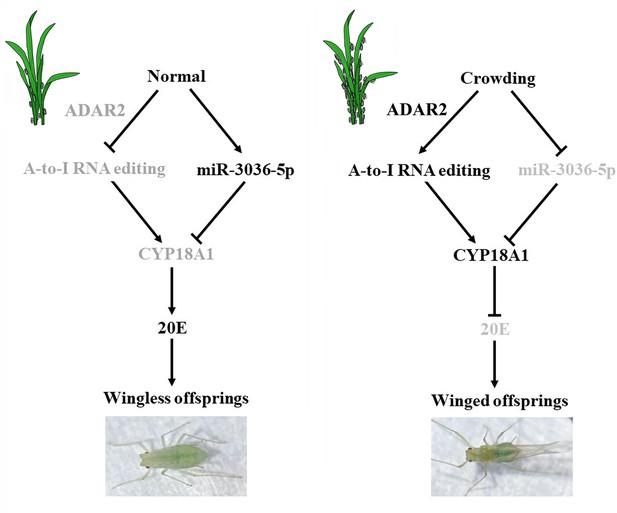
A-to-I RNA editing of CYP18A1 mediates transgenerational wing dimorphism in aphids
Figures
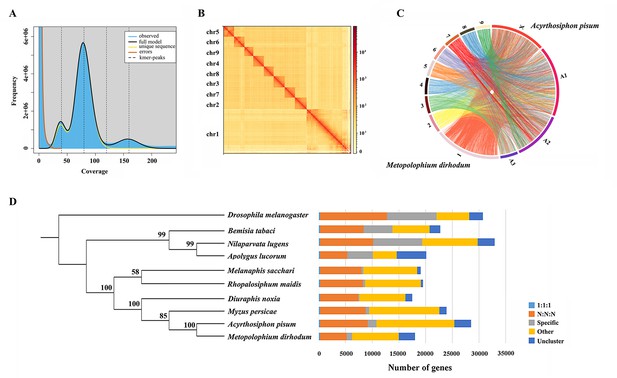
Assembled genome for Metopolophium dirhodum.
(A) k-mer (K=17) distribution of Illumina genome sequencing reads. (B) Hi-C contact heat map of the assembled genome. (C) Chromosome-level synteny analysis between M. dirhodum and Acyrthosiphon pisum. (D) Maximum likelihood phylogeny of M. dirhodum and nine other insect species based on a concatenated alignment of the conserved single copy orthologues. The histograms are subdivided to represent different categories of orthology: 1:1:1 (single copy orthologous genes in communal gene families); N:N:N (multiple copy orthologous genes in communal gene families); specific (genes from unique gene families from each species); other (genes that do not belong to any of the above mentioned orthologous categories); uncluster (genes that do not cluster to any families).
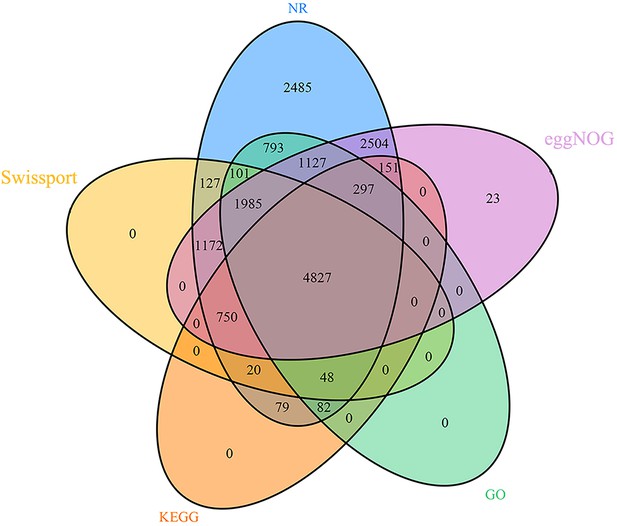
Venn diagram of functional annotation based on five databases for Metopolophium dirhodum.
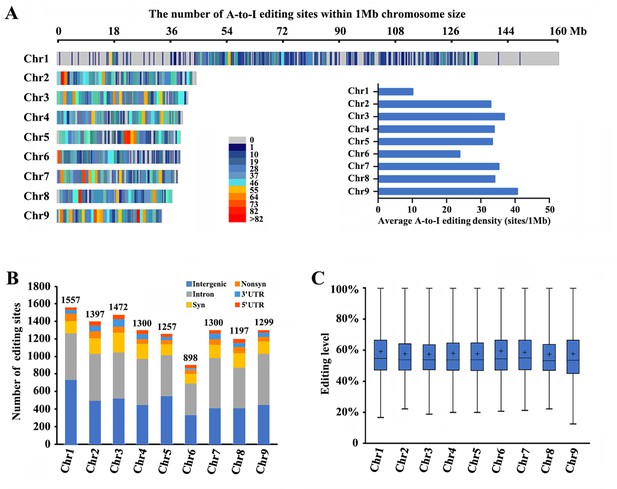
Landscape of A-to-I editomes in Metopolophium dirhodum.
(A) Density distribution map of A-to-I RNA-editing sites on nine chromosomes. (B) Number and distribution of the detected A-to-I editing sites over different genic regions. (C) Average editing levels of the detected A-to-I editing sites on nine chromosomes.
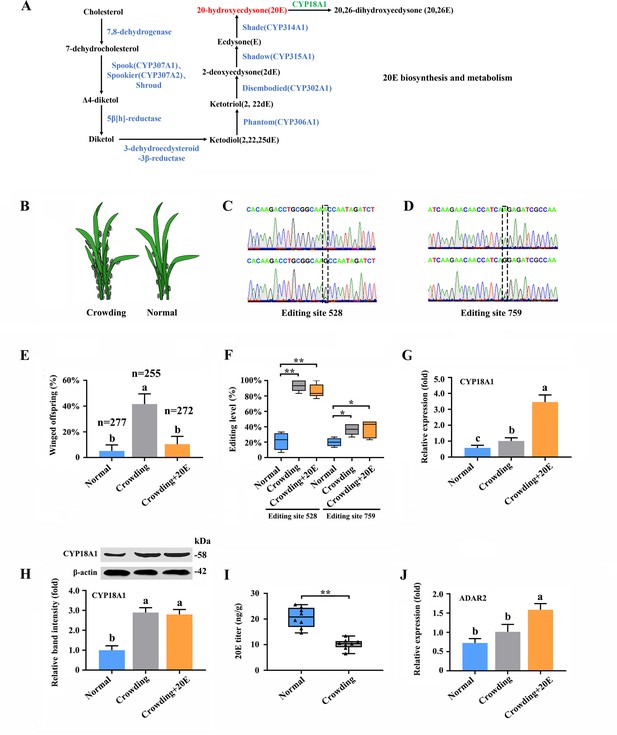
A-to-I RNA editing on CYP18A1 is linked to transgenerational wing dimorphism under crowding condition in Metopolophium dirhodum.
(A) 20-hydroxyecdysone biosynthesis and metabolism pathway. (B) Schematic diagram for normal and crowding conditions. (C, D) Representative chromatograms of the PCR product direct sequencing for the two synonymous A-to-I RNA-editing sites (editing site 528 and 759, 528th or 759th nucleotide in the CDS region of CYP18A1) on CYP18A1. (E) The proportion of editing individuals under normal, crowding, and crowding + 20E conditions. (F) The proportion of winged offspring under normal, crowding, and crowding + 20E conditions. (G, H) The expression level of CYP18A1 under normal, crowding and crowding + 20E conditions by RT-qPCR and western blot. (I) The 20E titers under normal and crowding conditions. (J) The expression level of ADAR2 under normal, crowding, and crowding + 20E conditions by RT-qPCR. Asterisks indicate significant differences between the treatment and the corresponding control (Student’s t-test, *0.01<p<0.05, **p<0.01). Different lowercase letters represent significant differences (one-way ANOVA followed by Tukey’s multiple comparison tests, p<0.05).
-
Figure 3—source data 1
Labeled file for the western blot analysis in Figure 3H (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig3-data1-v1.zip
-
Figure 3—source data 2
Uncropped file for the western blot analysis in Figure 3H (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig3-data2-v1.zip
-
Figure 3—source data 3
Labeled file for the western blot analysis in Figure 3H (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig3-data3-v1.zip
-
Figure 3—source data 4
Uncropped file for the western blot analysis in Figure 3H (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig3-data4-v1.zip
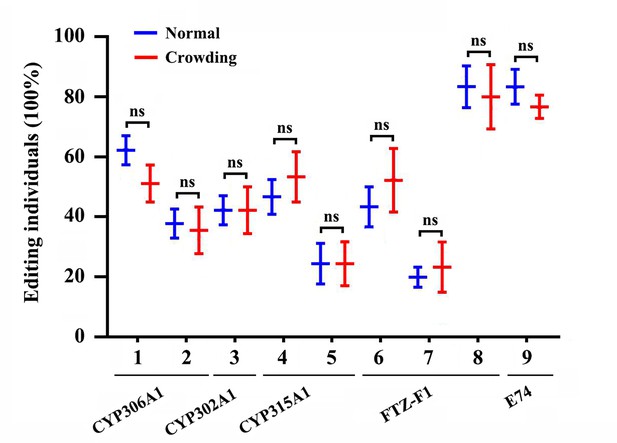
The proportion of A-to-I RNA-editing individuals for genes involved in 20E biosynthesis and metabolism under normal and crowding conditions.
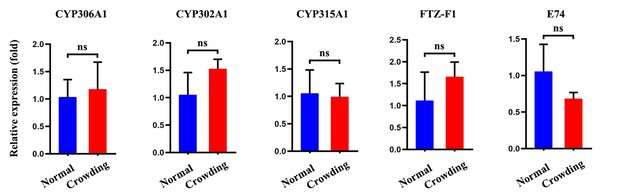
The relative expression for genes involved in 20E biosynthesis and metabolism under normal and crowding conditions.
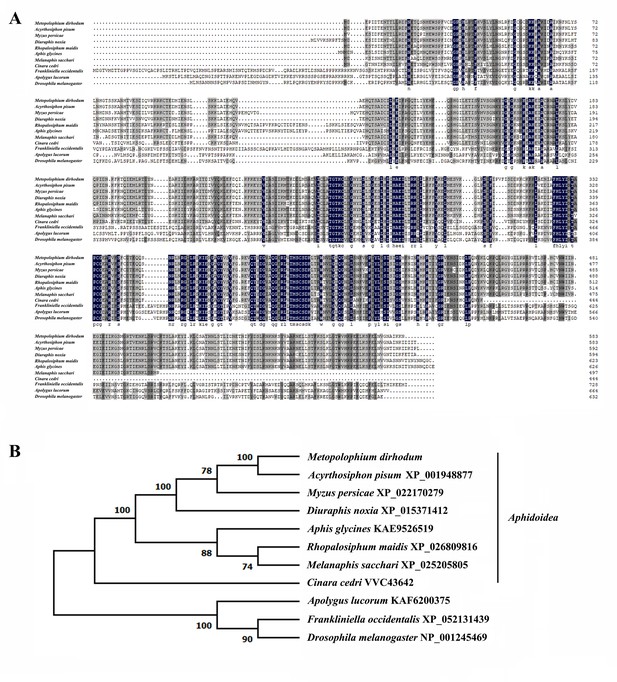
Bioinformatics analysis of the ADAR2 protein.
(A) Amino acid sequence alignment of ADAR2 homologs from Aphidoidea and other insect species. The identical and similar residues are respectively marked with black and gray backgrounds. (B) Phylogenetic tree constructed using the protein sequences of ADAR2 homologs from Aphidoidea and other insect species.
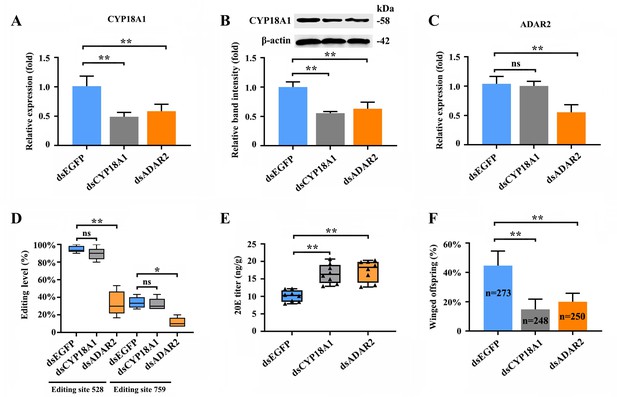
RNAi-mediated knockdown of ADAR2 could regulate the A-to-I RNA-editing level and expression of CYP18A1.
(A, B) The expression level of CYP18A1 under crowding condition treated with dsCYP18A1 and dsADAR2 after 48 h by RT-qPCR and western blot. (C) The expression level of ADAR2 under crowding condition treated with dsCYP18A1 and dsADAR2 after 48 h by RT-qPCR. (D) The proportion of editing individuals under crowding condition treated with dsCYP18A1 and dsADAR2 after 48 h. (E) The 20E titers under crowding condition treated with dsCYP18A1 and dsADAR2 after 48 h. (F) The proportion of winged offspring under crowding condition treated with dsCYP18A1 and dsADAR2 after 48 h. Asterisks indicate significant differences between the treatment and the corresponding control (Student’s t-test, *0.01<<0.05, **<0.01).
-
Figure 4—source data 1
Labeled file for the western blot analysis in Figure 4B (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig4-data1-v1.zip
-
Figure 4—source data 2
Uncropped file for the western blot analysis in Figure 4B (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig4-data2-v1.zip
-
Figure 4—source data 3
Labeled file for the western blot analysis in Figure 4B (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig4-data3-v1.zip
-
Figure 4—source data 4
Uncropped file for the western blot analysis in Figure 4B (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig4-data4-v1.zip
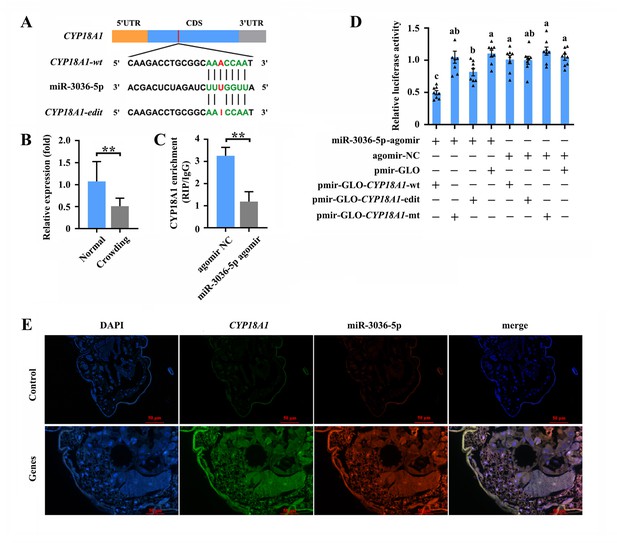
miR-3036-5p targets on CYP18A1 in Metopolophium dirhodum.
(A) The putative miR-3036-5p binding sites in CYP18A1. (B) The expression level of miR-3036-5p under normal and crowding conditions. (C) Interactions between miR-3036-5p and CYP18A1 determined by RNA-binding protein immunoprecipitation (RIP) in vivo. (D) Dual-luciferase reporter assays through co-transfection of miR-3036-5p agomir with recombinant pmirGLO vectors containing wild-type (wt), edited (edit) or mutated (mt) binding sites. (E) Co-localization of miR-3036-5p and CYP18A1 by fluorescence in situ hybridization (FISH) assay. Asterisks indicate significant differences between the treatment and the corresponding control (Student’s t-test, *0.01<p<0.05, **p<0.01). Different lowercase letters represent significant differences (one-way ANOVA followed by Tukey’s multiple comparison tests, p<0.05).
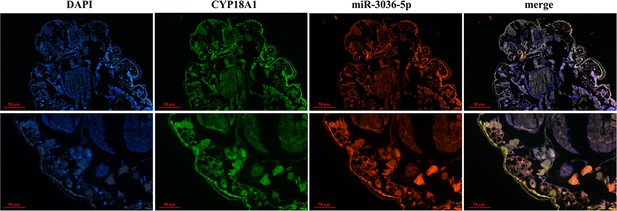
Co-localization of miR-3036-5p and CYP18A1 by fluorescence in situ hybridization (FISH) assay.
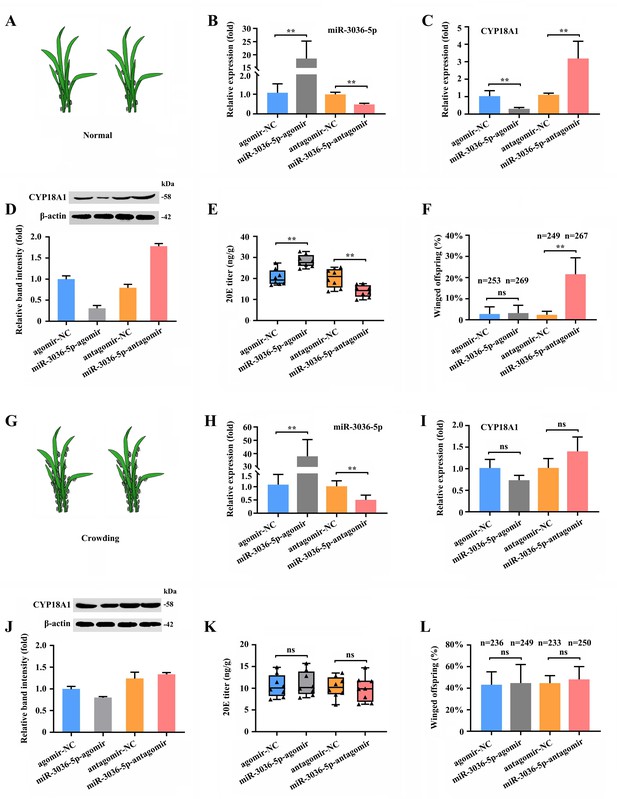
miR-3036-5p regulates transgenerational wing dimorphism by targeting CYP18A1 in Metopolophium dirhodum.
(A) Schematic diagram for normal condition. (B) The expression level of miR-3036-5p under normal condition treated with miR-3036-5p agomir or antagomir after 48 h. (C, D) The expression level of CYP18A1 under normal condition treated with miR-3036-5p agomir or antagomir after 48 h by RT-qPCR (C) and western blot (D). (E) The 20E titers under normal condition treated with miR-3036-5p agomir or antagomir after 48 h. (F) The proportion of winged offspring under normal condition treated with miR-3036-5p agomir or antagomir after 48 h. (G) Schematic diagram for crowding condition. (H) The expression level of miR-3036-5p under crowding condition treated with miR-3036-5p agomir or antagomir after 48 h. (I, J) The expression level of CYP18A1 under crowding condition treated with miR-3036-5p agomir or antagomir after 48 h by RT-qPCR (I) and western blot (J). (K) The 20E titers under crowding condition treated with miR-3036-5p agomir or antagomir after 48 h. (L) The proportion of winged offspring under crowding condition treated with miR-3036-5p agomir or antagomir after 48 h. Asterisks indicate significant differences between the treatment and the corresponding control (Student’s t-test, *0.01<p<0.05, **p<0.01).
-
Figure 6—source data 1
Labeled file for the western blot analysis in Figure 6D (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data1-v1.zip
-
Figure 6—source data 2
Uncropped file for the western blot analysis in Figure 6D (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data2-v1.zip
-
Figure 6—source data 3
Labeled file for the western blot analysis in Figure 6D (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data3-v1.zip
-
Figure 6—source data 4
Uncropped file for the western blot analysis in Figure 6D (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data4-v1.zip
-
Figure 6—source data 5
Labeled file for the western blot analysis in Figure 6J (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data5-v1.zip
-
Figure 6—source data 6
Uncropped file for the western blot analysis in Figure 6J (CYP18A1).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data6-v1.zip
-
Figure 6—source data 7
Labeled file for the western blot analysis in Figure 6J (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data7-v1.zip
-
Figure 6—source data 8
Uncropped file for the western blot analysis in Figure 6J (β-actin).
- https://cdn.elifesciences.org/articles/96540/elife-96540-fig6-data8-v1.zip
Tables
Assembly features for genomes of Metopolophium dirhodum and other Aphidinae insects.
| Genome assembly/species | M. dirhodum | S. graminum | S. miscanthi | R. maidis | A. pisum | E. lanigerum | D. noxia | M. persicae | A. gossypii |
|---|---|---|---|---|---|---|---|---|---|
| Level | Chr. | Chr. | Chr. | Chr. | Chr. | Chr. | Scaf. | Scaf. | Scaf. |
| No. chr. | 9 | 6 | 9 | 4 | 5 | 6 | - | - | - |
| Size (Mb) | 447.8 | 499.2 | 397.9 | 326 | 541.1 | 330 | 393 | 347.3 | 294 |
| No. contig | 296 | 276 | 1148 | 689 | 60,623 | 12,703 | 49,357 | 6044 | 22,569 |
| Contig N50 (bp) | 8,194,998 | 28,074,450 | 1,638,329 | 9,046,396 | 28,192 | 165,675 | 12,578 | 218,922 | 45,572 |
| No. scaf. | 68 | 22 | 656 | 220 | 23,924 | 7929 | 5641 | 4021 | 4724 |
| Scaf. N50 (bp) | 39,359,500 | 104,490,323 | 36,263,045 | 93,298,903 | 518,546 | 4,427,088 | 397,774 | 435,781 | 437,960 |
| No. gene | 18,003 | 13,353 | 16,006 | 17,629 | 36,195 | 28,186 | 19,097 | 23,910 | 14,694 |
Additional files
-
Supplementary file 1
Statistics for sequencing data for Metopolophium dirhodum assembly.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp1-v1.xlsx
-
Supplementary file 2
Overview of chromosome length of Metopolophium dirhodum assembly.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp2-v1.xlsx
-
Supplementary file 3
Gene family clusters of 10 species.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp3-v1.xlsx
-
Supplementary file 4
Chromosome-level synteny analysis between Metopolophium dirhodum and Acyrthosiphon pisum.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp4-v1.xlsx
-
Supplementary file 5
Overview of the A-to-I RNA-editing sites in Metopolophium dirhodum.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp5-v1.xlsx
-
Supplementary file 6
A-to-I RNA-editing sites for genes involved in 20E biosynthesis and metabolism pathway.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp6-v1.xlsx
-
Supplementary file 7
All primers used in this study.
- https://cdn.elifesciences.org/articles/96540/elife-96540-supp7-v1.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/96540/elife-96540-mdarchecklist1-v1.docx