Untargeted pixel-by-pixel metabolite ratio imaging as a novel tool for biomedical discovery in mass spectrometry imaging
Figures
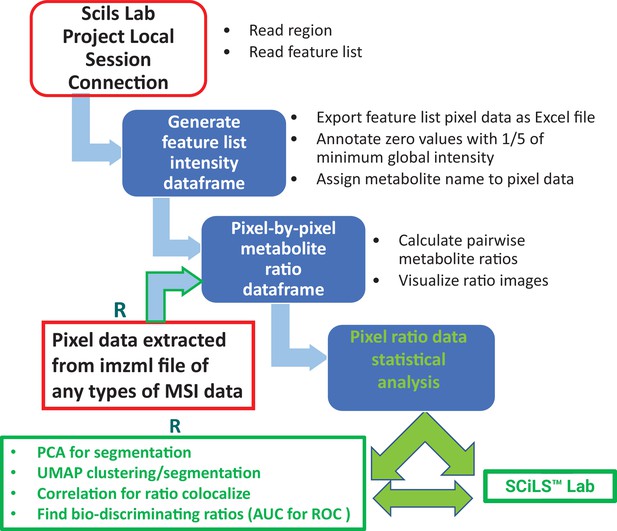
Schematic workflow of metabolite ratio imaging using pixel data from Bruker SCiLS Lab API or imzml file from any MSI data source.
For SCiLS lab MSI data, the execution of R codes starts with connecting a local SCiLS lab session project in R studio. Installation of SCiLS lab is required for this connection. After pixel data for a feature list is generated and exported as an Excel csv file, SCiLS lab installation is not required for downstream ratio imaging and data analysis, except for the potential writing of a spot image with labels back to SCiLS lab. For other types of MSI data, R code execution extracts pixel data from imzml file for subsequent ratio imaging and analysis.
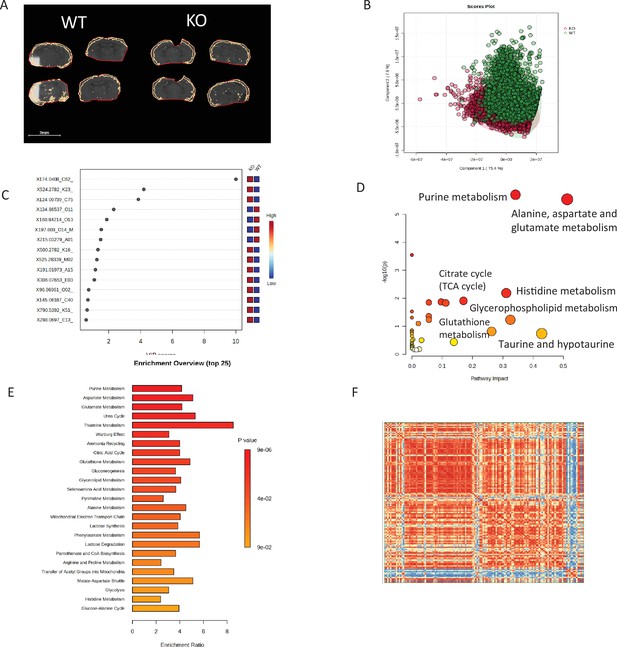
Application of pixel-by-pixel exemplary MALDI MS imaging data to metabolomics multivariate analysis using free or commercially available software package.
(A) ROIs of COX 10 KO (left 4 sections) and WT (right 4 sections) brain outer cortex; (B) PLS-DA score plot showing classification of COX 10 KO and WT cortex pixels using Metaboanalyst 5.0 (https://www.metaboanalyst.ca/MetaboAnalyst/ModuleView.xhtml). (C) Variable importance in projection (VIP) scores of metabolites from PLS-DA shown in (B). The colored boxes on the right indicate the relative concentrations of the corresponding metabolite in COX 10 KO and WT. (D) Matched pathways showing p values from the pathway enrichment analysis and pathway impact values from the pathway topology analysis. Differential metabolites imaged in cortex of COX 10 KO vs WT brain and metabolite quarries from KEGG mouse metabolic pathway were used in this analysis. (E) Metabolite set enrichment analysis showing enrichment ratio of metabolic pathway metabolites. Differential metabolites from cortex of COX 10 KO vs WT and the Small Molecule Pathway Database (SMPDB) were used for this analysis. The enrichment analysis resembles gene set enrichment analysis in transcriptomics. Panel F: Metabolite correlation heatmap generated from differential metabolites in cortex of COX 10 KO and WT. Metaboanalyst 5.0 was used for analyses shown in (B-E) and Agilent MassProfiler Professional 15.1 was used for analysis in shown in (F).
Exemplary MALDI MSI images generated before (A) and after (B) missing value annotation and from reciprocal ratios (C–D).
MSI Data was acquired at 80 µm using NEDC as matrix.
-
Figure 1—figure supplement 2—source data 1
Source data contains pixel data used to plot the figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig1-figsupp2-data1-v1.xlsx
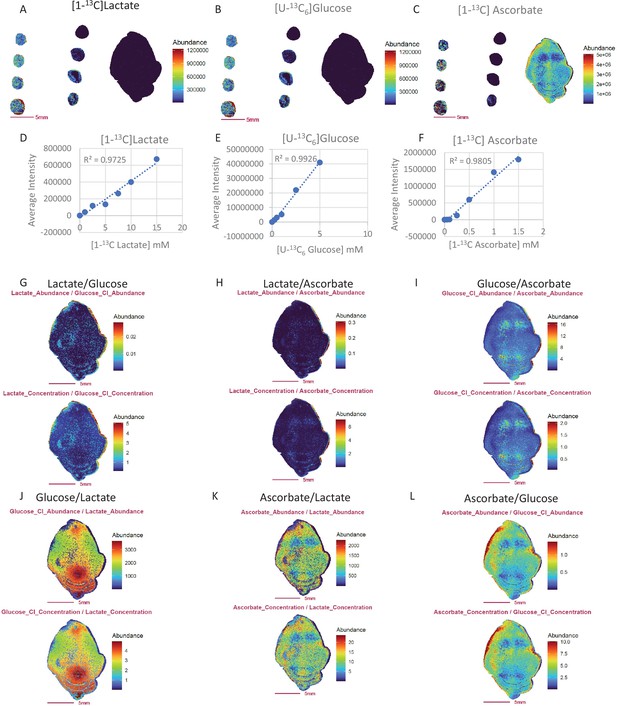
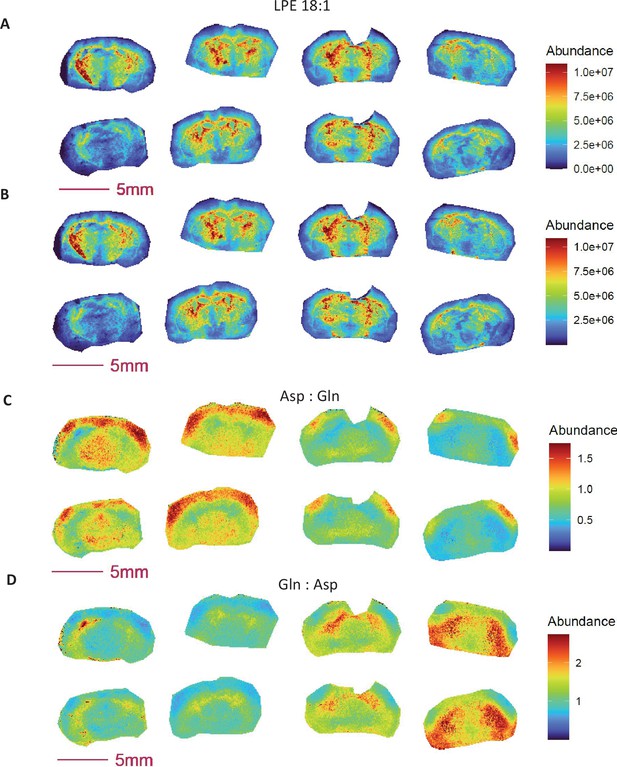
Ratio images obtained from metabolite ion abundance exhibit similar characteristics to those generated from absolute metabolite concentration.
(A-C) Images of mouse brain and mimetic brain sections spiked with stable isotope labeled [1- 13C] lactate (A), [U- 13C6]glucose (B) and [1- 13C]ascorbate (C). The concentrations of stable isotope standards from middle to left column of the slide are: 0, 1, 2.5, 5.0, 7.5, 10.0, 12.5, 15.0 mM for [1-13C] lactate; 0, 0.25, 0.5, 1.0, 2.5, 5.0, 7.5, 10 mM for [U-13C6]glucose; and 0, 0.05, 0.10, 0.25, 0.50, 0.75, 1.0, 1.5 mM for [1-13C]ascorbate. Panels (D-F) Calibration curves for [1-13C] lactate (D), [U-13C6]glucose (E) and [1- 13C]ascorbate (F). Panels (G-L) Ratio images from abundance ratio (upper image) verse concentration ratio (lower image).
-
Figure 1—figure supplement 3—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig1-figsupp3-data1-v1.xlsx
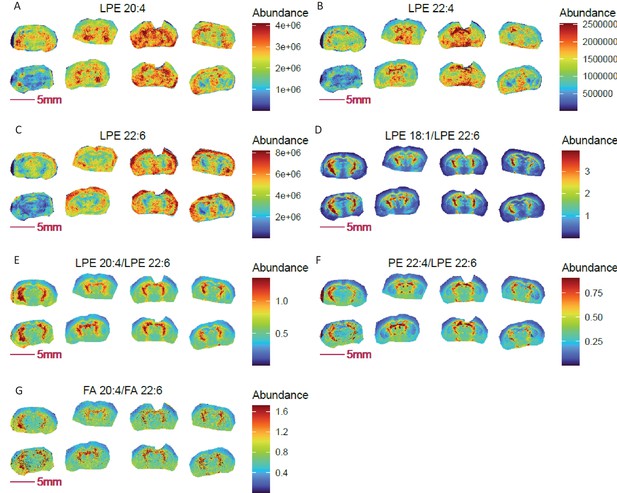
Pixel-by-pixel ratio image visualization reduces cryosection preparation artefacts and enables spatial metabolic discovery in COX 10 KO mouse brain (left 4 sections) compared to WT (right four sections).
(A-C) Blurry and diffused images of LPEs with ω3 (A–B) and ω6 (C) polyunsaturated fatty acyl chains; (D-F) sharper ratio images of ω3 fatty acid containing LPE 20:4 and LPE 22:4 to ω6 fatty acid containing LPE 22:6; (G) ratio image of ω6 arachidonic acid (FA 20:4) to ω3 docosahexaenoic acid (FA 22:6) is consistent with the identified abundance ratio distributions for LPE 20:4 and LPE 22:4 to LPE 22:6.
-
Figure 2—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig2-data1-v1.xlsx
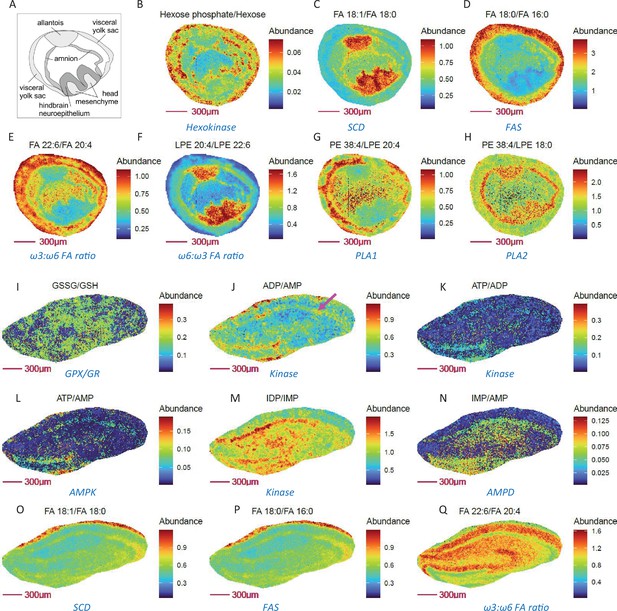
Metabolically relevant metabolite ratio pairs may probe enzymatic and metabolic pathway activity in E8.5 mouse embryo cryosections and mouse hippocampus at 10 µm spatial resolution.
(A) Annotated drawing of E 8.5 embryo tissue, derived from the section adjacent to imaged section. (B-H) Ratio of substrate/ product pairs of glycolysis, fatty acid and lipid metabolism that imply compartmented activities for hexokinase (B), fatty acid desaturation (C), fatty acid chain elongation (D), fatty acid composition (E–F) and phospholipase activity (G–H) in E 8.5 embryos. (I-Q) Ratio images showing relative oxidative stress (I), adenine nucleotide energy levels (J–L), purine nucleotide cycle activity (M–N), fatty acid desaturation (O), synthesis (P), ω3:ω6 FA ratio (Q) in neurons (purple arrow), compared to other cells in mouse hippocampus.
-
Figure 3—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig3-data1-v1.xlsx
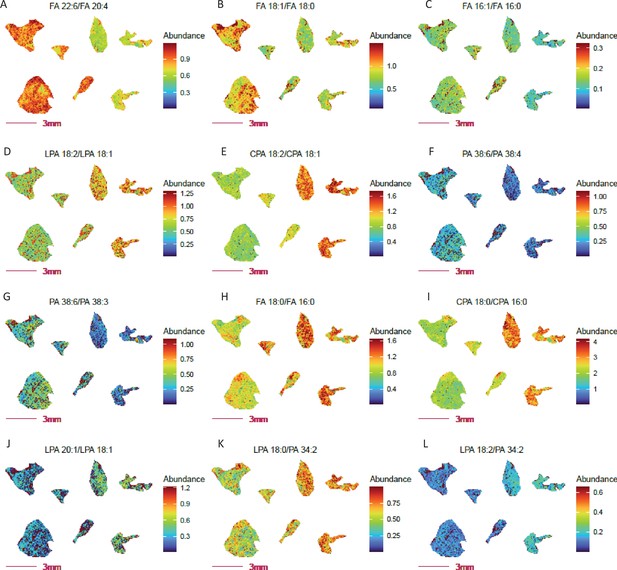
Altered fatty acid and lipid metabolism probed by ratio imaging in adipose tissue from COX 10 KO vs.WT mice.
COX 10 KO, left 4 sections; WT, right 3 sections. Imaging data was acquired at 80 µm spatial resolution. (A) Decreased abundance ratio of FA 22:6 (DHA, ω3 fatty acid) to FA 20:4 (AA, ω6 fatty acid) in COX 10 KO adipose tissue. Panels (B-C) Decreased fatty acid FA 16:0 and FA 18:0 desaturation in COX 10 KO adipose. (D-G) Altered desaturated fatty acid abundance ratios for lysophospolipids and phospholipids in COX 10 KO adipose tissue. Panels (H-J) Elevated fatty acid synthesis inferred by free fatty acid and phospholipids ratios, considering FA 18:0 to FA 16:0 (H–I) and FA 20:1 to 18:1 (L) in COX 10 KO adipose. Panels (K-L) Elevated PLA2 (K) and PLA1(L) activities implied by the elevated ratio of Lyso PA to PA in COX 10 KO adipose.
-
Figure 4—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig4-data1-v1.xlsx
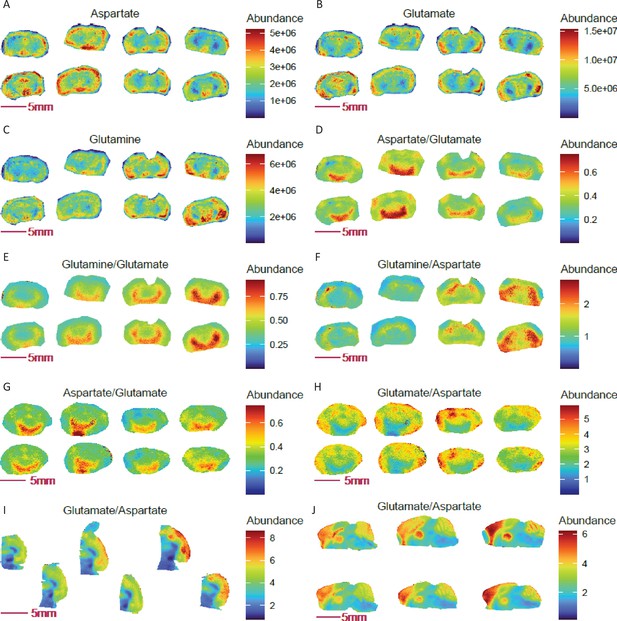
Ratio imaging uncovers novel mouse brain regions with genotype-specific and spatially resolved ratios of aspartate, glutamate, and glutamine.
(A-C) Images of glutamine, glutamate and aspartate in coronal sections of COX 10 KO (left 4 sections) and WT (right 4 sections) mouse brains. (D-F) Differential abundance ratios of aspartate, glutamate and glutamine observed in defined regions of the COX 10 KO and WT brains (p<0.0001, two-tailed Student t-test with Benjamini-Hochberg correction). Panels G-H: Reciprocal and differential aspartate to glutamate ratio and Arc-like NAC containing region was repeated in a separate group of COX 10 KO (left 4 sections) and WT (right four sections) mouse brains. (I-J) Enrichment in the glutamate to aspartate ratio in the cortex region of mouse brain, as revealed in horizonal and sagittal sections.
-
Figure 5—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig5-data1-v1.xlsx
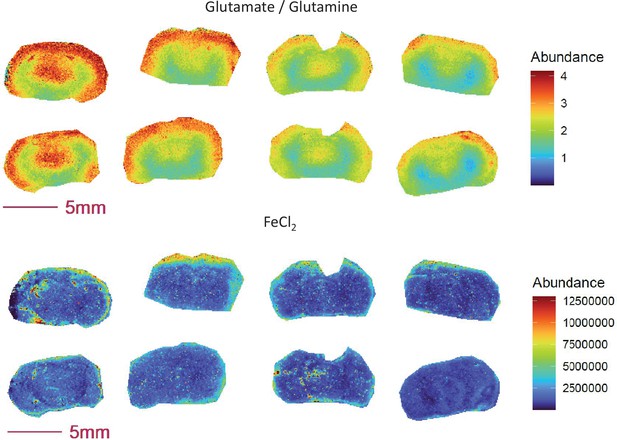
Ratio of glutamate to glutamine and Fe2+ abundance are co-enriched in the outer cortex of the COX10 KO (right 4 sections) and WT (left 4 sections) brain.
-
Figure 5—figure supplement 1—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig5-figsupp1-data1-v1.xlsx
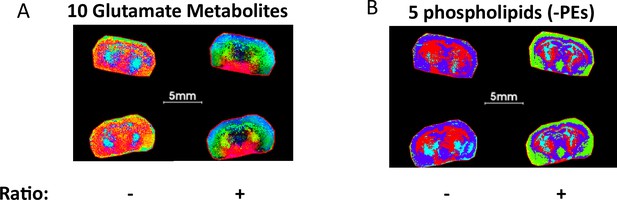
PCA segmentation of mouse brains comparing ratio and non-ratio metabolite pixel data.
(A) PCA considering the top 5 components from ratios of 10-glutamate related metabolites that reveal an unanticipated arc-like regional distribution within the hypothalamus, striatum and NAc, that is not detected by consideration of only non-ratio metabolite data. (B) PCA considering the top 5 components 5-LPE metabolites showing fine brain structure that is not fully represented by only non-ratio metabolite segmentation.
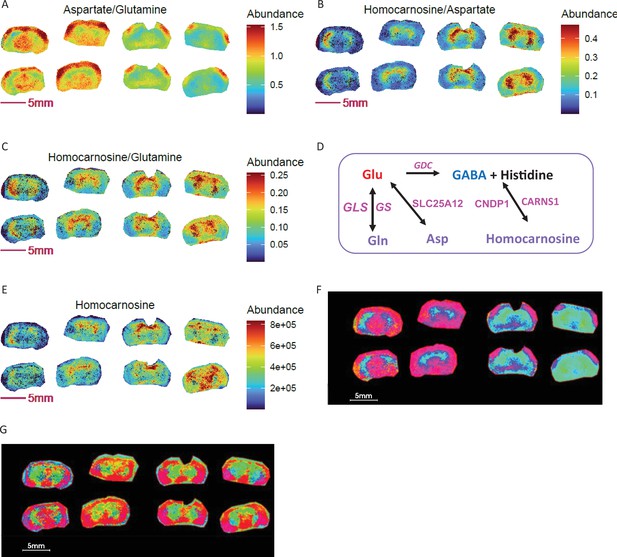
Metabolite ratios for segmentation and biomarker discovery in segmented ROIs.
KO: right 4 sections; WT: left 4 sections. (A-C) Ratio images of homocarnosine, glutamine and aspartate. (D) Enzymatic regulation and interconversions among brain glutamine, aspartate and homocarnosine. (E) Image of homocarnosine. (F) PCA with top 5 components and using ratios of homocarnosine, glutamine and aspartate shows differential ROI abundance shown in cyan blue in larger area of striatum, thalamus hypothalamus regions in KO compared to WT. (G) PCA with top 5 components using homocarnosine, glutamine, and aspartate pixel data show indistinguishable ROIs between genotypes.
-
Figure 7—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig7-data1-v1.xlsx
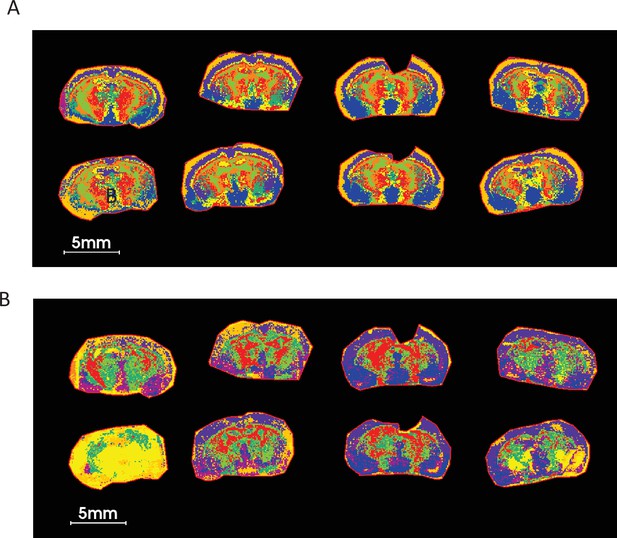
UMAP analysis using five LPEs ratio followed by K-mean clustering on COX KO and WT brain sections shows a much cleaner and biologically relevant ROIs compared to those of LPE non-ratio.
(A) UMAP clustering using ratios of five LPEs. (B) UMAP clustering using five LPE pixel data. The list of five LPEs is shown Supplementary file 4. Nine neighbors, two components with Euclidean distance as metric measurement of similarities were used in this UMAP analysis.
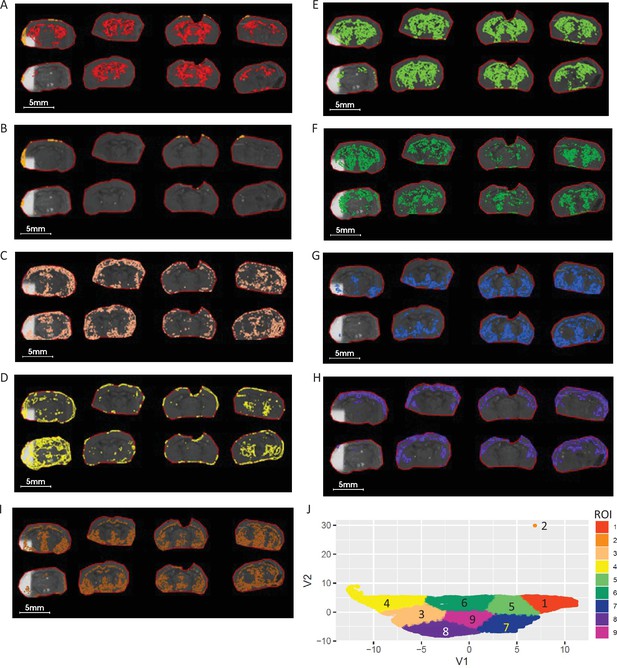
ROIs and UMAP scatter plots generated from non-ratio metabolite data.
(A-I) Nine segmented ROIs generated UMAP and K-mean clustering analysis corresponding to clustered regions of 1–9. (J) Scatterplot of UMAP first 2 components (V1 vs V2) projecting the distribution and similarity of nine segmented ROIs resulting in a incomplete and inconsistent ROIs among replicates. 5 LPE (Supplementary file 4) pixel data was used for UMAP analysis.
-
Figure 8—figure supplement 1—source data 1
Source data contains pixel data for UMAP score plot.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp1-data1-v1.csv
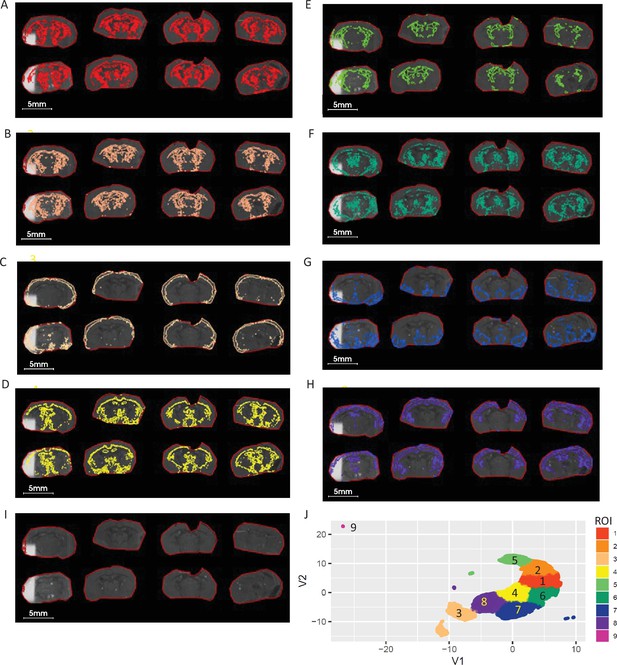
ROIs and UMAP scatter plots generated from metabolite ratio data.
(A-I) Nine segmented ROIs generated UMAP and K-mean clustering analysis corresponding to clustered regions of 1–9. (J) Scatterplot of UMAP first 2 components (V1 vs V2) projecting the distribution and similarity of nine segmented ROIs shows a distinct LPE profile between outer cortex (ROI 3) cortex (ROI 8) and ventral thalamus (ROI 5) striatum region(ROI2). Ratio of five LPE (Supplementary file 4) pixel data was used for UMAP.
-
Figure 8—figure supplement 2—source data 1
Source data contains data for UMAP score plot.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp2-data1-v1.csv
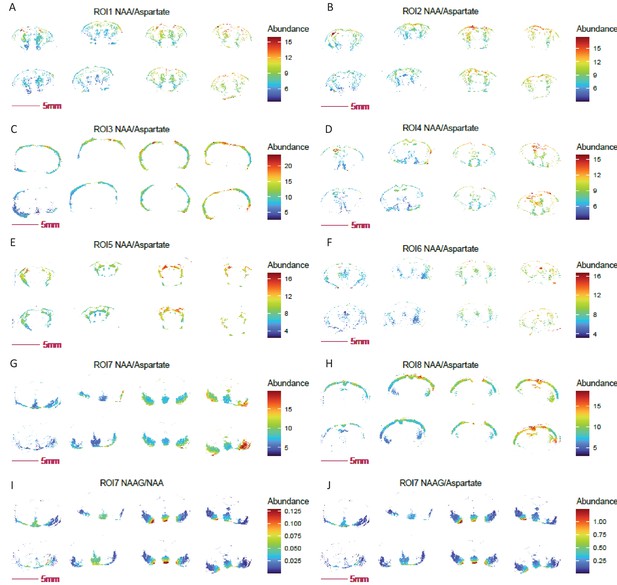
Ratio images of NAA, NAAG and aspartate for UMAP-generated ROIs.
(A-H) Ratio image of NAA to aspartate in eight UMAP-segmented ROIs. (I-J) Ratio images of NAAG to NAA (I) and NAAG (J) to aspartate in ROI 7 of UAMP segmented ROIs. COX10 KO, right four sections; WT, left four sections.
-
Figure 8—figure supplement 3—source data 1
Source data contains ROI pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp3-data1-v1.xlsx
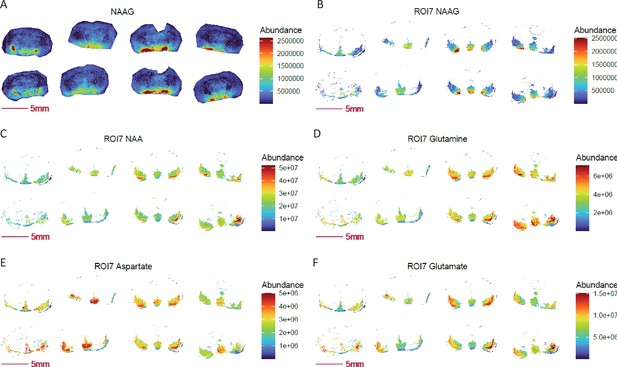
Images of NAAG and neurotransmitter related metabolites visualized in UMAP-generated ROI 7.
(A) NAAG is enriched and elevated in ROI 7–like region of COX 10 KO brain. (B-D) NAA,NAAG and glutamine are elevated in ROI 7 of COX 10 KO brain. (E) Decreased level of aspartate in ROI 7 of COX10 KO; (F) Glutamate shows no change in ROI 7 of COX 10 KO compared to WT. COX10 KO, right four sections; WT, left four sections.
-
Figure 8—figure supplement 4—source data 1
Source data contains pixel data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp4-data1-v1.xlsx
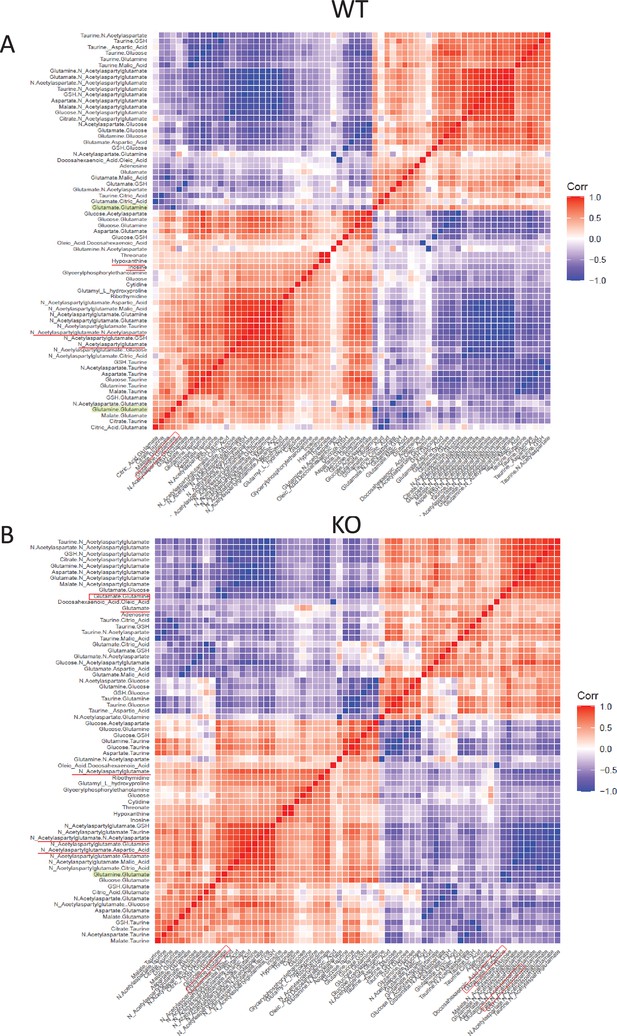
Pixel-by-pixel correlation of glutamine to glutamate ratio with a compendium of all detected metabolites and ratios exclusively in UMAP-generated ROI 7, comparing WT (A) to COX 10 KO (B) brain.
4 biological replicates for each genotype. Only significant correlations with a spearman correlation coefficient of greater than 0.4 were presented in the figure.
-
Figure 8—figure supplement 5—source data 1
Source data contains correlation matrix data for this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp5-data1-v1.xlsx
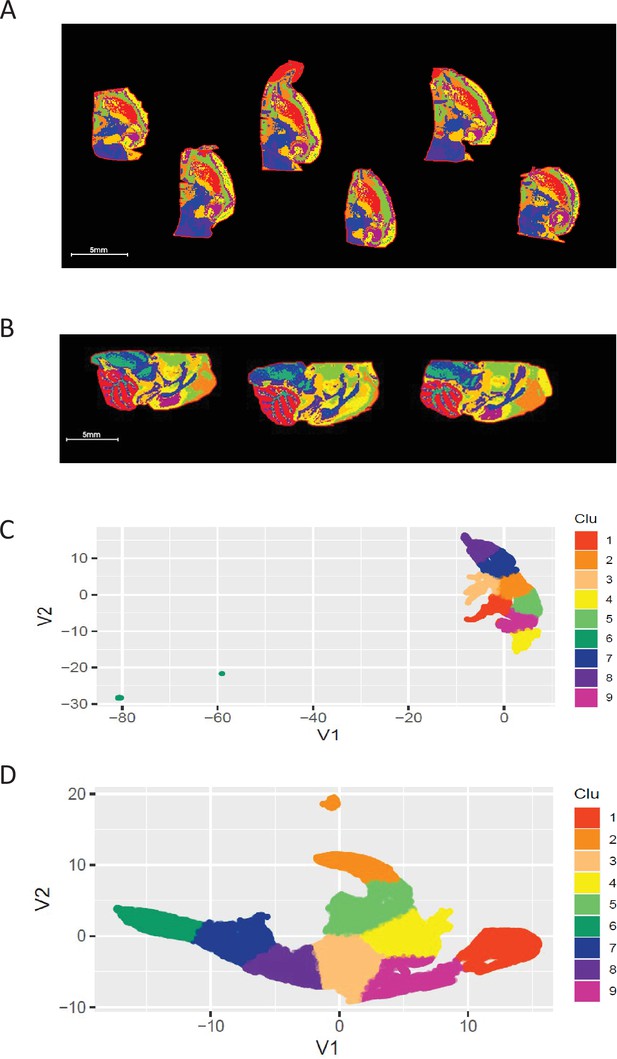
UMAP using five LPE ratios generated ROIs consistent with brain anatomy in two separate experiments with horizonal and sagittal brain sections.
(A-B) Overlay of nine segmented ROIs generated from UMAP and K-mean clustering analysis in horizonal hemisphere sections (A) and sagittal brain sections (B). (C-D) Scatterplot of UMAP first 2 components (V1 vs V2) projecting the distribution and similarity of 9 segmented ROIs in horizonal (C) and sagittal sections (D). See Supplementary file 4 for the list of five LPEs.
-
Figure 8—figure supplement 6—source data 1
Source data contains data for UMAP score plot in this figure.
- https://cdn.elifesciences.org/articles/96892/elife-96892-fig8-figsupp6-data1-v1.xlsx
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Biological sample (Mus musculus) | Mouse brain | Animal facility Weill Cornell Medicine | n/a | Fresh frozen |
| Biological sample (Mus musculus) | Mouse adipose | Animal facility Weill Cornell Medicine | n/a | Fresh frozen |
| Chemical compound, drug | N-(1-Naphthyl)ethylenediamine dihydrochloride | Sigma-Aldrich | Cat # 222488 | |
| Chemical compound, drug | 9-Aminoacridine | Millipore Sigma | Cat # 92817 | |
| Other | Indium tin oxide | ITO; (Delta Technologies) | Cat # CB-90IN-S111 | |
| Chemical compound, drug | 1,5-Diaminonaphthalene | Millipore Sigma | Cat # 56451 |
Spearman correlation of an unknown with ratio and non-ratio entities.
| Entity | Unknown [M-H] 375.23059 |
|---|---|
| CPA 18:1 | 0 86090633 |
| CPA 16:0 | 0 85862853 |
| Docosahexaenoic Acid | 0 84761078 |
| Palmitic Acid | 0 80091519 |
| LPA 18:1 | 0 78455912 |
| LPA 16:0 | 0 77663768 |
| Oleic Acid | 0 77371792 |
| PA 34:2 | 0 73231961 |
| PA 36:3 | 0 72919804 |
| Palmitoleic Acid | 0 72724047 |
| CPA 18:0 | 0 70128825 |
| LPE 16:0 | 0 69156441 |
| PA 38:6 | 0 68799022 |
| LPA 18:0 | 0 67264794 |
| 558.4283 | 0 67163649 |
| PE P 16:0 | 0 66456185 |
| Linoleic Acid | 0 65773742 |
| PI 36:2 | 0 65460109 |
| LPA 18:2/LPA 18:1 | –0 53780801 |
| PA 36:1/LPA 18:1 | –0 4549225 |
| PA 36:4/LPA 16:0 | –0 42353603 |
Brain glutamate to glutamine ratio positively correlates FeCl2 using combined ratio and non-ratio pixel data and spearman correlation.
| Metabolite or Metabolite ratio | Glutamate/Glutamine |
|---|---|
| Glutamate/Glutamine | 1 |
| N-Acetylaspartate/Glutamine | 0.69140746 |
| Glutamate/Glucose | 0.68318508 |
| Aspartate/Glutamine | 0.617246741 |
| Aspartate/Glucose | 0.571634522 |
| N-Acetylaspartate/Glucose | 0.487447557 |
| Glutathione/Glutamine | 0.482704981 |
| Glutamate/Malate | 0.477826203 |
| Glutamate/NAAG | 0.418208761 |
| Glutathione/Glucose | 0.414914055 |
| Glutamate/Taurine | 0.408270132 |
| Aspartate/NAAG | 0.40560898 |
| FeCl2 | 0.40137113 |
Additional files
-
Supplementary file 1
Comparison of untargeted ratio imaging R workflow to other ratio software.
- https://cdn.elifesciences.org/articles/96892/elife-96892-supp1-v1.docx
-
Supplementary file 2
Brain glutamine to glutamate ratio negatively correlates FeCl2 using combined ratio and non-ratio pixel data and spearman correlation.
- https://cdn.elifesciences.org/articles/96892/elife-96892-supp2-v1.pdf
-
Supplementary file 3
List of glutamate metabolism related metabolites for ratio generation and PCA segmentation of brain sections.
- https://cdn.elifesciences.org/articles/96892/elife-96892-supp3-v1.docx
-
Supplementary file 4
List of lyso PEs for ratio generation and PCA segmentation of brain sections.
- https://cdn.elifesciences.org/articles/96892/elife-96892-supp4-v1.docx
-
Supplementary file 5
Comparing correlation of neurotransmitter related metabolites and ratios in ROI7 of WT and KO brains.
- https://cdn.elifesciences.org/articles/96892/elife-96892-supp5-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/96892/elife-96892-mdarchecklist1-v1.docx





