A novel gene ZNF862 causes hereditary gingival fibromatosis
Figures
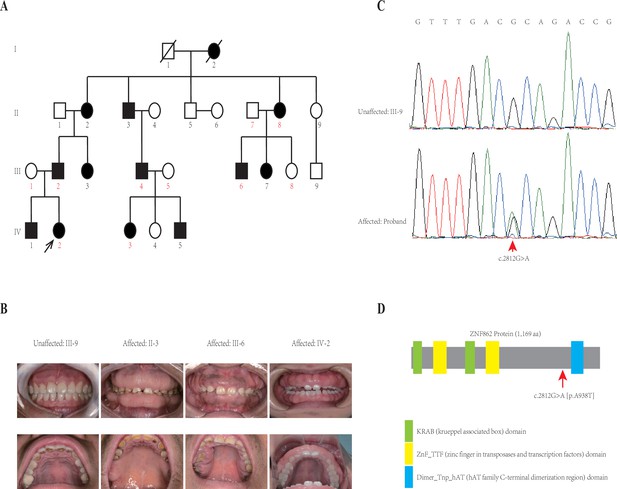
Pedigree and co-segregation analysis.
(A) The black arrow indicates the proband. The affected individuals are indicated with black-filled boxes in this family. Whole-exome sequencing (WES) was performed for the proband and nine other members (numbers indicated in red color). (B) The photographs of gingival overgrowth are revealed by intraoral examination in affected members compared with in the unaffected member in this family. (C) In this family, all affected individuals harbor the heterozygous variant (c.2812G > A) whereas the unaffected individuals are wild-type. (D) Schematic structure with putative domains of ZNF862 protein and the localization of the novel variant (red arrows). Green rectangles indicate KRAB (krueppel-associated box) domains; yellow rectangles indicate ZnF_TTF (zinc finger in transposases and transcription factors) domains; blue rectangle indicates Dimer_Tnp_hAT (hAT family C-terminal dimerization region) domain.
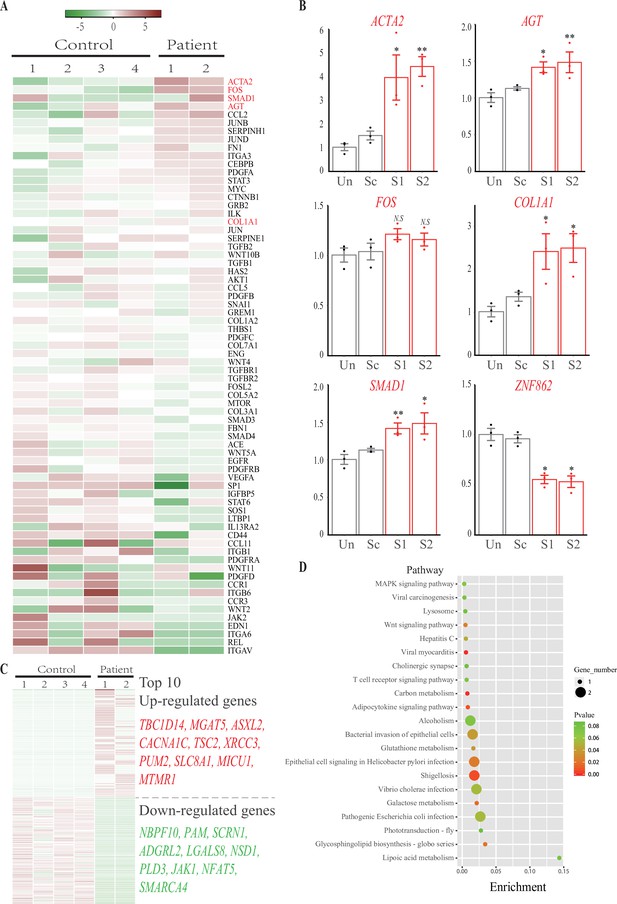
RNA sequencing and knockdown analysis.
(A) Heatmap of the 70 attested profibrotic genes expression profiling in RNA sequencing. Rows represent genes and columns represent samples. The heatmap is color-coded based on an average normalization; red represents high expression value and green represents low expression value. (B) Real-time PCR analysis of ACTA2, FOS, SMAD1, AGT, COL1A1, and ZNF862 mRNA abundance in untreated (Un), adenovirus delivered scrambler (Sc), adenovirus delivered ZNF862 short hairpin RNA (shRNA) 1 (S1), and adenovirus delivered ZNF862 shRNA 2 (S2) groups, respectively. Results are given as means ± SEM, dots indicate the relative values of gene expression levels. *p < 0.05, **p < 0.01 in comparison with Sc group. N.S. means not significant. (C) Heatmap of the 100 most up-regulated differentially expressed (DE) genes and 100 most down-regulated DE genes expression profiling in RNA sequencing. The color key is the same with (A). (D) Functional annotation chart of DE genes, the 20 most significantly enriched functions of DE genes were plotted. Functional pathways enrichment tests were based on KEGG database. Enrichment denotes the proportion of DE genes among the indicated pathway.
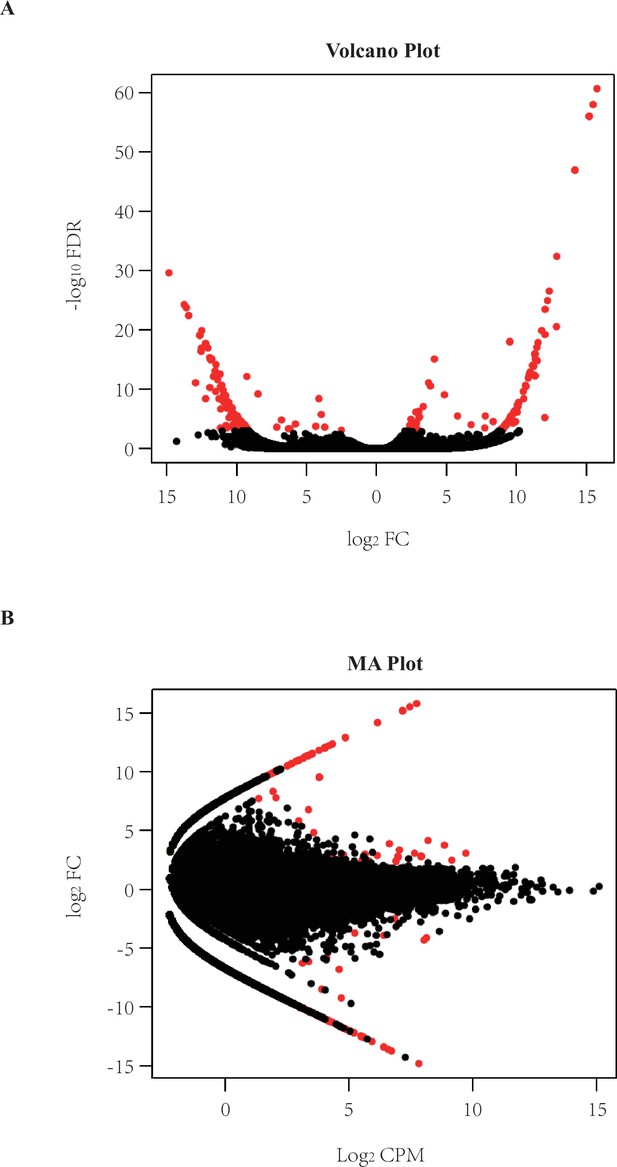
Transcriptome comparison between hereditary gingival fibromatosis (HGF) patients and controls.
(A) The transcriptomic comparison of HGF patients and controls yielded 597 differentially expressed (DE) transcripts (red dots) with a flexible cut-off (|log2 fold-change (FC)| ≥ 1.0 and false discovery rate (FDR) < 0.05), of which 355 up-regulated while 242 down-regulated DE transcripts were posited. The FC correlation and statistical significance is shown in the volcano plot. (B) The FC correlation and log2 counts per million mapped reads (CPM) is shown in the MA plot.
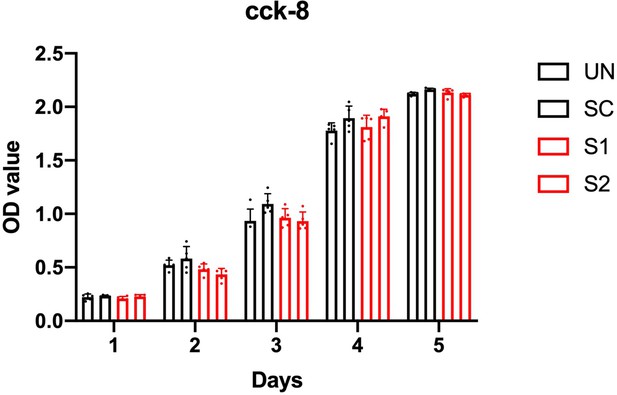
The proliferation of gingival fibroblasts.
CCK8 proliferation assay in untreated control (UN), adenovirus delivered scrambler (SC), adenovirus delivered ZNF862 short hairpin RNA (shRNA) 1 (S1), and adenovirus delivered ZNF862 shRNA 2 (S2) fibroblasts, respectively. Results are given as means ± SEM. *p < 0.05, **p < 0.01 in comparison with control group.
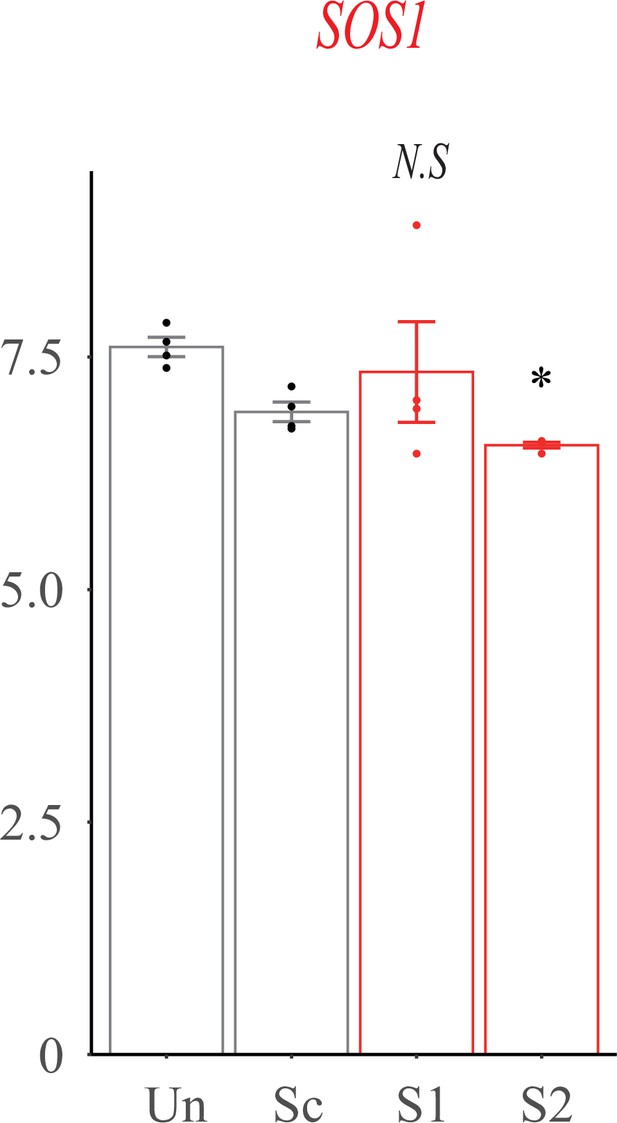
SOS1 expression in gingival fibroblasts.
Real-time PCR analysis of SOS1 mRNA abundance in untreated (Un), adenovirus delivered scrambler (Sc), adenovirus delivered ZNF862 short hairpin RNA (shRNA) 1 (S1), and adenovirus delivered ZNF862 shRNA 2 (S2) groups, respectively. Results are given as means ± SEM, dots indicate the relative values of gene expression levels. *p < 0.05, **p < 0.01 in comparison with Sc group. N.S. means not significant.
Tables
Clinical Characteristics of the individuals in this family.
NA = Not available.
| Members | Identified variant | Age at last exam | Gender | Gingival fibromatosis | Age of onset | Exposure to medication | Gingivectomy/recurrence | Other clinical finding |
|---|---|---|---|---|---|---|---|---|
| Ⅱ–1 | Wild type | 67 years | Male | − | – | No | -/- | No |
| Ⅱ–2 | p.A938T (ZNF862) | 70 years | Female | + | NA | No | -/- | Hypertension |
| II-3 | p.A938T (ZNF862) | 66 years | Male | + | NA | No | -/- | No |
| II-4 | Wild type | 66 years | Female | − | – | No | -/- | No |
| II-5 | Wild type | 64 years | Male | − | – | No | -/- | No |
| II-6 | Wild type | 64 years | Female | − | – | No | -/- | No |
| Ⅱ–7 | Wild type | 62 years | Male | − | – | No | -/- | No |
| Ⅱ–8 | p.A938T (ZNF862) | 60 years | Female | + | NA | No | -/- | No |
| Ⅱ–9 | Wild type | 54 years | Female | − | – | No | -/- | No |
| Ⅲ–1 | Wild type | 49 years | Female | − | – | No | -/- | No |
| Ⅲ–2 | p.A938T (ZNF862) | 47 years | Male | + | NA | No | -/- | No |
| Ⅲ–3 | p.A938T (ZNF862) | 45 years | Female | + (mild) | NA | No | -/- | No |
| Ⅲ–4 | p.A938T (ZNF862) | 41 years | Male | + | NA | No | -/- | No |
| Ⅲ–5 | Wild type | 36 years | Female | − | – | No | -/- | No |
| Ⅲ–6 | p.A938T (ZNF862) | 36 years | Male | + | NA | No | -/- | No |
| III-7 | p.A938T (ZNF862) | 34 years | Female | + | NA | No | -/- | No |
| III-8 | Wild type | 32 years | Female | − | – | No | -/- | No |
| Ⅲ–9 | Wild type | 30 years | Male | − | – | No | -/- | No |
| Ⅳ–1 | p.A938T (ZNF862) | 26 years | Male | + | 6–7 years | No | +/- | No |
| Ⅳ–2 | p.A938T (ZNF862) | 22 years | Female | + | 6–7 years | No | +/+ (mild) | No |
| Ⅳ–3 | p.A938T (ZNF862) | 13 years | Female | + | 2–3 years | No | -/- | No |
| Ⅳ–4 | Wild type | 11 years | Female | − | – | No | -/- | No |
| Ⅳ–5 | p.A938T (ZNF862) | 7 years | Male | + | 2–3 years | No | -/- | No |
Additional files
-
Supplementary file 1
The genes expression profiling and changes in patients over controls based on RNA sequencing.
- https://cdn.elifesciences.org/articles/66646/elife-66646-supp1-v3.xlsx
-
Supplementary file 2
A list of the 100 most up-regulated differentially expressed (DE) genes and 100 most down-regulated DE genes.
- https://cdn.elifesciences.org/articles/66646/elife-66646-supp2-v3.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/66646/elife-66646-transrepform1-v3.docx
-
Source code 1
Perl Code.
- https://cdn.elifesciences.org/articles/66646/elife-66646-code1-v3.zip





