Enhancing CRISPR prime editing by reducing misfolded pegRNA interactions
Figures
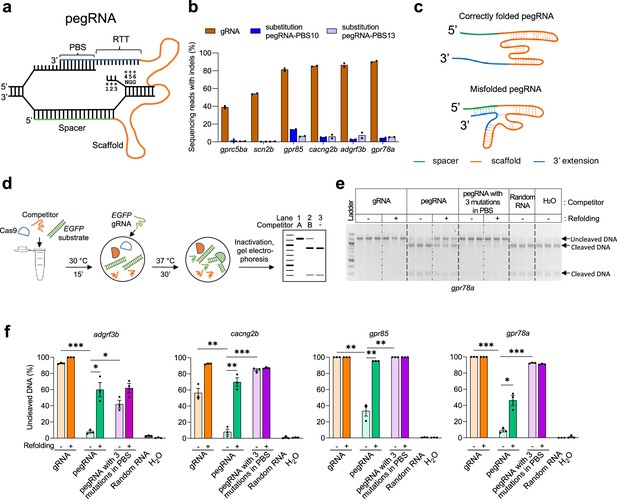
Improving in vitro SpCas9 binding efficiencies of pegRNA by refolding.
(a) Schematic illustrating the hybridization of a pegRNA and its target DNA. Four segments of a pegRNA are shown. PBS, Primer Binding Site; RTT, Reverse Transcriptase Template. Target DNA positions (as well as the corresponding sequences in the RTT) are numbered counting from the SpCas9-induced nick towards ‘NGG’, the protospacer adjacent motif (PAM). (b) Mutation frequencies induced by SpCas9 with gRNAs and pegRNAs in zebrafish. All pegRNAs carried a single nucleotide substitution at position +5 or+6, with RTT lengths of 14- or 15-nucleotide (nt), and PBS lengths of 10-nt (pegRNA-PBS10) or 13-nt (pegRNA-PBS13). Target loci are indicated at the bottom. SpCas9 protein was complexed with gRNA or pegRNA at a molar ratio of 1:2 (0.6 µM of gRNA or pegRNA). (c) Schematic illustrating hypothetical conformations of correctly folded and misfolded pegRNAs. The spacer is shown in green, Cas9-binding scaffold in orange and 3’ extension including PBS and RTT in blue. Dotted lines indicate potential base pairings. (d) Schematic illustrating the in vitro competition assay for Cas9 binding and substrate cleavage. Possible outcomes of the assay are shown in a representative gel. Lane 1 shows the addition of Competitor A with a high SpCas9-binding affinity resulting in 100% inhibition of cleavage of DNA substrates (1.2 kilobase pairs). Lane 2 shows the addition of Competitor B with a low SpCas9-binding affinity yielding a mix of uncleaved and cleaved (900 and 300 base pairs) DNA substrate. Lane 3 shows the reaction without any competitor resulting in 100% cleaved DNA products. (e) Agarose gel image showing the results of in vitro SpCas9 cleavage of DNA substrate in the presence of gRNA or pegRNAs targeting gpr78a as competitors, with or without refolding (indicated on top of the gel). Random RNA isolated from tolura yeast was used as a negative control. Assays were performed in triplicate. (f) Percentage of uncleaved DNA substrate in the presence or absence of competitor gRNA or pegRNA calculated using data from Figure 1—source data 1 and Figure 1—source data 2. Competitor gRNA and pegRNA target loci are indicated at the top. Competitor types are shown at the bottom. Dots represent individual data points, bars the mean and error bars ± s.e.m. Unpaired two-tailed t-test with equal variance was used to compare non-refolded gRNA vs non-refolded pegRNA, non-refolded pegRNA vs non-refolded pegRNA with three mutations in PBS, and non-refolded vs refolded pegRNAs. *p<0.05, **p<0.01, ***p<0.001.
-
Figure 1—source data 1
Original agarose gel images showing in vitro SpCas9 cleavage of EGFP DNA substrate in the presence of gRNAs or pegRNAs targeting gpr78 (a), adgrf3b (b), cacng2b (c), and gpr85 (d).
- https://cdn.elifesciences.org/articles/90948/elife-90948-fig1-data1-v1.pdf
-
Figure 1—source data 2
Labeled agarose gel images showing in vitro SpCas9 cleavage of EGFP DNA substrate in the presence of gRNAs or pegRNAs targeting adgrf3b (a), cacng2b (b), and gpr85 (c).
The percentage of uncleaved DNA substrate in the presence or absence of competitor gRNA or pegRNA calculated using this data is shown in Figure 1f.
- https://cdn.elifesciences.org/articles/90948/elife-90948-fig1-data2-v1.pdf
-
Figure 1—source data 3
pegRNA with/without triple mutations in PBS for in vitro assay.
The RT template (RTT) sequences are shown in red. The PBS sequences are underlined. The three mutations in PBS are shown in orange.
- https://cdn.elifesciences.org/articles/90948/elife-90948-fig1-data3-v1.pdf
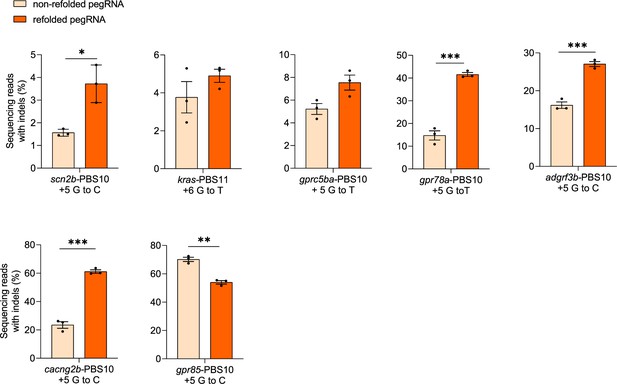
Indel frequencies of non-refolded and refolded pegRNAs with SpCas9 in zebrafish.
Indel frequencies in zebrafish induced by SpCas9 complexed with non-refolded or refolded pegRNAs at a molar ratio of 1:2 (1.8 µM of pegRNA). Target loci, PBS lengths, and pegRNA-specified edits are indicated at the bottom. Dots represent individual data points (n=3 biologically independent replicates, 5–10 embryos per replicate), bars the mean and error bars ± s.e.m. Results of unpaired two-tailed t-test with equal variance are shown in *p<0.05, **p<0.01, ***p<0.001.
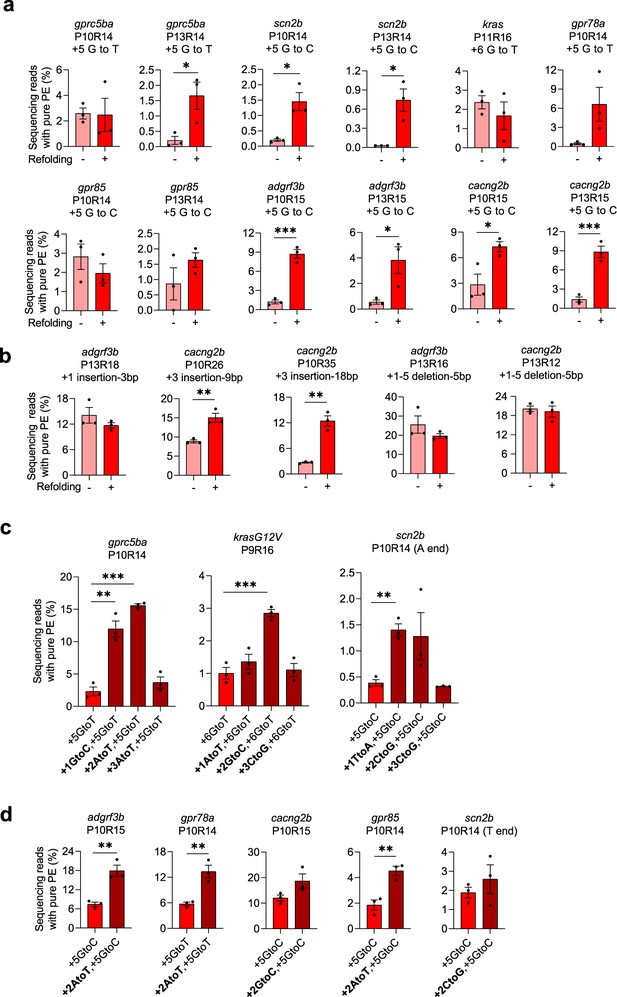
Improving prime editing efficiencies in zebrafish by pegRNA refolding and mutations in RTT.
(a–b) Pure PE frequencies of non-refolded and refolded substitution pegRNAs (a) and insertion or deletion pegRNAs (b) with PE2 in zebrafish. Target loci, PBS lengths (labeled as ‘P’ followed by the number of nucleotides), RTT lengths (labeled as ‘R’ followed by the number of nucleotides), and pegRNA-specified edits (denoted as the position of the edit followed by the edit) are shown at the top. Pure PE represents sequencing reads containing only the pegRNA-specified mutations. (c–d) Pure PE frequencies with refolded pegRNAs carrying additional RTT mutations (at +1,+2 or+3) and PE2 in zebrafish. Target loci, PBS, and RTT lengths are shown at the top and pegRNA-specified edits are shown at the bottom. All pegRNAs had 3 or 4 thymine (T) nucleotides at the 3’ end except for the ones labeled ‘A end’ for scn2b in which the terminal Ts were replaced with adenine (A) nucleotides. Dots represent individual data points (n=3 biologically independent replicates, 5–10 embryos per replicate), bars the mean and error bars ± s.e.m. *p<0.05, **p<0.01, ***p<0.001 (unpaired two-tailed t-test with equal variance).
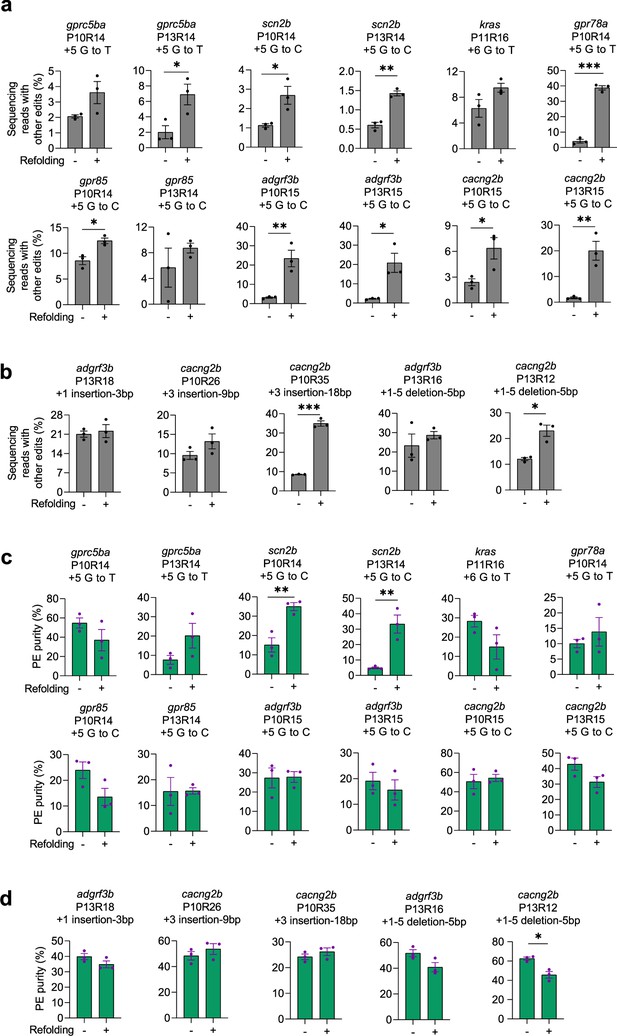
Purities of prime editing with PE2 and non-refolded or refolded pegRNAs in zebrafish.
(a–b) Frequencies of non-pure PE edits (labeled as other edits) mediated by PE2 with non-refolded and refolded substitution pegRNAs (a) and insertion or deletion pegRNAs (b). (c–d) PE purities (%) calculated as pure PE over total edit (pure and non-pure PE) frequencies for substitution pegRNAs (c) and insertion or deletion pegRNAs (d). Target loci, PBS lengths (labeled as ‘P’ followed by the number of nucleotides), RTT lengths (labeled as ‘R’ followed by the number of nucleotides), and pegRNA-specified edits are indicated at the top. Dots represent individual data points (n=3 biologically independent replicates, 5–10 embryos per replicate), bars the mean and error bars ± s.e.m. Results of unpaired two-tailed t-test with equal variance are shown in *p<0.05, **p<0.01, ***p<0.001.
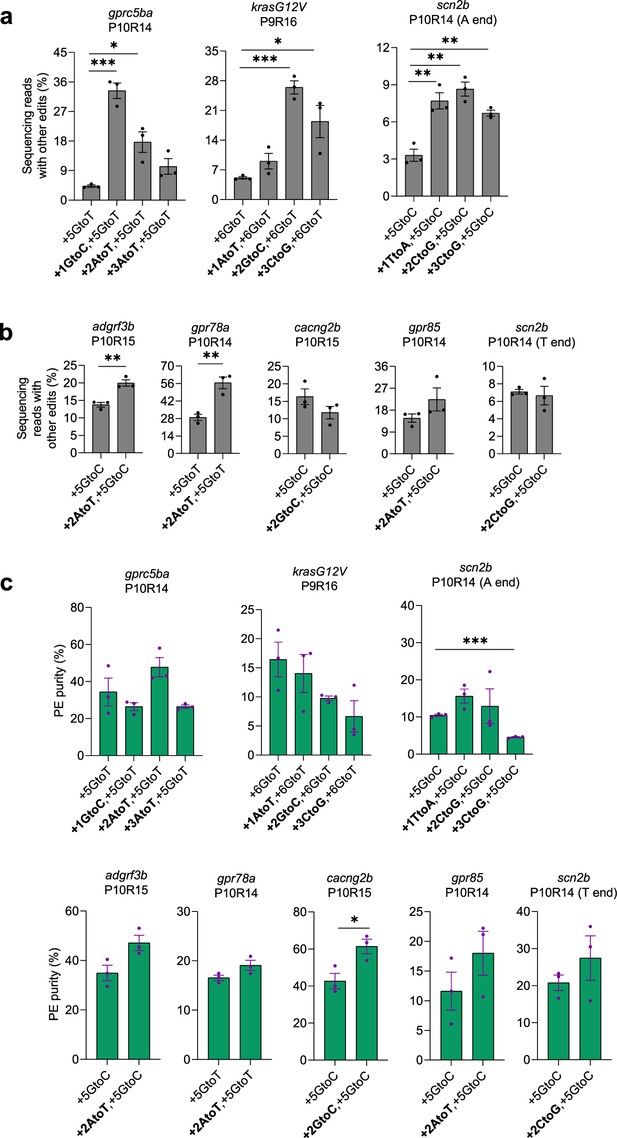
Purities of prime editing with PE2 and refolded pegRNAs carrying additional RTT mutations in zebrafish.
(a–b) Frequencies of non-pure PE edits (labeled as other edits) mediated by PE2 with refolded pegRNAs with or without an additional mutation in the RTT (at +1,+2 or+3). (c) PE purities calculated as pure PE over total edit (pure and non-pure PE) frequencies. Target loci, PBS lengths (labeled as ‘P’ followed by the number of nucleotides), and RTT lengths (labeled as ‘R’ followed by the number of nucleotides) are shown at the top. pegRNA-specified edits (denoted as the position of the edit followed by the edit) are shown at the bottom. Dots represent individual data points (n=3 biologically independent replicates, 5–10 embryos per replicate), bars the mean, error bars ± s.e.m. Results of unpaired two-tailed t-test with equal variance are shown in *p<0.05, **p<0.01, ***p<0.001.
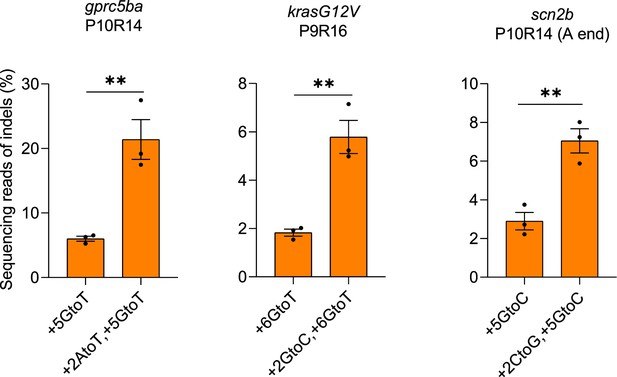
Indel frequencies in zebrafish induced by SpCas9 complexed with refolded pegRNAs with or without RTT mutation at +2 position.
SpCas9 protein and pegRNA were combined at a molar ratio of 1:2. (1.8 µM of pegRNA). Target loci, PBS lengths (labeled as ‘P’ followed by the number of nucleotides), and RTT lengths (labeled as ‘R’ followed by the number of nucleotides) are shown at the top. pegRNA-specified edits (denoted as the position of the edit followed by the edit) are shown at the bottom. Dots represent individual data points (n=3 biologically independent replicates, 5–10 embryos per replicate), bars the mean and error bars ± s.e.m. Results of unpaired two-tailed t-test with equal variance are shown in **p<0.01.
Additional files
-
Supplementary file 1
PE outcomes for pegRNAs with single and double substitutions and with or without refolding.
- https://cdn.elifesciences.org/articles/90948/elife-90948-supp1-v1.xlsx
-
Supplementary file 2
The target site, RT template, PBS and Cas9-binding scaffold sequences of pegRNAs.
- https://cdn.elifesciences.org/articles/90948/elife-90948-supp2-v1.xlsx
-
Supplementary file 3
PCR formulation and primers used in the DNA template construction for gRNAs and pegRNAs.
- https://cdn.elifesciences.org/articles/90948/elife-90948-supp3-v1.xlsx
-
Supplementary file 4
PCR primers for the generation of the EGFP DNA substrate.
- https://cdn.elifesciences.org/articles/90948/elife-90948-supp4-v1.xlsx
-
Supplementary file 5
PCR primers for deep sequencing.
- https://cdn.elifesciences.org/articles/90948/elife-90948-supp5-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/90948/elife-90948-mdarchecklist1-v1.pdf





