AI-based discovery and cryoEM structural elucidation of a KATP channel pharmacochaperone
Figures
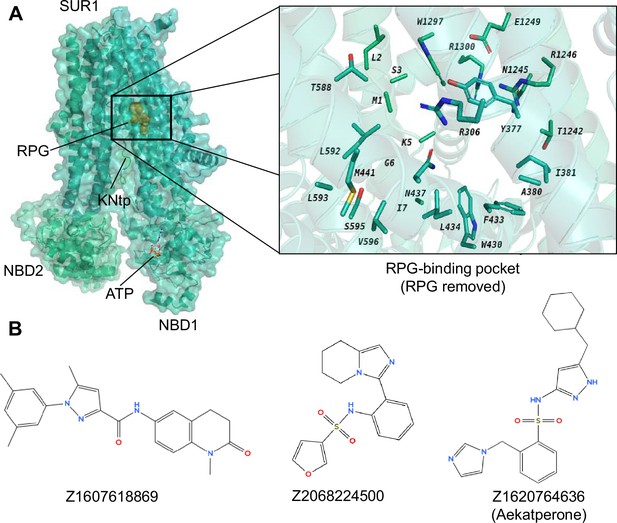
Identification of potential pharmacochaperones for pancreatic KATP channels through virtual screening and experimental validation.
(A) Left: Cryo-electron microscopy (cryoEM) structure of repaglinide (RPG)-bound pancreatic KATP channel (PDB ID: 6PZ9; only the SUR1 ABC core and Kir6.2-N terminal peptide [KNtp] are shown; RPG is shown in gold spheres and ATP bound at the nucleotide binding domain [NBD]1 of SUR1 is shown as red sticks). Right: A close-up view of the target site used for virtual screening of a library of ~2.5 million compounds (from Enamine), utilizing the AtomNet model (RPG is removed to show only the binding residues from SUR1). From this screening, a set of 96 ‘top scoring compounds’ was selected for experimental validation. (B) Chemical structures of the top three compounds, C24, C27, and C45, identified via experimental testing of pharmacochaperone effects on a KATP trafficking mutant (Figure 1—figure supplement 1) correspond to ZINC IDs Z1607618869, Z2068224500, and Z1620764636, respectively. These compounds are chemically distinct from one another. Note C27 and C45 both contain a sulfonamide group (-SO2-N-), which is different from the sulfonylurea group (-SO2-NH-CO-NH-) seen in known KATP inhibitors such as glibenclamide and tolbutamide.
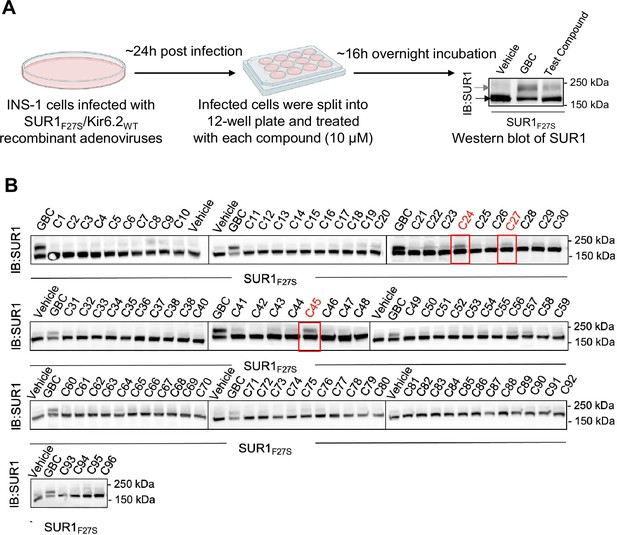
Experimental testing of the top 96 compounds identified from AtomNet-based virtual screening.
(A) Experimental workflow to screen for potential pharmacochaperone effects of the top scoring 96 compounds from virtual screening (see Supplementary file 1). INS-1 cells co-expressing the trafficking mutant SUR1F27S and Kir6.2WT were treated with 10 μM of 1 of the 96 compounds for 16 hr and harvested for western blot of SUR1. Cells treated with 0.1% DMSO served as a vehicle control, while cells treated with 10 μM glibenclamide (GBC) was used as a positive control. The lower band (black arrow) corresponds to the core-glycosylated immature SUR1, and the upper band (gray arrow) the complex-glycosylated mature SUR1. Molecular mass markers in kilodaltons (kDa) are shown on the right of the blots. (B) Western blots showing the effects of the 96 compounds tested, with the top three compounds (C24, C27, and C45) highlighted by red boxes. Experiments were conducted twice with similar results.
-
Figure 1—figure supplement 1—source data 1
PDF file containing original western blots for panel B, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig1-figsupp1-data1-v1.zip
-
Figure 1—figure supplement 1—source data 2
Original files for western blot analysis displayed in panel B.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig1-figsupp1-data2-v1.zip
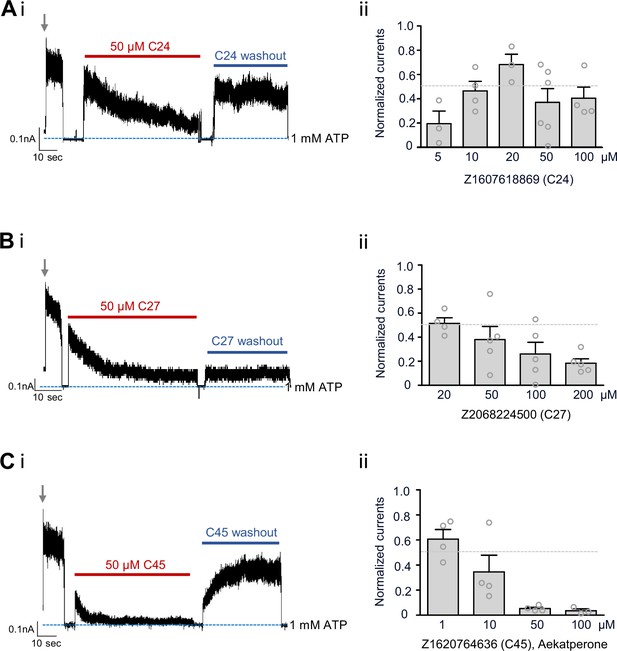
The top three compounds identified via western blot screening assay inhibit KATP channel activity.
(Ai, Bi, Ci) Representative inside-out patch-clamp recordings of COSm6 cells expressing wild-type (WT) SUR1/Kir6.2 channels. Patches were excised into K-INT/1 mM EDTA solution (indicated by the gray arrows), and then exposed to the same solution containing 50 μM of C24 (A), C27 (B), or C45 (C), followed by washout of the compounds to assess reversibility. Patches were exposed to K-INT/1 mM EDTA plus 1 mM ATP in between solution exchanges to monitor baseline (blue dashed line). Recordings were conducted at –50 mV and inward currents displayed as upward deflections. The vertical scale bar indicates current amplitudes in nanoamperes (nA), and the horizontal scale bar represents recording duration in seconds. (Aii, Bii, Cii) Steady-state inhibition by C24 (A), C27 (B), and C45 (C) at various concentrations indicated in the x-axis. Currents in the presence of the compounds at steady state (the end of compound exposure) were normalized to currents seen in K-INT/1 mM EDTA at the time of patch excision. Each bar represents the mean ± SEM of three to six patches (gray circles). The dotted gray line marks 50% inhibition.
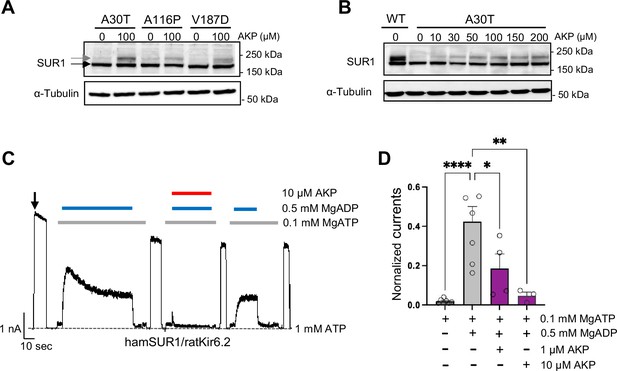
Aekatperone has dual pharmacochaperone and inhibitory actions on pancreatic KATP channels.
(A) Representative western blots of SUR1 from COSm6 cells co-transfected with cDNAs of wild-type (WT) Kir6.2 and trafficking mutants of SUR1 TMD0 domain, A30T, A116P, or V187D, and treated with either 0.1% DMSO (0 μM) or 100 μM Aekatperone (AKP) for 16 hr. The core-glycosylated immature SUR1 and the complex-glycosylated mature SUR1 are indicated by the black and gray arrows, respectively. The tubulin blot below serves as a loading control. (B) Representative western blots of SUR1 from COSm6 cells co-transfected with cDNAs of WT Kir6.2 and a SUR1 trafficking mutant A30T, and treated with either 0.1% DMSO (0 μM) or various concentrations of Aekatperone (10, 50, 100, 150, 200 μM) for 16 hr. WT SUR1 from cells co-transfected with WT Kir6.2 and WT SUR1 without Aekatperone treatment served as a control (left lane). (C) Representative recording from COSm6 cells co-transfected with hamster SUR1 and rat Kir6.2. Channels were exposed to K-INT solution upon patch excision (arrow) and exposed to solutions containing MgATP, MgADP, or Aekatperone as indicated by the bars above the recordings and the labels on the right. The patch was exposed to 1 mM ATP periodically to ensure the baseline has not shifted (gray dashed line). (D) Quantification of currents (normalized to currents in K-INT/1 mM EDTA at the time of patch excision) in various solutions from recordings such as that shown in (A). Each bar represents the mean ± SEM of at least three patches, with circles showing individual patches. *p<0.05 by one-way ANOVA and Dunnet’s post hoc test.
-
Figure 2—source data 1
PDF file containing original western blots for panels A and B, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig2-data1-v1.zip
-
Figure 2—source data 2
Original files for western blot analysis displayed in panels A and B.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig2-data2-v1.zip
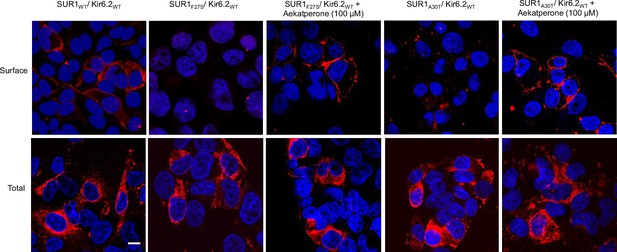
Aekatperone rescues surface expression of SUR1F27S and SUR1A30T trafficking mutants.
Top: Surface staining of COSm6 cells transiently co-transfected with Kir6.2WT and various FLAG-tagged (extracellular N-terminus) SUR1 constructs (WT, F27S, or A30T) treated with or without Aekatperone using M2 anti-FLAG mouse monoclonal antibody followed by Alexa Fluor 546-conjugated anti-mouse secondary antibody. Bottom: Total cellular staining of FLAG-tagged SUR1. Cells were fixed and permeabilized prior to incubation with primary and secondary antibodies. Images shown are stacked Olympic Fluoview confocal sections, with nuclei counterstained using DAPI. Scale bar, 10 μm.
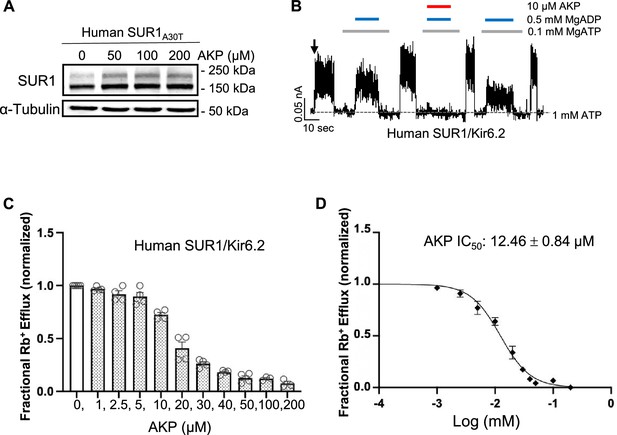
Aekatperone has the same pharmacochaperoning and inhibitory effects on human KATP channels.
(A) Representative western blot of SUR1 from COSm6 cells co-transfected with cDNAs of wild-type (WT) human Kir6.2 and human SUR1A30T trafficking mutant and treated with either 0.1% DMSO (0 μM) or various concentrations of Aekatperone (AKP) for 16 hr. (B) Representative recording from COSm6 cells co-transfected with human SUR1 and Kir6.2. Channels were exposed to K-INT solution upon patch excision (arrow) and exposed to solutions containing MgATP, MgADP, or Aekatperone as indicated by the bars above the recordings and the labels on the right. The patch was exposed to 1 mM ATP periodically to ensure the baseline has not shifted (gray dashed line). (C) Rb+ efflux assay results showing Aekatperone dose-dependently inhibits human WT KATP channels expressed in COSm6 cells and opened by metabolic inhibition (see Methods). The fractional Rb+ efflux was calculated by subtracting efflux in untransfected cells and normalizing to efflux in cells treated with 0.1% DMSO. (D) Aekatperone inhibition dose-response curve from data shown in (C) fitted with a Hill equation with variable slope using GraphPad Prism 10.
-
Figure 2—figure supplement 2—source data 1
PDF file containing original western blots for panel A, indicating the relevant bands and treatments.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig2-figsupp2-data1-v1.zip
-
Figure 2—figure supplement 2—source data 2
Original files for western blot analysis displayed in panel A.
- https://cdn.elifesciences.org/articles/103159/elife-103159-fig2-figsupp2-data2-v1.zip
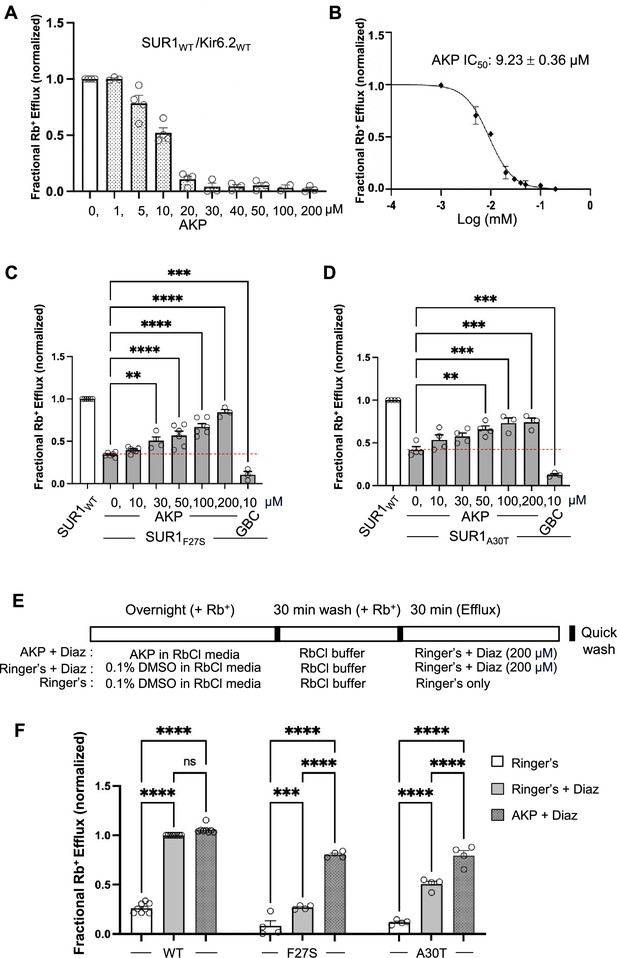
Functional recovery of mutant KATP channels rescued by Aekatperone.
(A) Rb+ efflux assay results showing Aekatperone (AKP) dose-dependently inhibited wild-type (WT) pancreatic KATP channels (hamster SUR1 and rat Kir6.2) expressed in COSm6 cells and opened by metabolic inhibition (see Methods). The fractional Rb+ efflux was calculated by subtracting efflux in untransfected cells and normalizing to efflux in cells treated with 0.1% DMSO. (B) Dose-response curve of Aekatperone inhibition from data shown in (A) fitted with a Hill equation with variable slope using GraphPad Prism 10. The IC50 is 9.23 µM±0.36 µM. Error bars represent the SEM. (C, D) Bar graphs showing dose-response enhancement in KATP channel activity as assessed by Rb+ efflux assay in COSm6 cells expressing two different trafficking mutations, SUR1F27S (A) and SUR1A30T (B). The cells were treated with varying concentrations of Aekatperone (10, 30, 50, 100, or 200 μM), glibenclamide (GBC) at 10 μM, or 0.1% DMSO as a vehicle control (0 μM Aekatperone) for 16 hr. Aekatperone and GBC was excluded from the efflux solutions during the efflux assay. Untransfected (UT) cells were included to quantify background Rb+ efflux, which was subtracted from other experimental readings. The data were normalized to the fractional Rb+ efflux of cells expressing WT channels. Error bars represent the SEM of at least three independent experiments (circles are individual data points from three to six different experiments). Statistical significance was performed using one-way ANOVA followed by Dunnett’s post hoc multiple comparison test, alpha = 0.05. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. A red dashed line is shown to indicate the basal efflux of the mutant channels under vehicle control conditions (in the absence of Aekatperone). (E) Schematic of experimental design for (F). COSm6 cells transfected with WT or various mutant channels were treated with 0.1% DMSO (vehicle control) or 100 Aekatperone (AKP) overnight in the presence of Rb+. Before efflux measurements, cells were washed in an RbCl containing buffer lacking AKP for 30 min. Efflux was then measured for 30 min in a Ringer’s solution±diazoxide (Diaz) at 200 μM. Note, diazoxide was included in Ringer’s solution during the efflux assay but not during the overnight incubation. (F) Rb+ efflux experiments showing overnight treatment with AKP enhances acute Diaz response in COSm6 cells expressing trafficking mutants. Each bar represents the mean ± SEM of at least three different biological repeats, with circles indicating individual data points, alpha = 0.05. *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001 by one-way ANOVA with Dunnett’s multiple comparisons test.
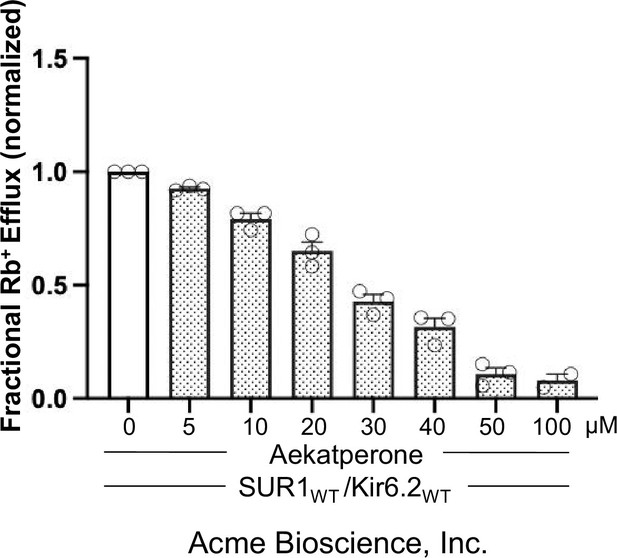
Acute inhibitory effect of Aekatperone purchased from Acme Bioscience, Inc.
Note the exact concentrations of Aekatperone may not precisely match those shown in Figure 3A due to the very small quantity received from Acme Bioscience, which made reconstitution less accurate.
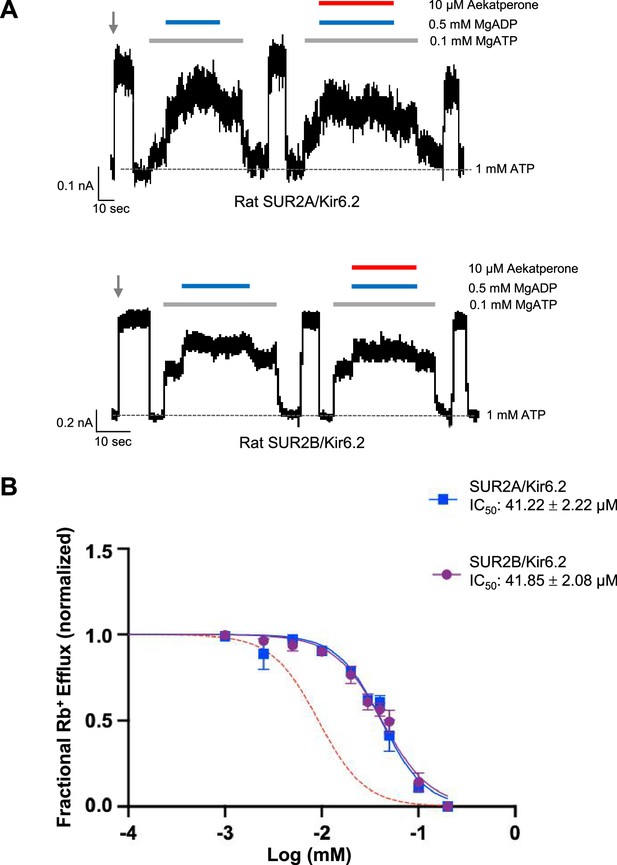
SUR2-containing KATP channels are less sensitive to Aekatperone inhibitory effects.
(A) Representative recordings from COSm6 cells expressing SUR2A/Kir6.2 (top) or SUR2B/Kir6.2 channels. Channels were exposed to K-INT solution upon patch excision (gray arrow) and exposed to solutions containing MgATP, MgADP, or Aekatperone as indicated by the bars above the recordings and the labels on the right. The patch was exposed to 1 mM ATP periodically to ensure the baseline has not shifted (gray dashed line). (B) Half-maximal inhibitory concentration (IC50) of Aekatperone on SUR2A/Kir6.2 or SUR2B/Kir6.2 channels transiently expressed in COSm6 cells Kir6.2 using Rb+ efflux assay. Data were fit with a Hill equation with variable slope using GraphPad Prism 10. The Aekatperone IC50 is 41.22 µM±2.22 µM (SEM) for SUR2A/Kir6.2 channels and 41.85 µM±2.08 µM (SEM) for SUR2B/Kir6.2 channels. The error bar for each data point represents the SEM. Red dotted line represents the dose-response curve of Aekatperone on SUR1/Kir6.2 channels from Figure 3B for comparison.
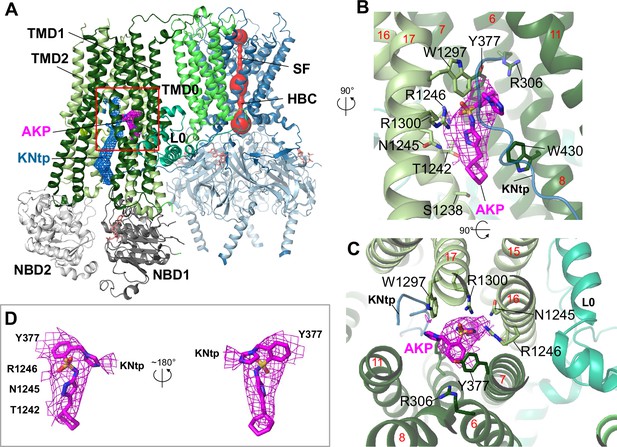
Structure of the KATP channel in complex with Aekatperone.
(A) Structural model of the pancreatic KATP channel in complex with Aekatperone showing SUR1 in NBD-separated conformation and the Kir6.2 pore (red) constricted at the helical bundle crossing (HBC) and the selectivity filter (SF) gates. Aekatperone is shown as purple mesh (0.08 V contour) and N-terminal domain of Kir6.2 (KNtp) as blue mesh (0.08 V contour). Only one SUR1 subunit attached to Kir6.2 core is shown based on the focused refinement cryo-electron microscopy (cryoEM) map of the Kir6.2 tetramer and a single SUR1 subunit. SUR1 transmembrane domain (TMD) 1 and 2 are colored in dark green and light green, respectively. NBD, nucleotide binding domain; L0, L0 loop of SUR1. (B, C) Close-up side view (B) and top-down view (C) of the Aekatperone binding site. Aekatperone cryoEM density and model are shown in magenta and SUR1 residues that interact with Aekatperone are shown as sticks. KNtp is shown as a blue main chain peptide. Red numbers indicate the numbers of helices of SUR1. (D) Aekatperone cryoEM density map (0.08 V contour) and model fitting in two different views. Note at the contour shown, some surrounding density from interacting SUR1 and Kir6.2 is included.
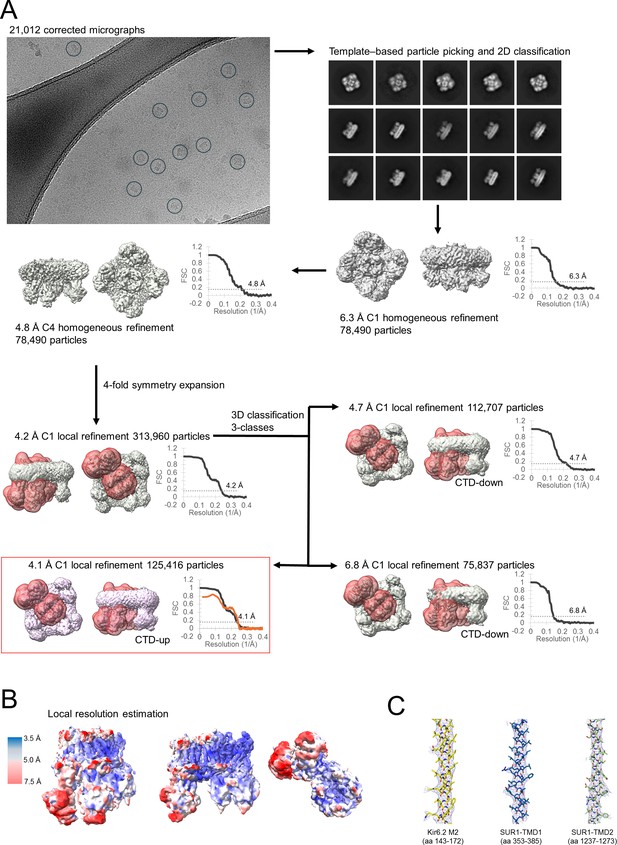
Cryo-electron microscopy (cryoEM) data processing.
(A) Workflow showing the total number of symmetry-expanded particles yielding the 4.1 Å C1 nonuniform refinement reconstruction is 125,416. The raw image on the top left corner serves to illustrate the particles selected (circles) for processing. The Fourier shell correlation (FSC) curve between two independent half-maps calculated for each refinement is located near the map (black curve). Where masks are applied, the mask used for its FSC calculations is shown in red. For the final map (red box), both FSC curve between two independent half-maps (black curve) and between model and map (orange curve; unmasked; PDB ID 9DFX and EMD-46820) are overlayed. For gold-standard FSC resolution cutoff (GSFSC) calculations, FSC automask was applied. FSC curves were calculated in Phenix and cryoSPARC. (B) Local resolution estimate for the reconstructed map at 0.08 V contour, with micelle density not shown, in side view (left, middle) and down-top view (right). No local filtering or local sharpening was used for visualization. Local resolution estimates were calculated in cryoSPARC and visualized in ChimeraX. (C) Representative close-up views of cryoEM density map (blue mesh, at 0.08 V contour) superposed with the structural model.
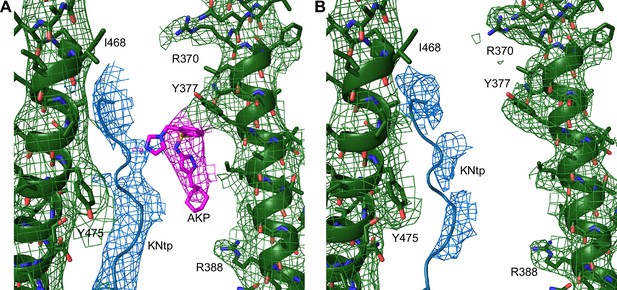
Comparison of AKP-bound and AKP-free structures.
(A) Aekatperone (AKP)-bound map (EMD-46820) superimposed with the structural model (PDB ID 9DFX) and (B) an AKP-free apo map (EMD-26320), also superimposed with the protein structural model (PDB ID 9DFX) for comparison.The cryo-electron microscopy (cryoEM) density in both maps was filtered to 4.5 Å, aligned in ChimeraX, normalized and displayed using PyMOL at 4.6 RMSD (equivalent to 0.08 V contour). The cryoEM density corresponding to AKP colored in magenta in (A) is absent in (B). The Kir6.2 N-terminal peptide (KNtp) density (modeled as a polyalanine chain) colored in blue is stronger in AKP-bound map than in the apo map. The cryoEM density corresponding to SUR1 is colored in green in both panels.
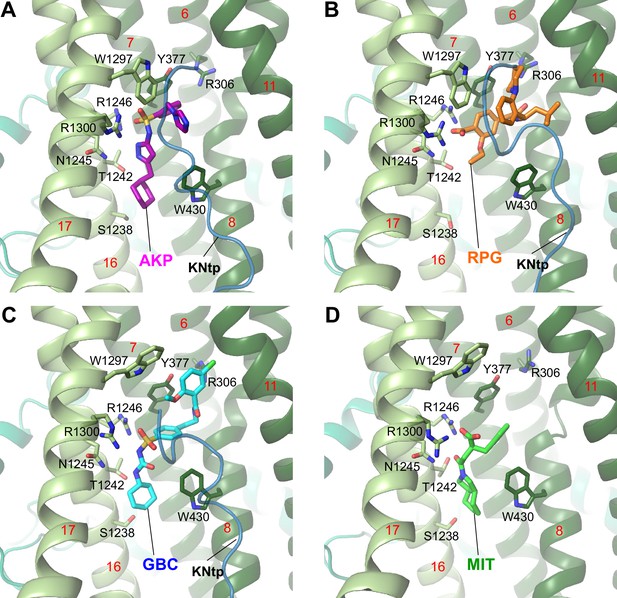
Binding site model for (A) Aekatperone (AKP), (B) repaglinide (RPG, PDB ID 7TYS), (C) glibenclamide (GBC, PDB ID 7U24) bound to pancreatic KATP channels, and (D) mitiglinide (MIT, PDB ID 7WIT) bound to a truncated SUR1 missing amino acids 2–208 in the absence of the Kir6.2 subunit.
In A–C, KNtp is shown as a blue peptide backbone.
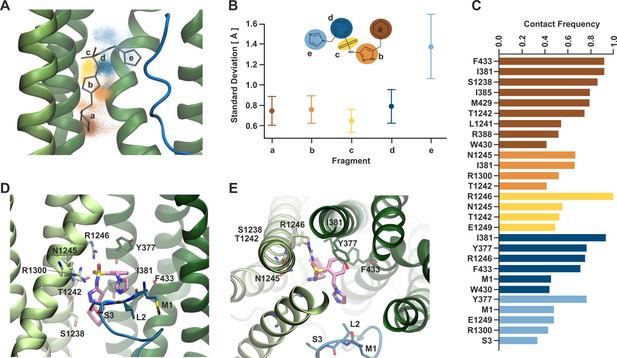
Dynamical behavior of Aekatperone in the SUR1 cavity investigated by molecular dynamics (MD) simulations.
(A) Scatter of the positions of the centers of mass of individual fragments of Aekatperone during the simulation. The initial position is marked as sticks. Green helices are TMD1/2 of SUR1, and the blue fragment is KNtp. (B) Standard deviation of the positions of the centers of mass of individual fragments during the simulation. The fragments are color-coded throughout the figure. (C) Frequency of staying in close contact with SUR1 and KNtp residues during the simulation, divided by individual fragments. (D, E) An example snapshot from the simulation of Aekatperone in the pocket showing its interactions with SUR1 and KNtp residues - side view (D) and top view (E).
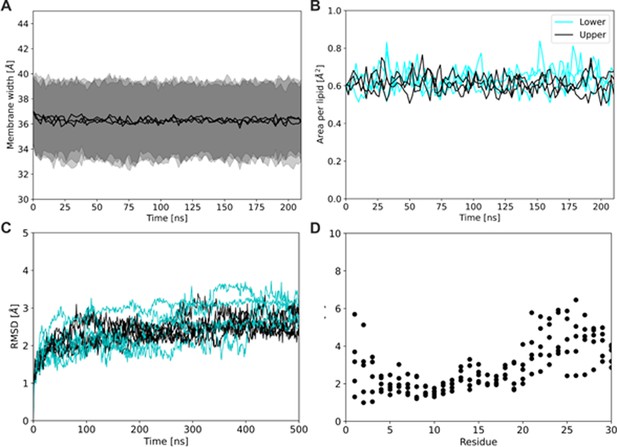
Properties of the system during the extended equilibration of the system (200 ns).
Calculated fluctuations in membrane thickness (A) and area per lipid indicating membrane density (black and cyan colors correspond to the upper and lower leaflet of the membrane, respectively) (B) suggest sufficient equilibration of the system with regard to the membrane. An additional 500 ns of simulation (5 repetitions) and the calculated RMSD values for the Kir6.2 tetramer and SUR1 protein (C, black and cyan respectively) indicate fairly good conformational stability of our system. Root mean square fluctuations for the N-terminal residues of Kir6.2 (D) located in the SUR1 pocket adjacent to the docked Aekatperone show that the first three amino acids of KNtp buried in the SUR1 pocket retained some conformational freedom and freely sampled the available space.
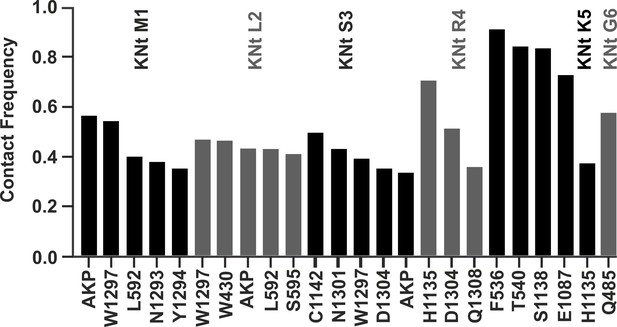
Close contact frequency between first six residues of KNtp (labeled as KNt followed by residue number) and its surroundings in the Aekatperone (AKP) binding pocket.
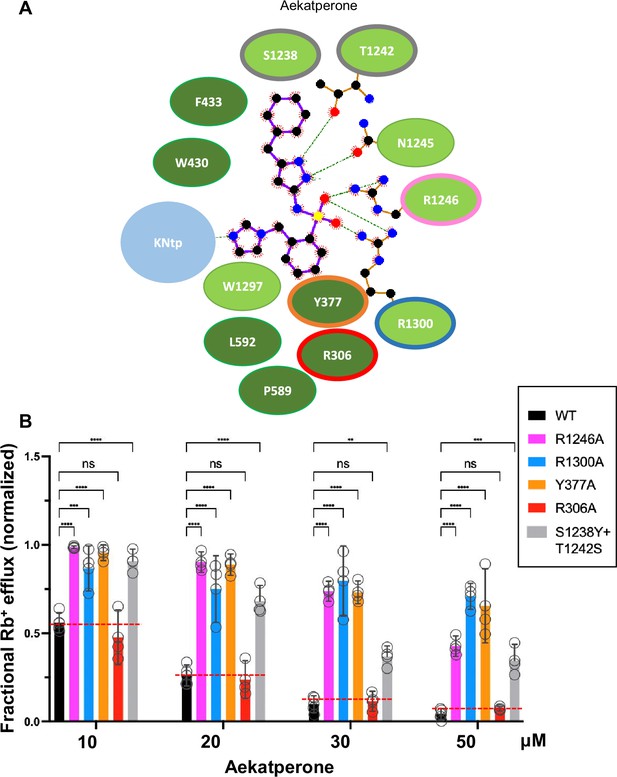
Mutagenesis studies supporting the Aekatperone binding site model derived from cryo-electron microscopy (cryoEM).
(A) 2D Aekatperone binding site model showing chemical interactions between the compound and surrounding SUR1 residues and KNtp. (B) Rb+ efflux experiments testing the effect of mutating select binding site residues on channel response to Aekatperone. Each bar is the mean and error bars represent the SEM of at least three independent experiments (individual data points shown as circles). Statistical significance is based on one-way ANOVA with Dunnett’s post hoc multiple comparisons test, with alpha = 0.05. **p<0.01, ***p<0.001, ****p<0.0001.
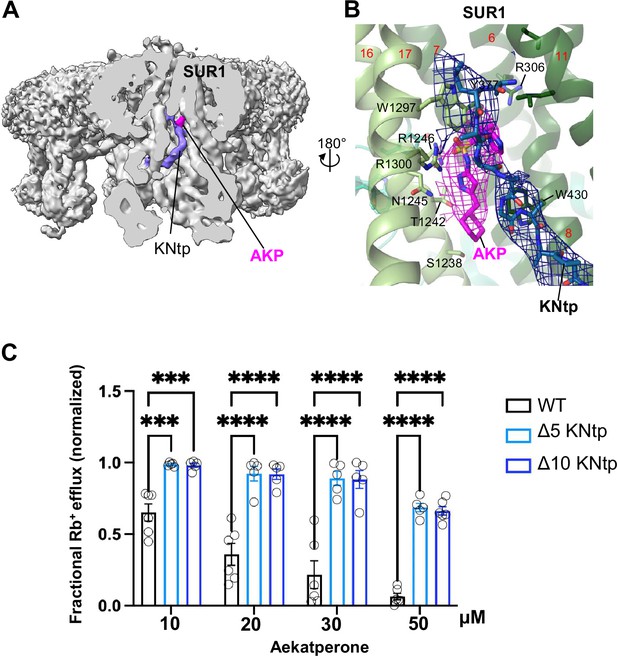
KNtp plays a role in Aekatperone-induced inhibitory effects on pancreatic KATP channel.
(A) Cryo-electron microscopy (cryoEM) density map of the pancreatic KATP channel in complex with Aekatperone (AKP), with the SUR1 subunit in the front shown in vertical sliced view. KNtp is highlighted in blue and Aekaperone in magenta. (B) Close-up view of the AKP (magenta) and KNtp (blue) structures overlayed with corresponding cryoEM densities (at 0.08 V) showing the close proximity of the imidazole group of Aekatperone to KNtp. (C) Comparison of the effects of Aekatperone on COSm6 cells expressing either wild-type (WT) KATP channels or channels with KNtp deletions of either 5 or 10 amino acids (Δ5 or Δ10 KNtp) using the Rb+ efflux assay. Rb+ efflux assays were performed in the presence of metabolic inhibitors with various concentrations of Aekatperone as indicated on the x-axis. The data were normalized against the fractional Rb+ efflux of untreated cells expressing either WT or KNtp mutant channels with metabolic inhibition but without Aekatperone. Each bar is the mean and error bars represent the SEM of five to six independent experiments (individual data points shown as light black circles). Statistical significance is based on one-way ANOVA and Tukey’s post hoc multiple comparisons test. Alpha = 0.05. **p<0.01, ***p<0.001, ****p<0.0001.
Additional files
-
Supplementary file 1
List of predicted binders that were subjected to biochemical and functional testing.
- https://cdn.elifesciences.org/articles/103159/elife-103159-supp1-v1.docx
-
Supplementary file 2
Cryo-electron microscopy (cryoEM) and model statistics of the Aekatperone-bound SUR1/Kir6.2 structure.
- https://cdn.elifesciences.org/articles/103159/elife-103159-supp2-v1.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/103159/elife-103159-mdarchecklist1-v1.docx





