEFRET = 11%) upon addition of 20 nM BRC7WT and 20 nM RAD51 that is not observed in BRC7T1980I or in the BRC2 peptides (WT and S1221P). Representative histograms do not include zero FRET values or photobleached portions of the FRET trajectories. A minimum of 250 smFRET trajectories were utilized to generate each histogram.
-
Figure 5—source data 1
Contains uncropped gel images (Figure 5B), EMSA band quantification (Figure 5C), biotin-DNA pulldown band quantification (Figure 5E), histogram data (Figure 5F).
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig5-data1-v3.zip
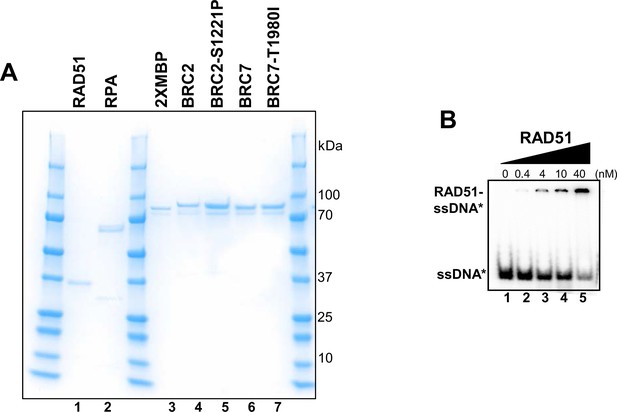
Purified proteins used for biochemical studies.
Optimization of RAD51-ssDNA EMSA. (A) Coomassie stained gel of 2XMBP-BRC fragments BRC2, BRC2-S1221P, BRC7, and BRC7-T1980I proteins purified from HEK 293T cells. Human RAD51 and RPA proteins purified from bacteria. (B) Increasing concentrations of purified RAD51 were incubated with radiolabeled dT40 and analyzed by EMSA.
-
Figure 5—figure supplement 1—source data 1
Contains original (uncropped) gel images.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig5-figsupp1-data1-v3.zip
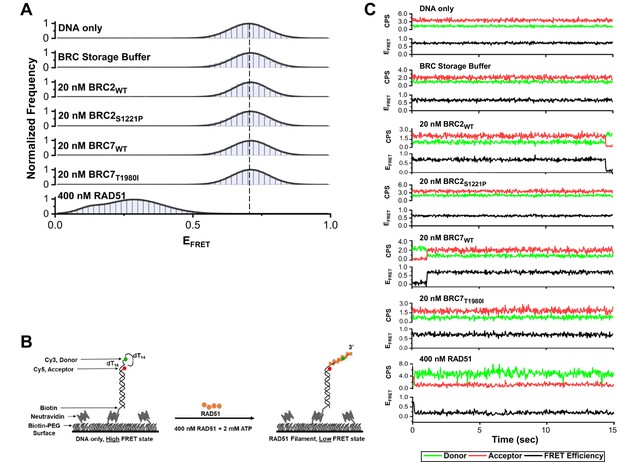
smFRET control assays highlighting BRC wildtype and mutant proteins (S1221P and T1980I) show no change in FRET efficiency in the absence of RAD51.
(A) Histograms depicting: DNA only, BRC storage buffer only, individual BRC repeat proteins in the absence of RAD51 protein, and 400 nM RAD51 alone. Representative histograms do not include zero FRET values or photobleached portions of the FRET trajectories. (B) Illustration of smFRET assay consisting of a 3’-tail DNA (30 nucleotide ssDNA overhang) tethered to a dsDNA substrate attached to the surface. (C) Representative smFRET trajectories for DNA only, BRC storage buffer, BRC peptides (wildtype and mutants), and RAD51 alone. At 400 nM RAD51, there is a notable decrease in acceptor intensity (red) indicating RAD51 binding and transition from a high FRET state (DNA-only) to a low FRET state (RAD51-bound). The donor traces are in green, acceptor in red, and FRET trajectories are shown in black.
-
Figure 5—figure supplement 2—source data 1
Contains data and traces for single molecule FRET.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig5-figsupp2-data1-v3.zip
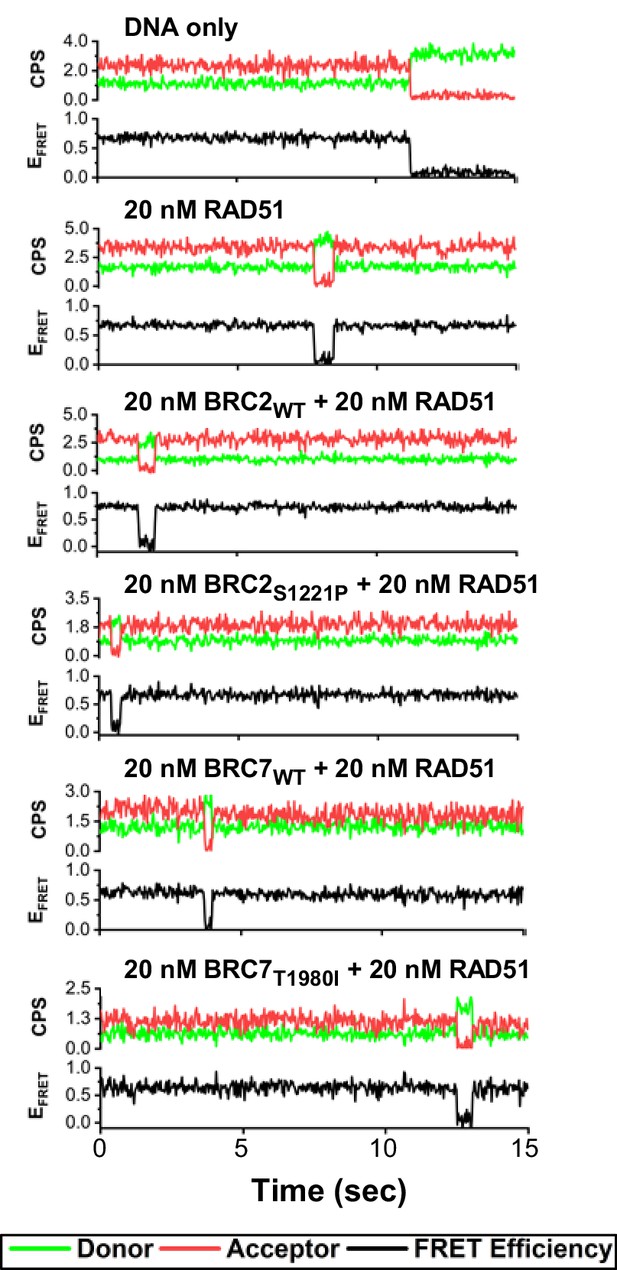
Representative trajectories corresponding to histograms in Figure 5F.
Representative individual smFRET trajectories for DNA only, 20 nM RAD51, 20 nM RAD51 in presence of 20 nM BRC2 (WT or S1221P), and 20 nM RAD51 in presence of 20 nM BRC7 (WT or T1980I). The donor traces are in green, acceptor in red, and FRET trajectories are shown in black.
-
Figure 5—figure supplement 3—source data 1
Contains data and traces for single molecule FRET.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig5-figsupp3-data1-v3.zip
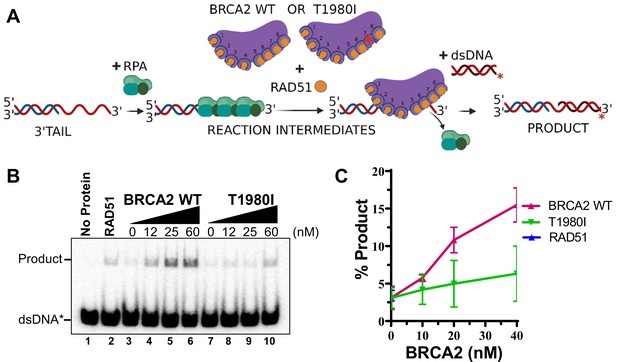
RAD51-mediated DNA strand exchange activity is stimulated by increasing amounts of WT BRCA2 but not T1980I.
(A) Diagram of DNA strand exchange assay. RPA was pre-bound to the 3’ tail DNA substrate for 5 min. Increasing amounts of WT BRCA2 or T1980I protein was then added in combination with RAD51 for 5 min, followed by the addition of radiolabeled donor dsDNA for 30 min. All reactions were incubated at 37 degrees. The reaction was then deproteinized and run on a 6% TAE polyacrylamide gel. (B) Autoradiograms of PAGE gels used to analyze the products of DNA strand exchange. Lane 1 is no protein control. Lane 2 is RAD51 in the absence of RPA or BRCA2. (C) Quantification of product formation from autoradiogram in (B); mean values were plotted. Errors bars represent the S.D. of three biological independent experiments.
-
Figure 6—source data 1
Contains original (uncropped) gel image (Figure 6B) and band quantitation (Figure 6C).
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig6-data1-v3.zip
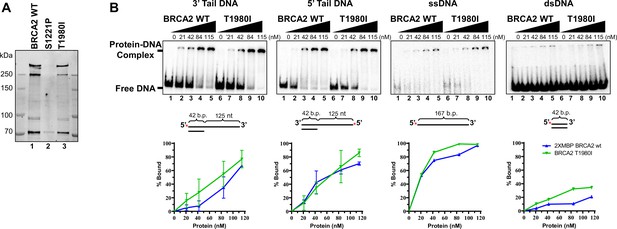
Purified full-length BRCA2 WT and T1980I proteins bind a panel of DNA substrates with equivalent efficiencies.
(A) Stain Free gel (4–15% gradient SDS-PAGE) of full-length 2XMBP-BRCA2 proteins purified from 293T cells. (B) EMSAs depicting increasing concentrations of BRCA2 WT and T1980I proteins incubated with 3’ Tail, 5’ Tail, ssDNA, and dsDNA substrates. Percentage of protein-DNA complex was calculated as the free DNA remaining in a lane relative to the protein-free lane (0% complex or 100% free DNA). Error bars are S.D. from 4 (3’tail) and 2 (5’tail) independent experiments.
-
Figure 6—figure supplement 1—source data 1
Contains original (uncropped) gel images and band quantitation.
- https://cdn.elifesciences.org/articles/79183/elife-79183-fig6-figsupp1-data1-v3.zip
Tables
Amino acid sequences of synthesized peptides.
| PEPTIDES | AMINO ACID SEQUENCES |
|---|---|
| BRC2 | NEVGFRGFYSAHGTKLNVSTEALQKAVKLFSDIEN |
| BRC2-S1221P | NEVGFRGFYPAHGTKLNVSTEALQKAVKLFSDIEN |
| BRC7 | SANTCGIFSTASGKSVQVSDASLQNARQVFSEIED |
| BRC7-T1980I | SANTCGIFSIASGKSVQVSDASLQNARQVFSEIED |
Amino acid sequences of BRC purified proteins.
| CONSTRUCTS | AMINO ACID SEQUENCE |
|---|---|
| 2xMBP BRC2 | YLTDENEVGFRGFYSAHGTKLNVSTEALQKAVKLFSDIENISEETSAEVHPISL* |
| 2xMBP BRC2-S1221P | YLTDENEVGFRGFYPAHGTKLNVSTEALQKAVKLFSDIENISEETSAEVHPISL* |
| 2xMBP BRC4 | RDEKIKEPTLLGFHTASGKKVKIAKESLDKVKNLFDEKEQGTSEI* |
| 2xMBP BRC4-T1526A | RDEKIKEPTLLGFHAASGKKVKIAKESLDKVKNLFDEKEQGTSEI* |
| 2xMBP BRC7 | GKLHKSVSSANTCGIFSTASGKSVQVSDASLQNARQVFSEIEDSTKQ* |
| 2xMBP BRC7-T1980I | GKLHKSVSSANTCGIFSIASGKSVQVSDASLQNARQVFSEIEDSTKQ* |
Oligonucleotide sequences.
| Oligonucleotides | Sequence |
|---|---|
| RJ-167-mer | 5′-CTG CTT TAT CAA GAT AAT TTT TCG ACT CAT CAG AAA TAT CCG TTT CCT ATA TTT ATT CCT ATT ATG TTT TAT TCA TTT ACT TAT TCT TTA TGT TCA TTT TTT ATA TCC TTT ACT TTA TTT TCT CTG TTT ATT CAT TTA CTT ATT TTG TAT TA TCC TTA TCT TAT TTA-3′ |
| RJ-5’Tail-167mer | 5’-ATT TAT TCT ATT CCT CTT TAT TTT CTC TGT TTA TTC ATT TAC TTA TTT TGT ATT AAT TTC CTA TAT TTT TTA CTT GTA TTT CTT ATT CAT TTA CTT ATT TTG TAT TAT CCT TAT TTA TAT CCT TTC TGC TTT ATC AAG ATA ATT TTT CGA CTC ATC AGA AAT ATC CG-3’ |
| RJ-167-mer complementary | 5’-TAA ATA AGA TAA GGA TAA TAC AAA ATA AGT AAA TGA ATA AAC AGA GAA AAT AAA GTA AAG GAT ATA AAA AAT GAA CAT AAA GAA TAA GTA AAT GAA TAA AAC ATA ATA GGA ATA AAT ATA GGA AAC GGA TAT TTC TGA TGA GTC GAA AAA TTA TCT TGA TAA AGC AG-3’ |
| RJ-PHIX-42–1 | 5′-CGG ATA TTT CTG ATG AGT CGA AAA ATT ATC TTG ATA AAG CAG-3′ |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Homo sapiens, Drosophila melanogaster, Mus musculus Felis catus, Rattus norvegicus, Strongylocentrotus purpuratus, Monodelphis domestica, Canis lupus familiaris, Xenopus tropicalis) | BRCA2 | Uniprot | P51587 Homo sapiens (Human); Q9W157 Drosophila melanogaster (Fruit fly); P97929Mus musculus (Mouse); Q864S8Felis catus (Cat); O35923Rattus norvegicus (Rat); A5A3F7 Strongylocentrotus purpuratus (Purple sea urchin); A4ZZ89Monodelphis domestica (Gray short-tailed opossum); Q8MKI9Canis lupus familiaris (Dog) (Canis familiaris); A4ZZ90Xenopus tropicalis (Western clawed frog) (Silurana tropicalis). | Alignment Figure 1 |
| Strain, strain background (Escherichia coli) | Stellar Competent Cells | Takara-Clontech | Cat# 636766 | Competent cells |
| Strain, strain background (Escherichia coli) | Rosetta(DE3)pLysS Competent Cells | Sigma-Aldrich | Cat# 70956 | |
| Strain, strain background (Escherichia coli) | E. coli Acella(DE3)/pCH1–RAD51o | Subramanyam and Spies, 2018 | ||
| Cell line (Homo-sapiens) | DLD1 | Horizon Discovery Hucl et al., 2008 | ||
| Cell line (Homo-sapiens) | DLD1 BRCA2 -/- | Horizon Discovery Hucl et al., 2008 | ||
| Cell line (Homo-sapiens) | HEK2093T | Jensen et al., 2010 | ||
| Recombinant DNA reagent | pBluescript BRCA2 (1–5286 bp) | This paper | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Recombinant DNA reagent | pUC57 BRCA2 (2141–9117 bp) Genescript | Synthetized by Genescript | ||
| Recombinant DNA reagent | pX330-LMNA-gRNA1 | addgene | 122507 | sgRNA plasmid targeting Lamin A |
| Recombinant DNA reagent | pCR2.1 Clover-LMNA Donor | Addgene | 122507 | Donor |
| Transfected construct (Homo-sapiens) | 2XMBP-BRC2 | This paper | YLTDENEVGFRGFYSAHGTKLNV STEALQKAVKLFSDIENISE ETSAEVHPISL* | Dr. Ryan Jensen laboratory. Material & Methods section |
| Transfected construct (Homo-sapiens) | 2XMBP-BRC2-S1221P | This paper | YLTDENEVGFRGFYPAHGTKLN VSTEALQKAVKLFSDIENISE ETSAEVHPISL* | Dr. Ryan Jensen laboratory. Material & Methods section |
| Transfected construct (Homo-sapiens) | 2XMBP-BRC7 | This paper | GKLHKSVSSANTCGIFSTASGKS VQVSDASLQNARQVFSEIEDSTKQ* | Dr. Ryan Jensen laboratory. Material & Methods section |
| Transfected construct (Homo-sapiens) | 2XMBP-BRC7-T1980I | This paper | GKLHKSVSSANTCGIFSIASGKSV QVSDASLQNARQVFSEIEDSTKQ* | Dr. Ryan Jensen laboratory. Material & Methods section |
| Transfected construct (Homo-sapiens) | 2XMBP BRCA2 WT | Jensen et al., 2010 | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Transfected construct (Homo-sapiens) | 2XMBP BRCA2 S1221P | This paper | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Transfected construct (Homo-sapiens) | 2XMBP BRCA2 T1346I | This paper | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Transfected construct (Homo-sapiens) | 2XMBP BRCA2 T1980I | This paper | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Peptide, recombinant protein | RPA | This paper, Anand et al., 2018; | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Peptide, recombinant protein | RAD51 | This paper, Subramanyam and Spies, 2018 | Dr. Ryan Jensen laboratory. Material & Methods section | |
| Peptide, recombinant protein | BRC2 peptide | Pierce, This paper | NEVGFRGFYSAHGTKLN VSTEALQKAVKLFSDIEN | Dr. Ryan Jensen laboratory. Material & Methods section |
| Peptide, recombinant protein | BRC2-S1221P peptide | Pierce, This paper | NEVGFRGFYPAHGTKL NVSTEALQKAVKLFSDIEN | Dr. Ryan Jensen laboratory. Material & Methods section |
| Peptide, recombinant protein | BRC7- peptide | Pierce, This paper | SANTCGIFSTASGKSVQ VSDASLQNARQVFSEIED | Dr. Ryan Jensen laboratory. Material & Methods section |
| Peptide, recombinant protein | BRC7- T1980I peptide | Pierce, This paper | SANTCGIFSIASGKSVQV SDASLQNARQVFSEIED | Dr. Ryan Jensen laboratory. Material & Methods section |
| Antibody | Anti-MBP (mouse monoclonal) | NEB | Cat# E8032L | IF (1:100) WB (1:1000) |
| Antibody | Anti-BRCA2 (rabbit polyclonal) | Abcam | Cat# ab27976 | IF (1:100) WB (1:1000) |
| Antibody | Anti-BRCA2 Nt (rabbit polyclonal) | Bethyl Laboratories | Cat# A303-434 | WB (1:1000) |
| Antibody | Anti-RAD51 (rabbit polyclonal) | Proteintech | Cat# 14961–1-AP, RRID:AB_2177083 | IF (1:100) WB (1:1000) |
| Antibody | Anti-RAD51 (mouse monoclonal) | Novus Biologicals | Cat# NB 100–148, RRID:AB_350083 | WB (1:1000) |
| Antibody | Rabbit Anti-RAD51 (rabbit polyclonal) | Abcam | Cat# ab63801, RRID:AB_1142428 | IF (1:100) |
| Antibody | Anti-BrdU (purified mouse monoclonal) | BD Biosciences | Cat# 347580, RRID:AB_10015219 | 1:10 |
| Antibody | Anti-BrdU (rat monoclonal) | Abcam | Cat# ab6326, RRID:AB_2313786 | 1:40 |
| Antibody | Anti-Rat Cy5 (goat polyclonal) | Abcam | Cat# ab6565, RRID:AB_955063 | 1:100 |
| Antibody | Anti-Mouse Cy3 (sheep polyclonal) | Sigma Aldrich | Cat# C2181, RRID:CVCL_K942 | 1:100 |
| Antibody | m-IgGκ BP-HRP | Santa Cruz Biotechnology | Cat# sc-516102, RRID:AB_2687626 | WB (1:10000) |
| Antibody | anti-rabbit IgG-HRP (goat polyclonal) | Santa Cruz Biotechnology | Cat# sc-2004, RRID:AB_631746 | WB (1:10000) |
| Antibody | anti-Rabbit IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 (goat polyclonal) | Thermo Fisher Scientific | Cat# A-11034, RRID:AB_2576217 | IF (1:10000) |
| Antibody | anti-Mouse IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 546 (goat polyclonal) | Thermo Fisher Scientific | Cat# A-11003, RRID:AB_2534071 | IF (1:10000) |
| Sequenced- based reagent | dT40 | Carreira and Kowalczykowski, 2011 | EMSA | 5’TTTTTTTTTTTTTTTTTTTTTT TTTTTTTTTTTTTTTTTT3’ |
| Sequenced- based reagent | RJ-167-mer | Jensen et al., 2010 | EMSA DNA strand exchange | 5′-CTG CTT TAT CAA GAT AAT TTT TCG ACT CAT CAG AAA TAT CCG TTT CCT ATA TTT ATT CCT ATT ATG TTT TAT TCA TTT ACT TAT TCT TTA TGT TCA TTT TTT ATA TCC TTT ACT TTA TTT TCT CTG TTT ATT CAT TTA CTT ATT TTG TAT TA TCC TTA TCT TAT TTA-3′ |
| Sequenced- based reagent | RJ-5’tail 167-mer | Jensen et al., 2010 | EMSA | : 5’-ATT TAT TCT ATT CCT CTT TAT TTT CTC TGT TTA TTC ATT TAC TTA TTT TGT ATT AAT TTC CTA TAT TTT TTA CTT GTA TTT CTT ATT CAT TTA CTT ATT TTG TAT TAT CCT TAT TTA TAT CCT TTC TGC TTT ATC AAG ATA ATT TTT CGA CTC ATC AGA AAT ATC CG-3’ |
| Sequenced- based reagent | RJ-PhiX-42–1 | Jensen et al., 2010 | EMSA DNA Strand Exchange | 5’-CGG ATA TTT CTG ATG AGT CGA AAA ATT ATC TTG ATA AAG CAG-3' |
| Sequenced- based reagent | PS4 | Jensen et al., 2010 | EMSA DNA Strand Exchange | 5’- TAATACAAAATAAGTAAATGA ATAAACAGAGAAAATAAAG –3' |
| Sequenced- based reagent | PS5 | Jensen et al., 2010 | EMSA DNA Strand Exchange | 5’- CTTTATTTTCTCTGTTTATTCATT TACTTATTTTGTATTA –3' |
| Chemical compound, drug | Jet Optimus | Polyplus Transfection | 117–07 | |
| Chemical compound, drug | Calcium Phosphate transfection | This paper | https://www.nature.com/articles/nmeth0405-319 | Dr. Ryan Jensen laboratory. Material & Methods section |
| Chemical compound, drug | Lipofectamine3000 | Thermo Scientific | L3000001 | |
| Chemical compound, drug | TurboFect reagent | Thermo Scientific | R0533 | |
| Chemical compound, drug | G418 | American Bio | AB05058-00020 | |
| Chemical compound, drug | protease inhibitor cocktail | Sigma Aldrich | 04693159001 | |
| Chemical compound, drug | amylose resin | NEB | E8021 | |
| Chemical compound, drug | Hydroxyurea | Sigma Aldrich | 127-07-1 | |
| Chemical compound, drug | Benzonase | Millipore | 9025-65-4 | |
| Chemical compound, drug | MMC | Sigma Aldrich | M4287 | |
| Chemical compound, drug | Cisplatin | Sigma Aldrich | 15663-27-1 | |
| Chemical compound, drug | Olaparib | ApexBio | AZD2281, Ku-0059436 | |
| Chemical compound, drug | Talazoparib-BMN673 | ApexBio | A4153 | |
| Chemical compound, drug | DTT | Sigma Aldrich | 3483-12-3 | |
| Chemical compound, drug | IdU | Sigma Aldrich | I7125 | |
| Chemical compound, drug | CldU | Sigma Aldrich | C6891 | |
| Chemical compound, drug | FluorSave reagent | Calbiochem | 345789 | |
| Software, algorithm | ClustalIX | Larkin et al., 2007 | http://www.ch.embnet.org/software/ClustalW.html | |
| Software, algorithm | Bioedit | Bioedit (Hall, 1999) | https://itservices.cas.unt.edu/software/bioedit725 | |
| Software, algorithm | Snapgene 5.3.1. | Snapgene 5.3.1. | from Insightful Science; available at snapgene.com | |
| Software, algorithm | Prism Graph Pad version 9.0 | https://www.graphpad.com/updates/prism-900-release-notes | ||
| Software, algorithm | PyMOL Molecular Graphics System, Version 1.3 Schrödinger, LLC. | Schrodinger, L. (2010) The PyMOL Molecular Graphics System | ||
| Software, algorithm | Image J | Schneider et al., 2012 | https://imagej.nih.gov/ij/ | |
| Software, algorithm | ImageQuant TL 8.0 image analysis software | http://www.hhmi.umbc.edu/downloads/Imaging%20support%20GE/IQ%20TL%20collateral/IQTL_UserManual%208.pdf | ||
| Software, algorithm | Image Lab Software 6.1 (Bio-Rad) | https://www.bio-rad.com/en-us/product/image-lab-software?ID=KRE6P5E8Z&WT.mc_id=220411034041&WT.knsh_id=_kenshoo_clickid_&gclid=Cj0KCQjwr4eYBhDrARIsANPywCiJded2fEKfZWaf_-Fxki9ynO1VFgLb3kkN-gJ-A3u4baPwfvKj2ZQaApk-EALw_wcB | ||
| Software, algorithm | ChemiDoc MP imaging system XRS+ (Bio-Rad) | https://www.bio-rad.com/en-us/product/chemidoc-mp-imaging-system?ID=NINJ8ZE8Z&gclid=Cj0KCQjwr4eYBhDrARIsANPywCjmMNj5mMKTHZkXXj4iOrhnRLMo8JRt4nwLib3XTupvbHi2Q2CRJnkaAvp6EALw_wcB | ||
| Software, algorithm | Cell profiler software following counting and scoring pipeline | Carpenter et al., 2006 | ||
| Other | PDB structure BRC4 | Pellegrini et al., 2002 | 1NOW | Swiss model BRC2-7 |
| Other | DAPI stain | Thermo Fisher | D1306 | (1 µg/mL) |
| Other | NucRed Live 647 ReadyProbes Reagent | Thermo Fisher | R37106 | (1 µg/mL) |