Glial-dependent clustering of voltage-gated ion channels in Drosophila precedes myelin formation
Figures
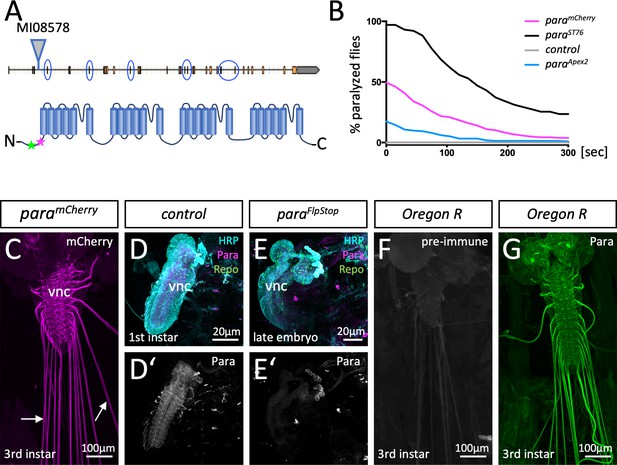
Localization of Paralytic (Para) voltage-gated ion channels in the larval nervous system.
(A) Schematic view on the para gene. Alternative splicing at the circled exons results in the generation of more than 60 Para isoforms. All isoforms share a common N-terminus. Here, the MiMIC insertion MI08578 allows tagging of the endogenous para gene. The peptide sequence AEHEKQKELERKRAEGE (positions 33–49) that was used for immunization is indicated by a green star, the MiMIC insertion is indicated by a magenta star. (B) Homozygous paramCherry, paraApex2 or paraST76 flies were tested for temperature-induced paralysis. The recovery time is indicated. (C) Third instar larval paramCherry nervous system stained for Cherry localization. ParamCherry is detected in the ventral nerve cord (vnc) and diffusely along peripheral nerves (arrows). (D,D’) Affinity purified anti-Para antibodies detect a protein in the CNS of dissected 24-hr-old wild type first instar larvae. (E,E’) No protein is found in the CNS of dissected age-matched para mutant animals. (F) Third instar larval nervous system stained with the pre-immune control. (G) Third instar larval nervous system stained with affinity purified anti-Para antibodies. Scale bars are as indicated.
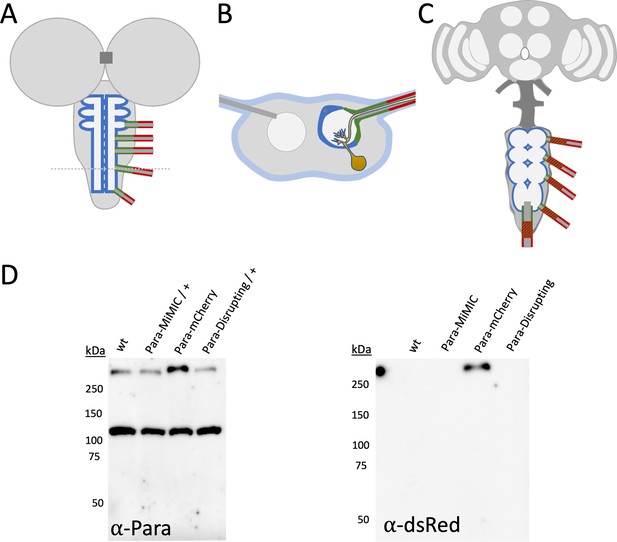
Ensheathing- / wrapping glia in Drosophila and characterization of Para protein.
(A–C) Schematic representation of the larval (A,B) and the adult Drosophila nervous system (C). The ensheathing glia is labeled in blue, the ensheathing/wrapping glia is labeled in green, the wrapping glia is shown in red. The tract glia of the adult nervous system is shown in green and red stripes. The tract glia likely corresponds to the ensheathing/wrapping glia but the exact lineage relationship is not known. (D) Western blot of protein lysates of adult heads. Purified anti-Paralytic (Para) antibodies detect a band of 105 kDa and a band of >250 kDa in size. The size of the >250 kDa protein band increases in paramCherry heads compared to wild type control as well as paraMiMIC heads, indicating that this band corresponds to the Para protein. Note that elevated levels of the endogenous Para::mCherry fusion protein are detected. Anti-dsRed antibodies detect only the ParamCherry fusion protein.
-
Figure 1—figure supplement 1—source data 1
Four images of western blots, with and without marker bands, are provided.
- https://cdn.elifesciences.org/articles/85752/elife-85752-fig1-figsupp1-data1-v2.zip
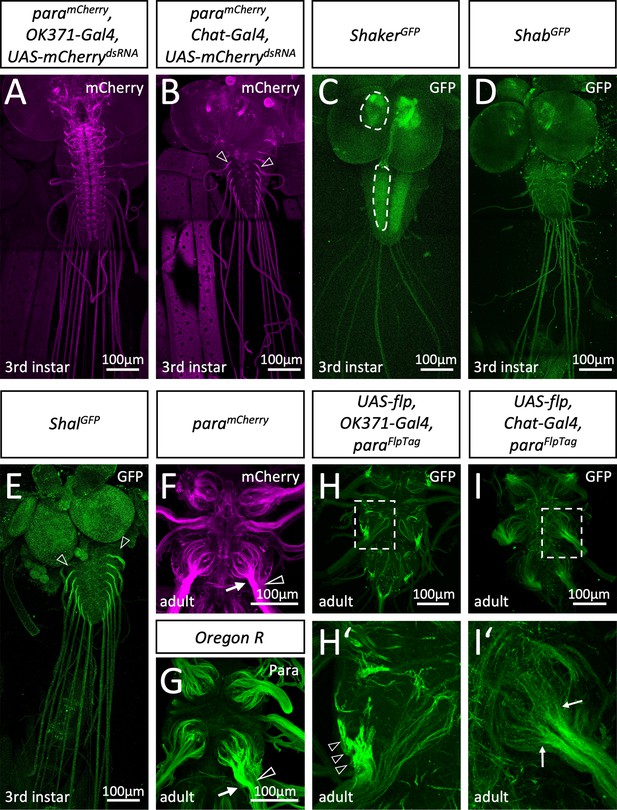
Differential localization of voltage-gated ion channels in Drosophila.
(A) Third instar larvae with the genotype [paramCherry; OK371-Gal4, UAS-mCherrydsRNA]. paramCherry expression is suppressed in all glutamatergic neurons and thus, ParamCherry localization along axons of cholinergic sensory neurons becomes visible. (B) Third instar larvae with the genotype [paramCherry; Chat-Gal4, UAS-mCherrydsRNA]. Here, expression of paramCherry is suppressed in all cholinergic neurons which reveals Paralytic (Para) localization in motor neurons. Note the prominent Para localization at the CNS/PNS transition point (arrowheads). (C) Third instar larval shakerGFP nervous system stained for GFP localization. Shaker is found in the neuropil (dashed areas). (D) Third instar larval shabGFP nervous system stained for GFP localization. Shab is distributed evenly along all peripheral axons. (E) Third instar larval shalGFP nervous system stained for GFP localization. Shal localizes similar as Para on motor axons. Scale bars are 100 µm. (F) Adult paramCherry ventral nerve cord stained for Para localization. ParamCherry localizes prominently along segments of peripheral nerves (arrow) as they enter thoracic neuromeres. Note that some axons entering the CNS neuropil show only a weak Para signal (open arrowhead). (G) Control (Oregon R) adult ventral nerve cord stained for Para protein localization using purified anti-Para antibodies. Note the differential localization of Para along axons entering the nerve (arrow, open arrowhead). (H) Ventral nerve cord of an adult fly with the genotype [paraFlpTag-GFP; Ok371-Gal4, UAS-flp]. The boxed area is shown enlarged in (H’). The arrowheads point to high density of Para. (I) Ventral nerve cord of an adult fly with the genotype [paraFlpTag-GFP; Chat-Gal4, UAS-flp]. The boxed area is shown enlarged in (H’). Note that Para localization is reduced as soon as axons enter the neuropil (arrows). Scale bars are as indicated.
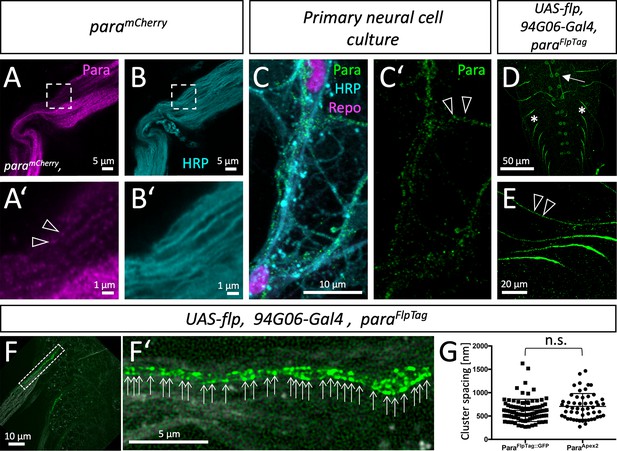
Clustered localization of Paralytic (Para) along motor axons.
(A) High-resolution Airyscan analysis of ParamCherry and (B) HRP localization in an adult nerve. The boxed area is shown in higher magnification below (A’,B’). Note the clustered appearance of ParamCherry, clusters are about 0.6–0.8 µm apart (arrowheads). (C,C’) Primary wild type neural cells cultured for 7 days stained for Repo (magenta) to label glial nuclei, HRP (cyan) to label neuronal cell membranes and anti-Para antibodies (green). The Para protein localizes in a dotted fashion. (D) Ventral nerve cord of a third instar larva with the genotype [paraFlpTag-GFP; 94G06-Gal4, UAS-flp]. The arrow points to a single neuronal cell body found in every hemineuromer. (E) Higher magnification of single ParaGFP expressing axons. Note the dotted arrangement of ParaGFP along motor axons (arrowheads). (F,F’) Ventral nerve cord of a third instar larva with the genotype [paraFlpTag-GFP; 94G06-Gal4, UAS-flp] imaged with super-resolution. The dashed box is shown in high magnification in (F’). Arrows point to clusters of Para protein. Scale bars are as indicated. (G) Quantification of Para cluster distance using super-resolution imaging (ParaFlpTag::GFP, average distance is 620 nm, n=91 clusters on 3 axons, 2 larvae) or electron microscopy (ParaApex2, average distance is 706 nm, n=64 clusters on 8 axons 4 animals, Mann-Whitney U-test, p=0.0747, two-tailed). Scale bars are as indicated.
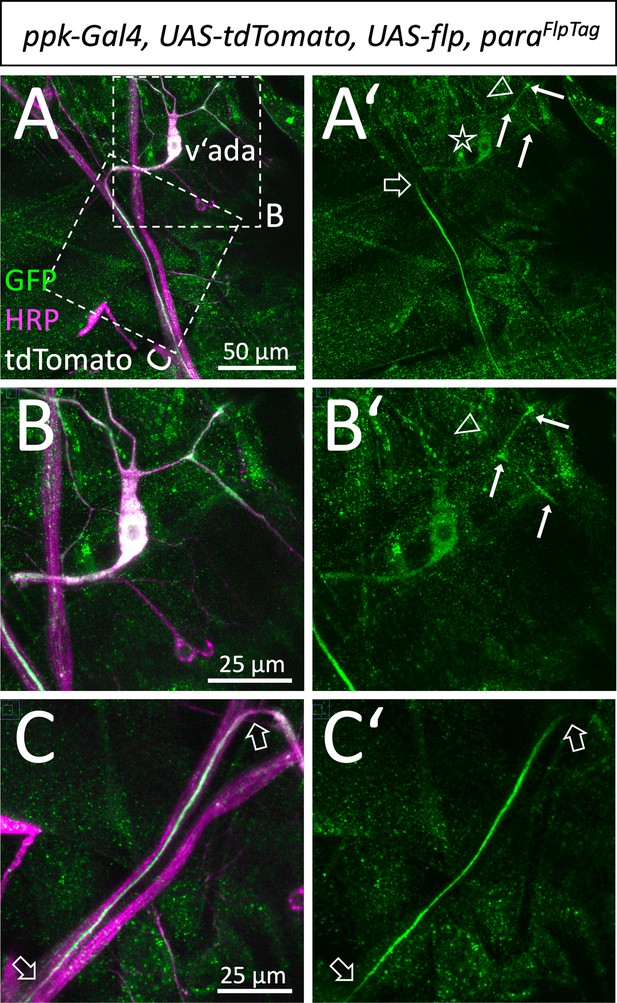
Localization of Paralytic (Para) along sensory axons.
(A,A’) Ventral pickpocket expressing sensory neuron (v’ada) of a third instar larva with the genotype [paraFlpTag-GFP; ppk-Gal4, UAS-flp, UAS-tdTomato] stained for GFP (green), HRP (magenta), and tdTomato (white). The dashed boxes are shown in higher magnification in (B,C). The asterisk denotes the position of the neuronal cell soma. The filled arrows indicate localized Para along some of the dendritic processes. The open arrowhead points to a dendritic process lacking Para localization. Note that Para localization along the descending axon becomes prominent only after about 50 µm (open arrow). (B,B’) Magnification of the neuronal soma attached dendrites. (C,C’) Descending axon of the v’ada neuron. Note that the strong Para signal starts 50 µm distal to the cell soma and fades out after 100 µm (open arrows). Scale bars are as indicated.
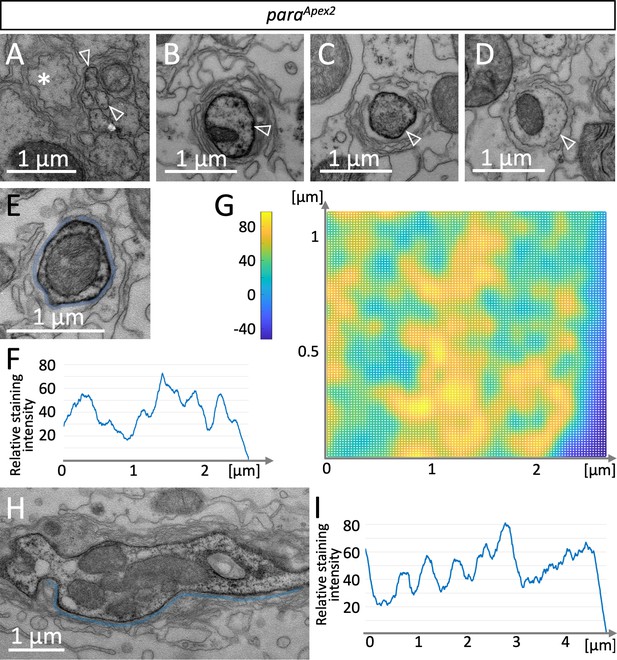
A glial lacunar system surrounds the axon initial segment.
(A) Weak Paralytic (Para) expression can be detected on paraApex2 expressing small axons (arrowheads) running in fascicles within the nerve. (B–D) Cross sections through the same axon at various positions. Distance between individual sections (B,C) is 15 µm, distance between (C,D) is 6.5 µm. Note the intense labeling of the axonal membrane is changing between the different sections. (E) Cross section, to determine the staining intensity along the membrane (below the blue line), a corresponding ROI was defined and (F) quantified using Fiji. (G) Surface plot of ParaApex2 distribution along 16 consecutive axonal cross sections. For details, see Materials and methods. The intensity of diaminobenzidine (DAB) precipitates is transformed to different colors. Note that Para clusters are organized in two longitudinal lines across the axonal membrane surface. (H) Longitudinal section of a paraApex2 expressing axon. The staining intensity along the membrane (above the blue line) was quantified using Fiji. (I) Staining intensity of the membrane stretch shown in (H). Note the regular increase in staining intensity every 0.6–0.8 µm. For quantification see Figure 3G. Scale bars are as indicated.
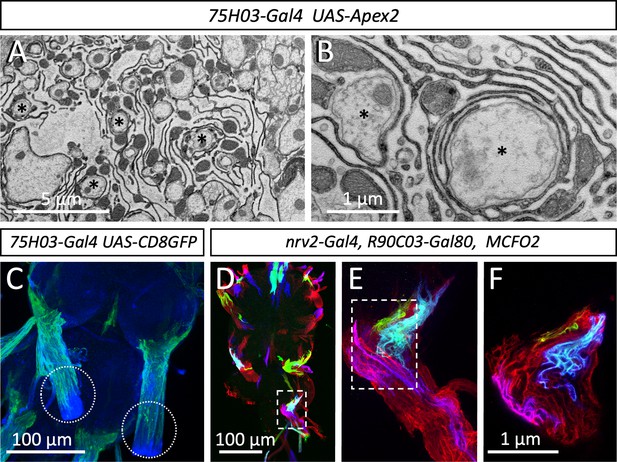
Organization of the lacuna forming tract glial.
(A,B) Apex2 expression directed by 75H03-Gal4. Axons (asterisks) are engulfed by lacunar structures that are largely formed by the tract glia. (C) Maximum projection of a confocal image stack. 75H03-Gal4 directed expression of GFP labels the ensheathing/wrapping or tract glia. Note that GFP expression ends proximal to the dissection cut (white dashed circles). (D–F) multicolor flipout (MCFO2) analysis of the nrv2-Gal4, R90C03-Gal80 positive wrapping glia. Note that glial cells tile the nerve roots with no gaps in between. Scale bars are as indicated.
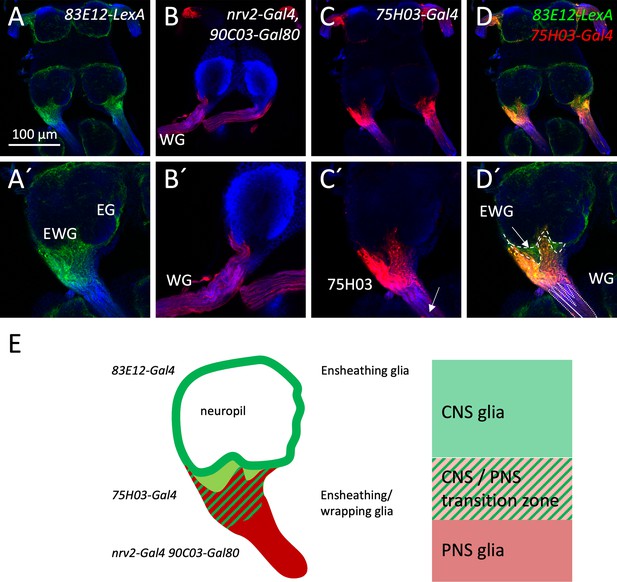
Glial cell types involved in lacuna formation.
(A–C) The tract glial cells as defined by 75H03-Gal4 UAS-tdTomato activity also express the CNS ensheathing glia marker 83E12-LexA LexAop-CD8::GFP. (D) The PNS wrapping glia marker nrv2-Gal4 90C03-Gal80 UAS-mCherry labels cells that overlap in their expression domain with the tract glial cells. HRP (blue) labels neuronal membranes. Scale bar is 100 µm. (E) Schematic summary of central and peripheral wrapping glial cells in Drosophila. The neuropil is covered by the ensheathing glia. The peripheral axons are wrapped by the peripheral wrapping glia. The 75H03-Gal4 positive glial cells are located in between these two glial cell populations.
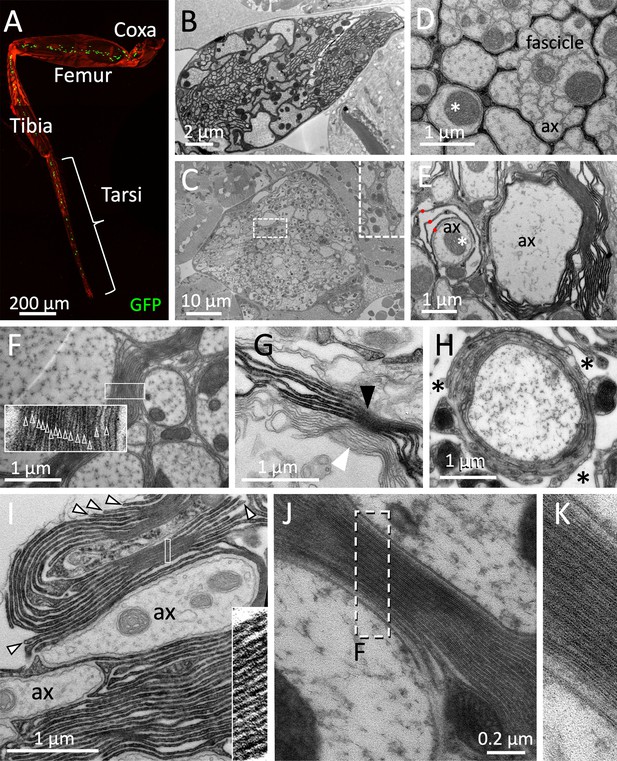
Drosophila wrapping glia form myelin.
(A) Drosophila leg of a 3-week-old fly with wrapping glial nuclei in green, the cuticle is stained by autofluorescence, the genotype is [nrv2-Gal4, UAS-lamGFP]. (B–K) Electron microscopic images of sections taken from 3-week-old female flies. (B) Section at the level of the femur. (C) Electron microscopic section at the level of the coxa. In some areas, an increased amount of glial membranes can be detected close to large caliber axons (box with white dashed lines, enlarged as an inlay). (D,E,G) Cross sections through a 2 weeks adult leg of a fly with the genotype [75H03-Gal4, UAS-Myr-Flag-Apex2-NES]. Glial cell processes are stained by the presence of Apex2 which generates an osmiophilic diaminobenzidine (DAB) precipitate. (D) Small caliber axons (ax) are engulfed by a single glial process as fascicle. Larger axons are individually wrapped (asterisk). (E) Large caliber axons are surrounded by glial membrane stacks. The asterisk denotes an axon engulfed by a few glial wraps (red dots). (F) Up to 15 densely packed membrane sheets are found (see inlay for enlargement). (G) Darkly stained tract glia membrane stacks (black arrowhead) can be found next to unlabeled membrane stacks (white arrowhead), suggesting that myelin-like structures can be derived from both, central and peripheral wrapping glial cells. (H) High-pressure freezing preparation showing a single axon covered by myelin-like membrane sheets in a lacunar area (asterisks). (I) Note the bulged appearance of the growing tip of the glial cell processes that form the myelin-like structures (arrowheads). The inlay shows a highly organized membrane stacking. (J,K) High-pressure freezing preparation of prefixed samples to reduce tissue preparation artifacts. Note the compact formation of membrane layers. The white dashed area is shown in (K). Scale bars are as indicated.
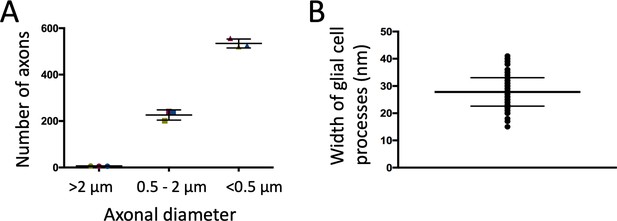
Count of axons in Drosophila leg nerves and measurement of glial cell process width.
(A) About 760 axons innervate the leg. The majority is smaller than 0.5 μm in diameter, very few ones are larger than 2 μm. (B) The width of glial cell processes is about 28 nm and very regular.
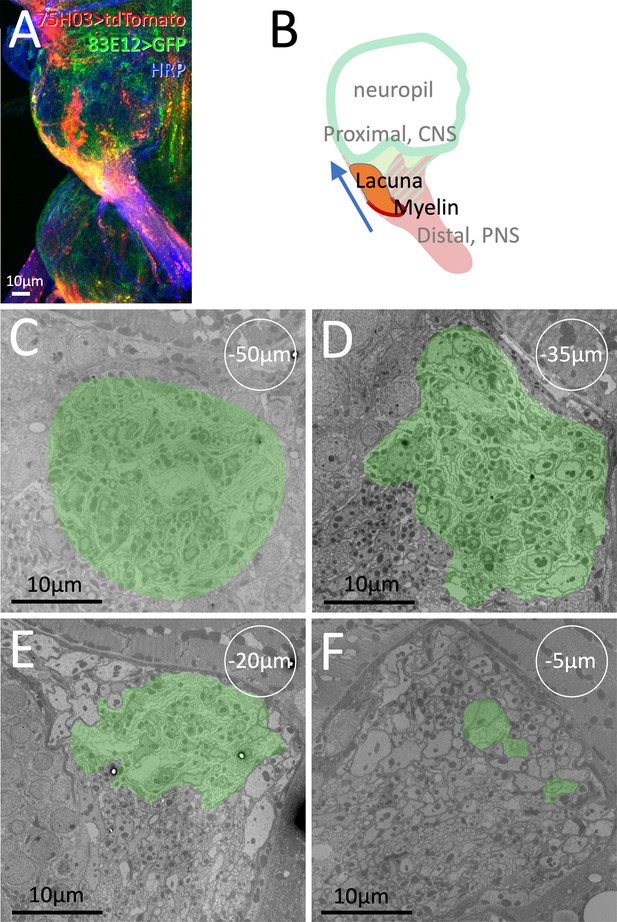
Extent of the lacunar system.
(A) CNS/PNS boundary of the mesothoracic neuromere of an adult ventral nerve cord with the genotype [75H03-Gal4, UAS-tdTomato, 83E12-lexA, lexAop-CD8GFP] imaged for tdTomato, GFP, and HRP expression. (B) Schematic representation of the image shown in (A) with the position of the lacunar region indicated. (C–F) Examples of a serial section series taken every 5 µm for 40–65 µm. The green shading indicates the lacunar region. The numbers in circles show relative distances to the first distal section with lacunar structures. Scale bars are as indicated.
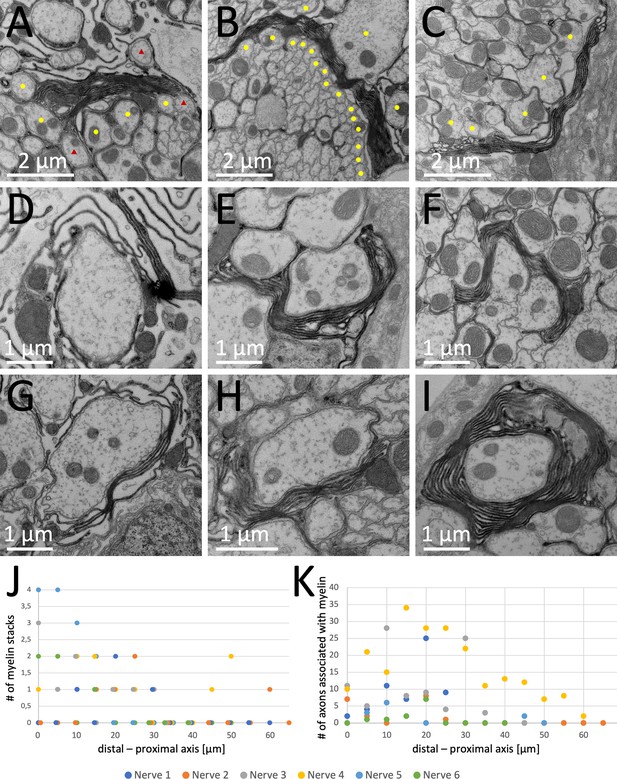
Quantification of myelin distribution in the leg nerve.
(A–I) Examples of myelin-like structures of the leg nerve. (A) Myelin-like sheets can be found separating the lacunar region from small caliber axons. (B) Myelin-like sheets separate large caliber axons from small caliber axons. (C) Myelin-like sheets can be found toward the blood-brain barrier. (D) Myelin-like sheets are rarely found in the lacunar region without close contact to axons. (E,F) Myelin-like sheets can partially wrap larger axons. (G–I) Myelin-like sheets can be found with different complexity around single large caliber axons. (J) Quantification of the number of myelin-like stacks detected in a specific section plane (see Figure 3). The value set as 0 corresponds to the distal most point where lacunar structures were detected. Progression of sections is toward the CNS (proximal). (K) Quantification of the number of axons contacting the myelin-like stacks.
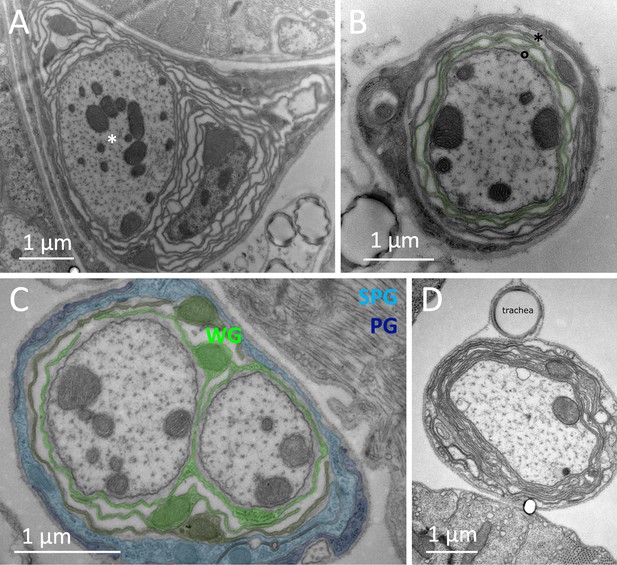
Multilayered myelin-like structures are formed around single axons in the adult nervous system.
(A) Loosely wrapped glial membranes around one single axon (asterisk). The spacing of the glial membranes resembles the glial lacunae. (B) Wrapping around a single axon. The green shaded glial cell process wraps spirally around the central axon. The ends are denoted by the asterisk and the circle. (C) Simple wrapping around single axons. The shading indicates the different glial cell types present in the nerve: wrapping glia, WG; perineurial glia, PG; subperineurial glia SPG. (D) Tight wrapping around a single axon. Unlike the image shown in (A) a close apposition of glial membranes is noted. Scale bars are as indicated.
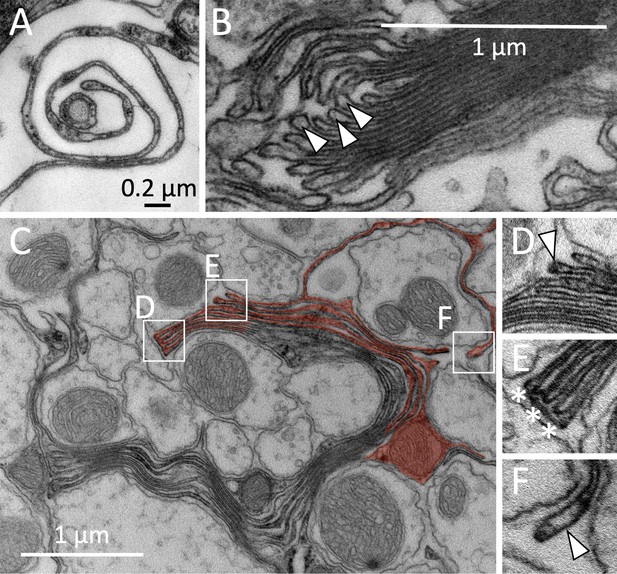
Formation of myelin-like structures in the adult CNS of Drosophila.
(A) High-pressure freezing preparation. Spiral growth of a glial cell process. (B) Membrane stack formed by a wrapping glial cell. Note the bulbed growing tips of the glial membrane sheets (arrowheads). (C–F) Myelin-like membrane sheets can be connected by comb-like structures. (C) Overview of a multilayered membrane stack around several axons, red shading highlights some of the glial membrane sheets. The arrowhead indicates a bulb structure at the end of the glial membrane sheet (D). In some cases, the ends of the membrane sheets are connected by comb-like structures (asterisks) (E). Growing tip of a wrapping glial cell process that navigated around an axon (ax) (F). Scale bars are as indicated.
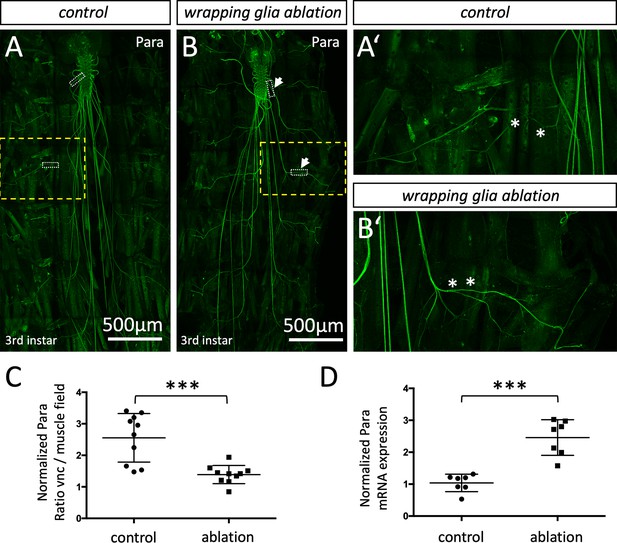
Localization of the voltage-gated sodium channel depends on glia.
(A,A’) Third instar larval filet preparation with the genotype [nrv2-Gal4, UAS-CD8-GFP; R90C03-Gal80] showing the localization of Paralytic (Para) as detected using the anti-Para antibody in a control larva. (B,B’) Third instar larval filet preparation with the genotype [nrv2-Gal4, UAS-hid; R90C03-Gal80] showing the localization of Para as detected using the anti-Para antibody in a wrapping glia ablated larva. The white dashed boxes were used for quantification of Para fluorescence intensity in the CNS/PNS transition zone in relation to its expression in the muscle field area. The yellow boxed areas are shown in higher magnification (A’,B’). Note the increased localization of Para along the peripheral nerve at the level of the muscle field (asterisks). Scale bars are as indicated. (C) Quantification of Para fluorescence intensity in the CNS/PNS transition area and the muscle field area in control and wrapping glia ablated larvae (n=10 larval filets, 3 nerves/filet). To exclude a possible influence seen in individual animals, the average fluorescence intensities along nerves of each individual were compared. Note, Para distributes more evenly along the axon in the absence of wrapping glia (p=0,0003; Mann-Whitney U-test). (D) Quantification of para mRNA expression using qRT-PCR in control and wrapping glia ablated larvae (n=7, with 15–20 brains each). para ct-values were normalized to ct-values of control gene, RPL32. Note, the significant increase in para mRNA expression upon wrapping glia ablation (p=0,0006, Mann-Whitney U-test). Scale bars are as indicated.
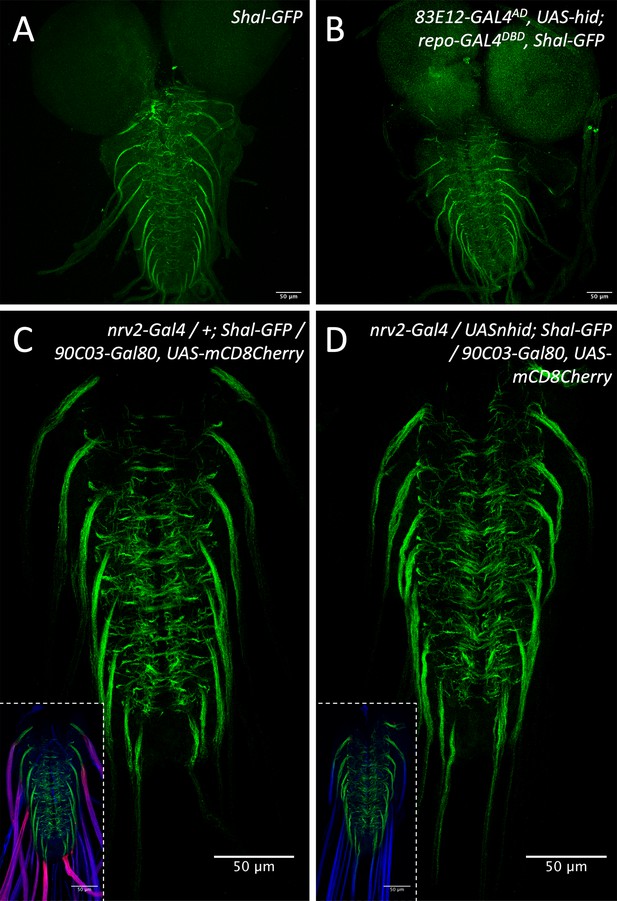
Glia ablation does not affect the localization of the voltage-gated potassium channel Shal.
(A) Control larva with an endogenously tagged Shal potassium channel. Shal predominantly localizes to the axon initial segment. (B) Upon ablation of the ensheathing glia, no change in Shal localization is detected. (C) Control larva. The inlay shows co-staining for wrapping glial cell processes (magenta) and HRP to detect neuronal membranes. (D) Upon ablation of wrapping glia, no change in Shal localization is detected. Scale bars are as indicated.
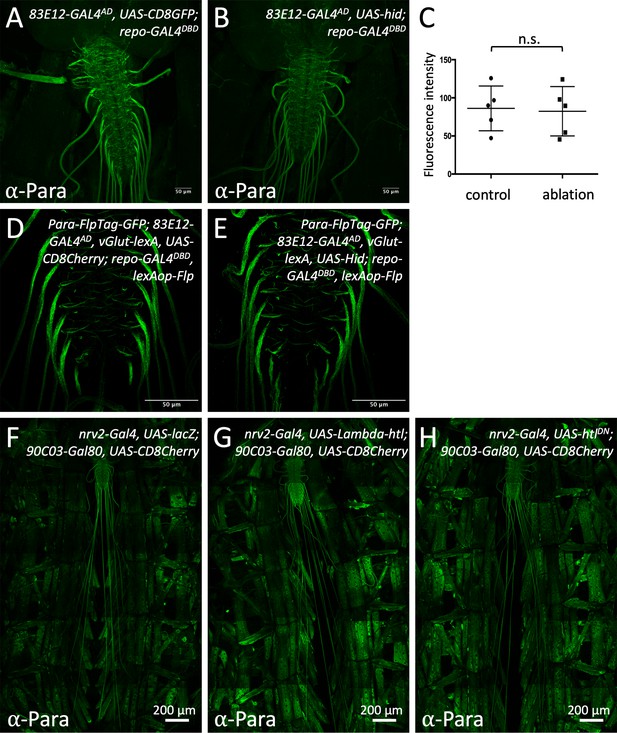
Ablation of central ensheathing glia does not affect positioning of Paralytic (Para) at the axon initial segment (AIS).
CNS preparations of third instar larvae of the genotypes indicated are shown. (A) Control larva, expressing CD8GFP under the control of the split Gal4 driver [83E12-Gal4AD, repo-Gal4DBD, UAS-CD8GFP] specific for ensheathing glial cells stained for Para protein expression. (B) Upon ablation of the ensheathing glia following expression of the proapoptotic gene hid, no change in the Para expression levels is detected. (C) Quantification of the Para fluorescence intensity in control and ensheathing glia ablated larvae (n=5 larval brains, 10–16 nerves/brain). To exclude influence of individual animals 10–16 nerves per individual were measured and analysis was performed on the mean fluorescence intensity of all values from one animal (p=0,8016, Mann-Whitney U-test). (D) Control larvae for ensheathing glia ablation using the FlpTag approach. The GFP encoding exon was flipped in all motor neurons using [vGlut-lexA, lexAop-Flp]. Note, the pronounced localization of ParaGFP at the AIS-like domain of the nerve. (E) Upon ablation of the ensheathing glial cells, no change in Para localization in motor axons can be detected. (F–H) Filet preparations of third instar larvae stained for Para localization. Control larva (F). Upon expression of activated FGF-receptor Heartless no change in Para localization is noted (G). Upon expression of dominant negative Heartless, no change in Para localization is noted (H). Scale bars are as indicated.
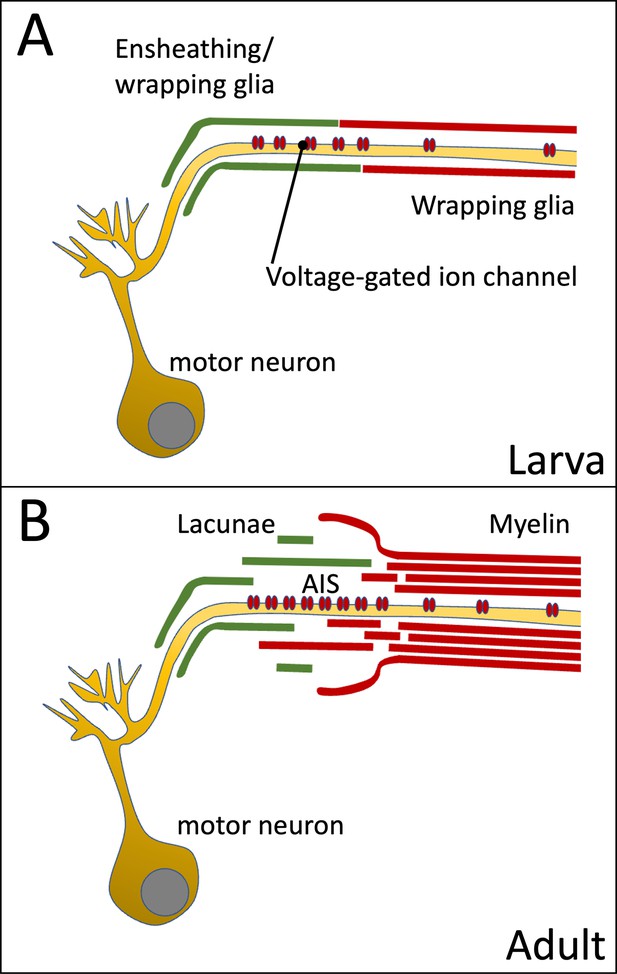
Organization of the axon initial segment (AIS) in Drosophila motor axons.
Voltage-gated sodium channels are preferentially positioned at the AIS of the motor axon. (A) In the larval nervous system positioning is mediated by the peripheral wrapping glia. (B) In adults these cells form myelin-like structures, which fray out in the lacunae which represent a reservoir possibly needed for ion homeostasis during sustained action potential generation.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Drosophila melanogaster) | y[1] w[*] Mi{MIC}MI08578a Mi{MIC}MI08578b | Bloomington Drosophila Stock Center | BDSC51087 | |
| Genetic reagent (Drosophila melanogaster) | y[1] w[*] Mi{FlpStop}para[MI08578-FlpStop.D]/FM7c | Bloomington Drosophila Stock Center | BDSC67680 | |
| Genetic reagent (Drosophila melanogaster) | Para-mCherry | This study | Figures 1, 2 | |
| Genetic reagent (Drosophila melanogaster) | Para-Apex | This study | Figures 1, 5 | |
| Genetic reagent (Drosophila melanogaster) | Para-FlpTag GFP/Fm7i | Fendl et al., 2020 | ||
| Genetic reagent (Drosophila melanogaster) | y[1] w[*]; Mi{PT-GFSTF.1}Shal[MI00446-GFSTF.1] | Bloomington Drosophila Stock Center | BDSC60149 | |
| Genetic reagent (Drosophila melanogaster) | y[1] w[*] Mi{PT-GFSTF.2}Sh[MI10885-GFSTF.2]/FM7j, B[1] | Bloomington Drosophila Stock Center | BDSC59423 | |
| Genetic reagent (Drosophila melanogaster) | y[1] w[*]; Mi{PT-GFSTF.1}Shab[MI00848-GFSTF.1]/TM6C, Sb[1] Tb[1] | Bloomington Drosophila Stock Center | BDSC60514 | |
| Genetic reagent (Drosophila melanogaster) | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=VALIUM20-mCherry}attP2 | Bloomington Drosophila Stock Center | BDSC35785 | |
| Genetic reagent (Drosophila melanogaster) | UAS-CD8GFP II; R90C03Gal80 III | Kottmeier et al., 2020 | ||
| Genetic reagent (Drosophila melanogaster) | UAS-Hid/CyOw; R90C03Gal80 III | Kottmeier et al., 2020 | ||
| Genetic reagent (Drosophila melanogaster) | UAS-lacZ NLS II | Y. Hirmoi | ||
| Genetic reagent (Drosophila melanogaster) | UAS-lambda-Htl | Gisselbrecht et al., 1996 | ||
| Genetic reagent (Drosophila melanogaster) | UAS-htlDN II | Bloomington Drosophila Stock Center | BDSC5366 | |
| Genetic reagent (Drosophila melanogaster) | UAS-CD8GFP II | Bloomington Drosophila Stock Center | BDSC5137 | |
| Genetic reagent (Drosophila melanogaster) | UAS-CD8mCherry II | Bloomington Drosophila Stock Center | BDSC 27391 | |
| Genetic reagent (Drosophila melanogaster) | UAS-Hid II | Igaki et al., 2000 | ||
| Genetic reagent (Drosophila melanogaster) | UAS-Flp | Bloomington Drosophila Stock Center | BDSC 4539 | |
| Genetic reagent (Drosophila melanogaster) | UAS-Myr-Flag-APEX2-NES86Fb III | This study | Figures 6, 7 | |
| Genetic reagent (Drosophila melanogaster) | pBPhsFLP2::pEST/I;; UAS HA, FLAG, V5, OLLAS/III | Nern et al., 2015 | ||
| Genetic reagent (Drosophila melanogaster) | ChAT-Gal4 | Salvaterra and Kitamoto, 2001 | ||
| Genetic reagent (Drosophila melanogaster) | OK371-Gal4 | Mahr and Aberle, 2006 | ||
| Genetic reagent (Drosophila melanogaster) | GMR94G06Gal4 III | Bloomington Drosophila Stock Center | BDSC40701 | |
| Genetic reagent (Drosophila melanogaster) | Nrv2Gal4 II | Sun et al., 1999 | ||
| Genetic reagent (Drosophila melanogaster) | Nrv2Gal4 II; R90C03Gal80 III | Kottmeier et al., 2020 | ||
| Genetic reagent (Drosophila melanogaster) | Nrv2Gal4 II; R90C03Gal80, UAS-CD8Cherry III | Kottmeier et al., 2020 | ||
| Genetic reagent (Drosophila melanogaster) | GMR83E12_AD II; Repo4.3_DBD III | Bittern et al., 2021 | ||
| Genetic reagent (Drosophila melanogaster) | GMR75H03-Gal4 III | Bloomington Drosophila Stock Center | BDSC39908 | |
| Genetic reagent (Drosophila melanogaster) | ppkGal4, AUS-tdTomato III | Herzmann et al., 2017 | ||
| Genetic reagent (Drosophila melanogaster) | lexAop-Flp III | Bloomington Drosophila Stock Center | BDSC55819 | |
| Genetic reagent (Drosophila melanogaster) | w[*]; TI{2A-lexA::GAD}VGlut[2A-lexA]/CyO | Bloomington Drosophila Stock Center | BDSC84442 | |
| Genetic reagent (Drosophila melanogaster) | ParaST76 | Bloomington Drosophila Stock Center | BDSC26701 | |
| Genetic reagent (Drosophila melanogaster) | Oregon R | Bloomington Drosophila Stock Center | BDSC5 | |
| Antibody | Anti-Para N-term (rabbit, polyclonal) | This study | IF (1:1000), WB (1:1000) Figures 1, 2 | |
| Antibody | Anti-dsRed (rabbit, polyclonal) | Takara | Cat#632496 RRID:AB_10013483 | IF (1:1000) |
| Antibody | Anti-GFP (chicken, polyclonal) | Abcam | Cat#ab13970 RRID:AB_300798 | IF(1:500) |
| Antibody | Anti-GFP (rabbit, polyclonal) | Invitrogen | Cat#A-11122 RRID:AB_221569 | IF(1:1000) |
| Antibody | Anti-Rumpel (rabbit, polyclonal) | Yildirim et al., 2022 | IF(1:1000) | |
| Antibody | Anti-Repo (mouse, monoclonal) | Hybridoma Bank | Cat#8D12 RRID: AB_528448 | IF(1:5) |
| Antibody | Anti-V5 (rabbit, polyclonal) | Sigma-Aldrich | Cat#V8137-.2MG RRID:AB_261889 | IF(1:500) |
| Antibody | Anti-HA (mouse, monoclonal) | Covance BioLegend | Cat#MMS-101R RRID:AB_291262 | IF(1:1000) |
| Antibody | Anti-Flag (rat, monoclonal) | Novus Biologicals | Cat#NBP1-06712SS RRID:AB_162598 | IF(1:200) |
| Antibody | FluoTag-X4 anti-GFP (Alpaca, monoclonal) | NanoTag Biotechnologies | Cat#N0304 RRID:AB_2905516 | IF(1:500) |
| Antibody | FluoTag-X4 anti-RFP (Alpaca, monoclonal) | NanoTag Biotechnologies | Cat#N0404 RRID:AB_2744638 | IF(1:500) |
| Antibody | Anti-rabbit Alexa 488 (goat, polyclonal) | Thermo Fisher | Cat#A-11008 RRID:AB_143165 | IF(1:1000) |
| Antibody | Anti-rabbit Alexa 568 (goat, polyclonal) | Thermo Fisher | Cat#A-11011 RRID:AB_143157 | IF(1:1000) |
| Antibody | Anti-chicken Alexa 488 (goat, polyclonal) | Thermo Fisher | Cat#A-11039, RRID:AB_2534096 | IF(1:1000) |
| Antibody | Anti-mouse Alexa 488 (goat, polyclonal) | Thermo Fisher | Cat#A-11001, RRID:AB_2534069 | IF(1:1000) |
| Antibody | Anti-HRP Alexa 647 (goat, polyclonal) | Thermo Fisher | Cat#123-605-021, RRID:AB_2338967 | IF(1:500) |
| Antibody | Anti-rabbit HRP (goat, polyclonal) | Invitrogen | Cat#31460 RRID:AB_228341 | WB(1:5000) |
| Recombinant DNA reagent | pBS-KS-attB1-2-PT-SA-SD-0-mCherry | Drosophila Genome Research Centre | DGRC#1299 | |
| Recombinant DNA reagent | pBS-KS-attB1-2-PT-SA-SD-0-Apex2 | This study | Generation of transgenic fly | |
| Sequence-based reagent | BamH1 Apex2 fwd | This study | PCR primer | AAGGATCCGGAAAGTCTTACCCAACTGT |
| Sequence-based reagent | BamH1 Apex2 rev | This study | PCR primer | AAGGATCCGGCATCAGCAAACCCAAG |
| Sequence-based reagent | MiLF | Venken et al., 2011 | PCR primer | GCGTAAGCTACCTTAATCTCAAGAAGAG |
| Sequence-based reagent | MiLR | Venken et al., 2011 | PCR primer | CGCGGCGTAATGTGATTTACTATCATAC |
| Sequence-based reagent | mCherry-Seq fwd | PCR primer | ACGGCGAGTTCATCTACAAG | |
| Sequence-based reagent | mCherry-Seq rev | PCR primer | TTCAGCCTCTGCTTGATCTC | |
| Sequence-based reagent | Apex253_rev1 | This study | PCR primer | AGCTCAAAATAGGGAACTCCG |
| Sequence-based reagent | Apex286_fwd1 | This study | PCR primer | TACCAGTTGGCTGGCGTTGTT |
| Sequence-based reagent | Para qPCR Primer | Thermo Fisher Scientific | Cat#4331182 Dm01813740_m1 | |
| Sequence-based reagent | RPL32 qPCR Primer | Thermo Fisher Scientific | Cat#4331182 Dm02151827_g1 | |
| Commercial assay or kit | RNeasy Kit | QIAGEN | Cat#74104 | |
| Commercial assay or kit | Quantitect Reverse Transcription Kit | QIAGEN | Cat#205313 | |
| Commercial assay or kit | Taqman gene expression assay, Universal Master Mix II, with UNG | Thermo Fisher Scientific | Cat#4440038 | |
| Software algorithm | GraphPad PRISM | GraphPad Software, USA | Version 6.0 | |
| Software algorithm | Fiji | https://imagej.net/software/fiji/ | ||
| Software algorithm | ZEN Software | Zeiss | Black version | |
| Software algorithm | Affinity Photo | Serif (Europe) | ||
| Software algorithm | MATLAB | The MathWorks, Inc | ||
| Software algorithm | Photoshop CS6 | Adobe |





