A cell wall synthase accelerates plasma membrane partitioning in mycobacteria
Figures
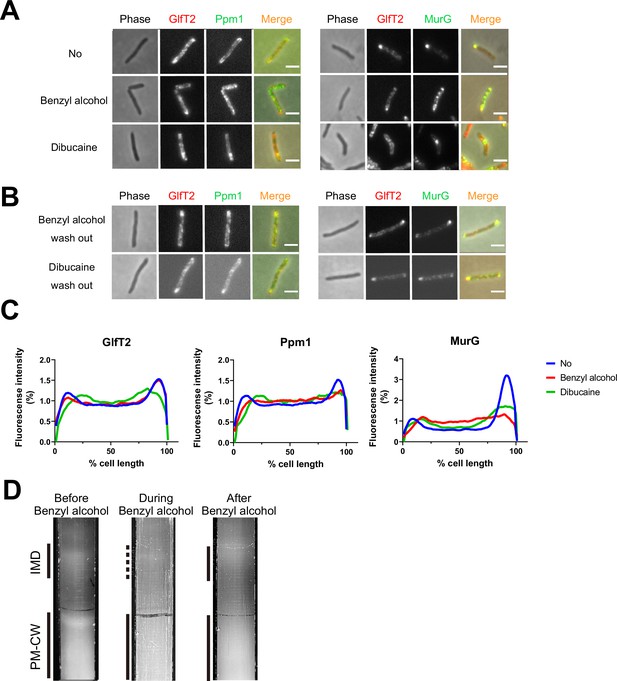
Delocalization of inner membrane domain (IMD) proteins by membrane-disrupting chemicals in M. smegmatis.
mCherry-GlfT2, Ppm1-mNeonGreen, and MurG-Dendra2 are functional fluorescent protein fusions to well-established, IMD-associated proteins (Hayashi et al., 2016; Hayashi et al., 2018; García-Heredia et al., 2021). IMD proteins were imaged after 1 hr benzyl alcohol or 1 hr dibucaine treatment, (A), and again 12 hr after washout, (B). Pictures are representative of three independent experiments. Scale bars, 2.5 μm. (C) Fluorescence distributions of the fusion proteins after chemical treatment were calculated from three independent experiments. Lines show the average of all cells (50 < n < 75). Signal was normalized to cell length and total fluorescence intensity. Cells were oriented such that the brighter poles are on the right-hand side of the graph. See ‘Materials and methods’ for details. (D) Lysates from M. smegmatis taken before, during, and 3 hr after benzyl alcohol treatment were sedimented in sucrose density gradients and imaged.
-
Figure 1—source data 1
Datasets for plot profiles of inner membrane domain (IMD) protein localizations.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig1-data1-v2.zip
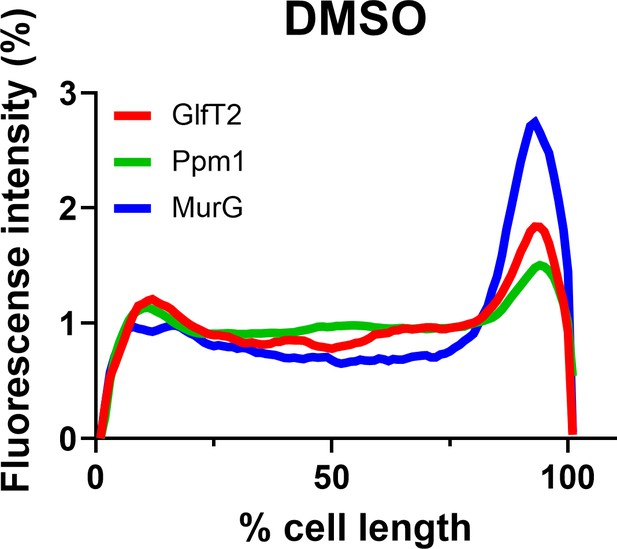
The localization of inner membrane domain (IMD) proteins was not changed by Dimethyl sulfoxide (DMSO) in M. smegmatis (compare to Figure 1C).
mCherry-GlfT2, Ppm1-mNeonGreem, and MurG-Dendra2 are functional fluorescent protein fusions to well-established, IMD-associated proteins (García-Heredia et al., 2021; Hayashi et al., 2018; Hayashi et al., 2016). Lines show the average of total cells (n = 102).
-
Figure 1—figure supplement 1—source data 1
Dataset for plot profile.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig1-figsupp1-data1-v2.zip
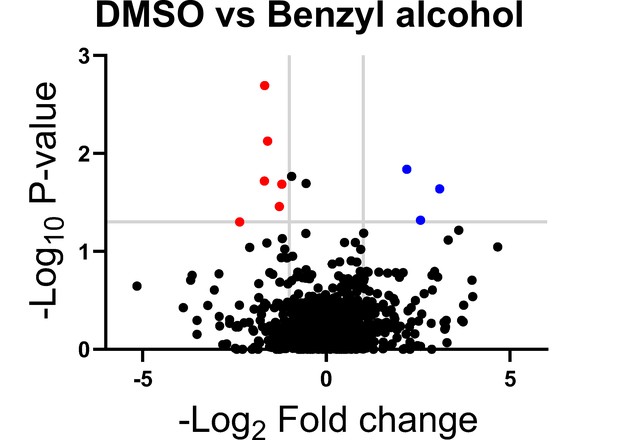
Genes in which transposon insertions are under (red, left) or over (blue, right) represented in M. smegmatis exposed to benzyl alcohol relative to DMSO vehicle control.
A transposon library was treated with benzyl alcohol or DMSO for 1 hr. Benzyl alcohol was washed away and bacteria were resuspended in Middlebrook 7H9 growth medium. The OD600 was then adjusted to 0.01 and bacteria were incubated for an additional 16–24 hr to an OD600 of ~1.0. The library was then collected for DNA sequencing. Transposon insertion counts presented relative to counts ratio (treated/control) per gene and the corresponding p-values calculated by Mann–Whitney U-test (y-axis) from n = 3 independent experiments. The horizontal gray line indicates p<0.05; the vertical gray lines indicate two-fold change.
-
Figure 2—source data 1
Raw analysis data of Tnseq.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig2-data1-v2.zip

Modest contribution of DivIVA to M. smegmatis recovery from benzyl alcohol.
Left, wild-type M. smegmatis, and right, DivIVA depletion depletion occurs upon addition of anhydrotetracycline (aTc; García-Heredia et al., 2021; Meniche et al., 2014). Following exposure to benzyl alcohol or DMSO vehicle control, M. smegmatis were washed and resuspended in growth medium in the absence of anhydrotetracycline (aTc). Cells were adjusted to OD600 0.01 and incubated ± 100 ng/mL aTc and 0.0015% of resazurin. Data combined from two independent experiments performed with six technical replicates each. Similar results were obtained when cells were pre-incubated with aTc for 2 hr prior to benzyl alcohol exposure (not shown).
-
Figure 2—figure supplement 1—source data 1
Dataset for resazurin growth curve.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig2-figsupp1-data1-v2.zip
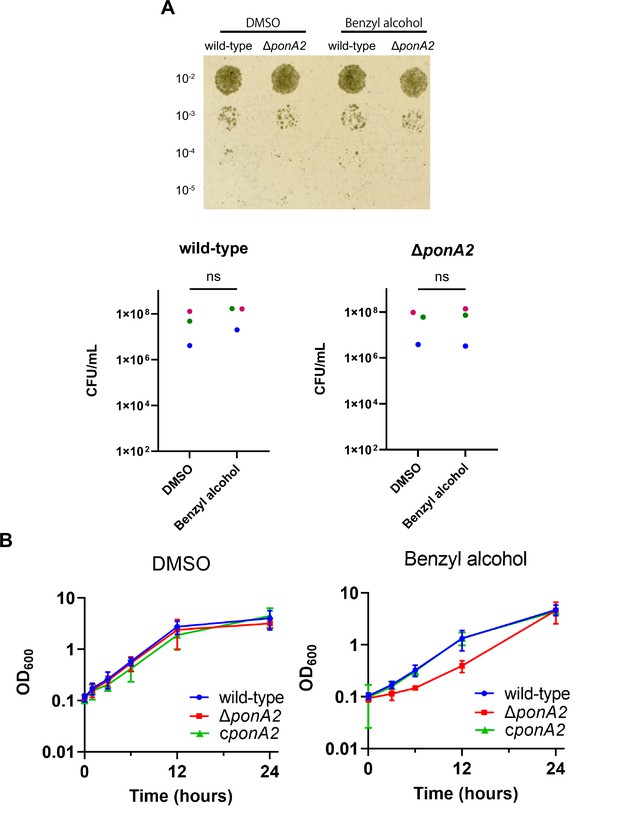
PonA2 contributes growth recovery from benzyl alcohol.
(A) Top, wild-type or ΔponA2 M. smegmatis were treated with benzyl alcohol for 1 hr, then 10-fold serial dilutions were spotted on Middlebrook 7H10 agar. The image is representative of three independent experiments. Bottom, colony-forming units (CFUs) were calculated from three biological replicates. Colors correspond to same-day replicates. ns, no statistically significant difference by Mann–Whitney U-test. p=0.4 or p>0.99 respectively. (B) Benzyl alcohol- or DMSO vehicle-treated wild-type, ΔponA2, or complemented strain (cponA2) were washed three times then grown in Middlebrook 7H9 medium. Lines show the average and SD obtained from three independent experiments.
-
Figure 3—source data 1
Colony-forming unit (CFU) and OD600 value for growth curves.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig3-data1-v2.zip

PonA2 is dispensable for survival during benzyl alcohol treatment.
Wild type, ΔponA2, or complemented M. smegmatis (cponA2) were treated with benzyl alcohol at indicated concentrations for 1 hr, then 10-fold serial dilutions were spotted on Middlebrook 7H10 agar. Image is representative of two independent experiments.
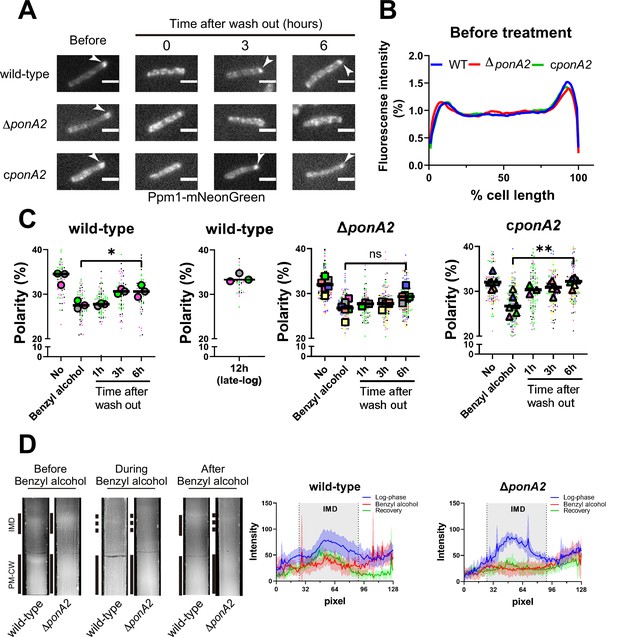
PonA2 promotes membrane repartitioning after benzyl alcohol treatment.
(A) Fluorescence imaging of M. smegmatis expressing Ppm1-mNeonGreen before benzyl alcohol treatment or after benzyl alcohol washout. Arrowheads indicate subpolar foci of Ppm1-mNeonGreen. Scale bars, 2.5 µm. Pictures are representative of three independent experiments. (B) Fluorescence of cells imaged as in (A) were quantitated from three independent experiments as in Figure 1C. Lines show the average of all cells (50 < n < 69). (C) The percentage of signal associated with the distal 15% of rod-shaped cells is quantified to indicate polarity of fluorescence distribution. Each color in the super plots (Lord et al., 2020) represents an independent biological replicate. Smaller symbols are the polarities of each cell, and larger symbols are the means of the polarity in each replicate. Statistical significance was determined by the Kruskal–Wallis test, followed by Dunn’s multiple-comparison test. ns, no statistically significant difference (p=0.3313); *p=0.0130; **p=0.0037. Data were obtained from three or six independent experiments. (D) Lysates from M. smegmatis at indicated time points were sedimented in a sucrose density gradient. Representative images of the collection tubes after sucrose gradient fractionation at left. Densitometry of the membranous material (highlighted next to the tubes with solid and dashed gray lines) shown in the right panel. The lines are the average pixel values derived from three distinct lines across images obtained from a representative experiment. The lighter areas are the standard deviations. Images of wild-type sucrose gradient fractionations are repeated from Figure 1D for clarity.
-
Figure 4—source data 1
Raw values for plot profiles of inner membrane domain (IMD) protein, raw values for the super plots, and raw values of densitometry.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig4-data1-v2.zip

Following exposure to benzyl alcohol or DMSO vehicle control, cells were washed, resuspended in Middlebrook 7H9, and incubated with 0.0015% of resazurin.
Data were obtained from three independent experiments and are means of biological duplicates or triplicates. Data for wild-type and ∆ponA2 are reproduced from Figure 6A for comparison.
-
Figure 4—figure supplement 1—source data 1
Dataset for resazurin growth curve of mutants.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig4-figsupp1-data1-v2.zip
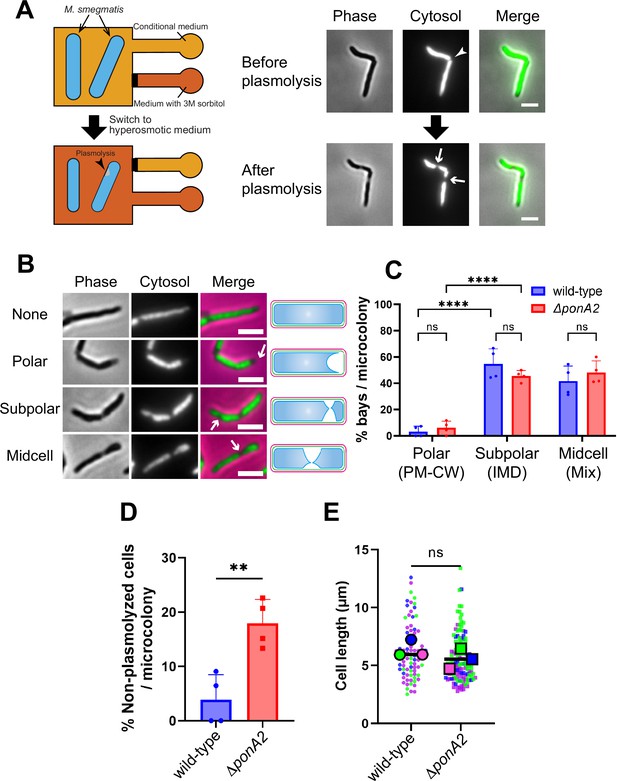
Membrane–cell wall interactions are weaker in subpolar regions.
(A) Left, schematic of the microfluidics plasmolysis assay. GFP-expressing M. smegmatis were subjected to hyperosmotic shock (3 M sorbitol) in microfluidics device and imaged before, during, and after shock. Right, images before and after hyperosmotic shock. Cells were perfused with 7H9 medium for 5 min and then perfused with 7H9 + 3 M sorbitol for 5 min. Cells were imaged in phase (50 ms) and GFP (50 ms) every 30 s. An arrowhead indicates the septum while arrows indicate sites of plasmolysis. Scale bars, 2.5 μm. Images are representative of four independent experiments. (B) Cytoplasmic GFP enables visualization and quantitation of plasmolysis bays. Arrows indicate representative sites of plasmolysis. Scale bars, 2.5 μm. Images are representative of two independent experiments. (C) Analysis of the sites of plasmolysis (as defined in [B]) in wild-type (n = 85) and ΔponA2 (n = 101) M. smegmatis from two biological replicates performed in duplicate. For this analysis, we removed cells that did not plasmolyse. Statistical significance was determined by the two-way ANOVA test, followed by Šídák’s multiple-comparisons test. ****p<0.0001, ns, p>0.99. Polar, subpolar, and midcell bays spatially correlate with the PM-CW, inner membrane domain (IMD), and a mixture of PM-CW and IMD, respectively (García-Heredia et al., 2021; Hayashi et al., 2018; Hayashi et al., 2016; Prithviraj et al., 2023). (D) The proportion of cells that did not plasmolyse in (C) was calculated for wild-type (n = 90) and ΔponA2 (n = 123) M. smegmatis. Statistical significance was determined by the Mann–Whitney U-test. **p=0.045. (E) Cell lengths were analyzed for wild-type (n = 84) and ΔponA2 (n = 132) M. smegmatis from three biological replicates. Each color in the super plots (Lord et al., 2020) represents an independent biological replicate. Smaller symbols are the lengths of individual cells, and larger symbols are the means of each replicate. Statistical significance was determined by the Mann–Whitney U-test. ns, p=0.4.
-
Figure 5—source data 1
Raw values of the proportions of where plasmolysis bay were seen, population without plasmolysis, and cell length.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig5-data1-v2.zip
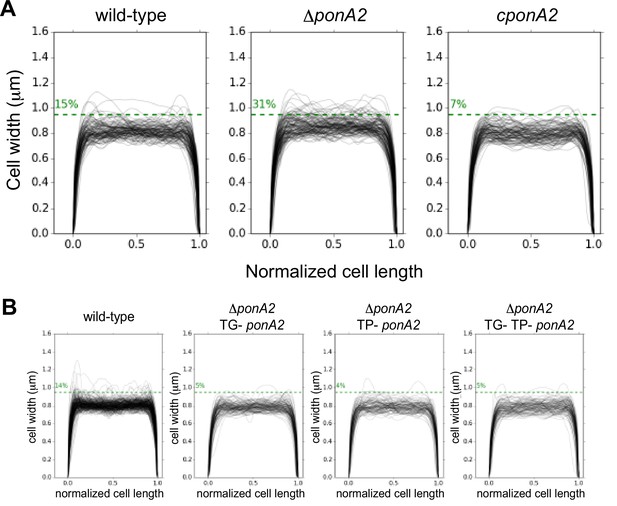
M. smegmatis width profiles of cells along the normalized cell length were determined by Oufti (Nguyen et al., 2007; Paintdakhi et al., 2016) and a Python script here (copy archived at Sparks, 2023).
(A) Cell widths of wild-type (n = 116), ∆ponA2 (n = 124), and complemented ∆ponA2 (cponA2; n = 107). (B) Cell widths of wild-type (n = 278), ∆ponA2 complemented with TG- ponA2 (n = 120), TP- ponA2 (n = 101), or TG- TP- ponA2 (n = 109) were quantified from three independent experiments.
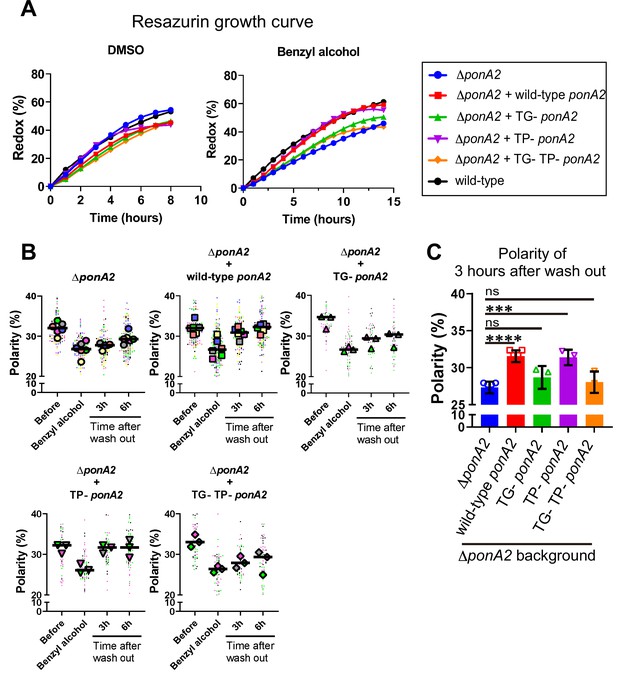
Transglycosylase domain of PonA2 accelerates regrowth and membrane repartitioning post-benzyl alcohol.
(A) Following exposure to benzyl alcohol or the DMSO vehicle control, cells were washed, resuspended in Middlebrook 7H9, and incubated with 0.0015% of resazurin. Data were obtained from three independent experiments and are means of biological duplicates or triplicates. (B) Inner membrane domain (IMD) marker (Ppm1-mNeonGreen) polarities were assessed in mutants before, during, or 3- or 6 hr after benzyl alcohol treatment as in Figure 4C. Data from ΔponA2 and ΔponA2 complemented with wild-type ponA2 are repeated from Figure 4C for ease of comparison. Each color in the super plots (Lord et al., 2020) represents an independent biological replicate. Smaller symbols are the polarities of each cell, and larger symbols are the means of each replicate. (C) The polarity of 3 hr time points from (B) were compiled and compared across mutants. The bar graph shows the average and standard deviation. Statistical significance was determined by the Kruskal–Wallis test, followed by Dunn’s multiple-comparison test. ns, no statistically significant difference (p=0.2538 or 0.7601 respectively); ****p<0.0001; ***p=0.0002.
-
Figure 6—source data 1
Raw values of resazurin growth curve, super plots, and polarity of 3 hr after wash out.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig6-data1-v2.zip

Protein alignments of PBP1a of E. coli and PonA1 and PonA2 of M. smegmatis.
As in M. smegmatis ponA1 (Kieser et al., 2015b), the true start codon of ponA2 is located in the upstream of the annotation in MycoBrowser (Kapopoulou et al., 2011). That is, M. smegmatis ponA2 starts from 87 base pairs upstream of MSMEG_6201. Red shading indicate conserved amino acids. Arrows and blue squares indicate the amino acids that were mutated. Amino acid sequences were aligned using Jalview2 (Waterhouse et al., 2009).
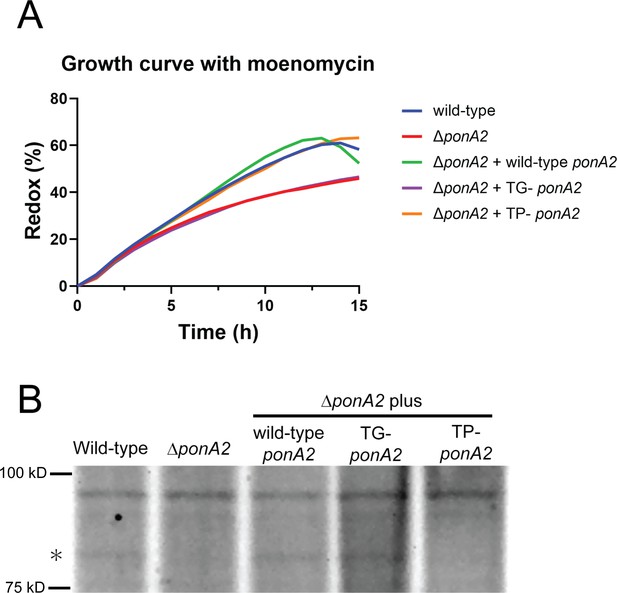
Enzymatic activity of catalytic inactive mutants.
(A) Mutation in conserved ponA2 transglycosylase domain sensitizes M. smegmatis to moenomycin. Log-phase bacteria were adjusted to OD600 = 0.1 and incubated with 15.625 mg/mL moenomycin and 0.0015% of resazurin. Representative data from two independent experiments. (B) Mutation in conserved ponA2 transpeptidase domain abolishes transpeptidase activity. 30 μg of whole-cell lysates were incubated with 100 nmol of Bocillin-FL for 30 min at 37℃ then separated by SDS-PAGE. Gel fluorescence was visualized by Amersham ImageQuant 800 system (Cytiva). Star at left indicates PonA2.
-
Figure 6—figure supplement 2—source data 1
Dataset for resazurin growth curve of moenomycin-treated mutants.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig6-figsupp2-data1-v2.zip
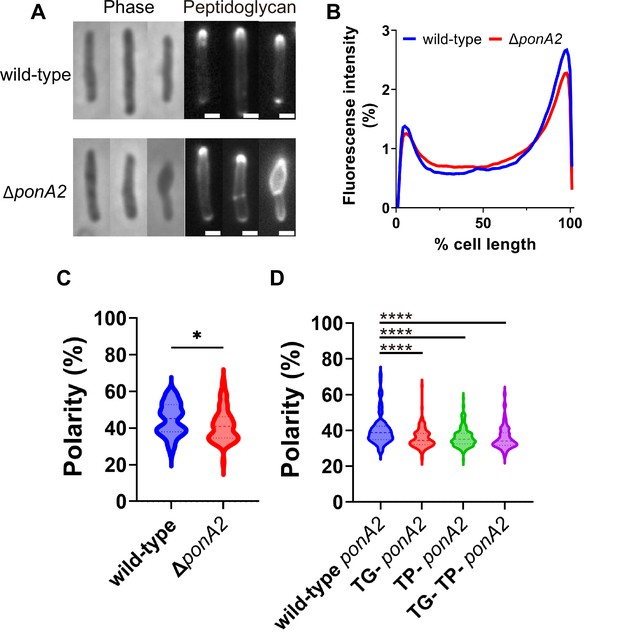
PonA2 localizes peptidoglycan synthesis and maintains cell morphology.
(A) Nascent peptidoglycan in wild-type or ∆ponA2 M. smegmatis was labeled for 15 min (~10% generation) with alkyne-d-alanine-d-alanine (alkDADA) followed by copper-catalyzed alkyne-azide cycloaddition (CuAAC) with AFDye488 Azide. Scale bars, 1 µm. Pictures are representative of three independent experiments. (B) Wild-type and ∆ponA2 strains were labeled as in (A) and subcellular fluorescence was quantitated as in Figure 1C. Lines show the average of all cells (50 < n < 68) obtained by three independent experiments. (C) The percentage of signal associated with the distal 15% of rod-shaped cells quantified to indicate polarity of fluorescence distribution. Statistical significance was determined by Mann–Whitney U-test. p-value, *p=0.0227. (D) Polarity ratio of catalytically inactive mutants. Statistical significance was determined by the Kruskal–Wallis test, followed by Dunn’s multiple-comparison test. Data were obtained from the three independent experiments. ****p<0.0001.
-
Figure 7—source data 1
Raw values of plot profiles and polarity of peptidoglycan synthesis.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig7-data1-v2.zip
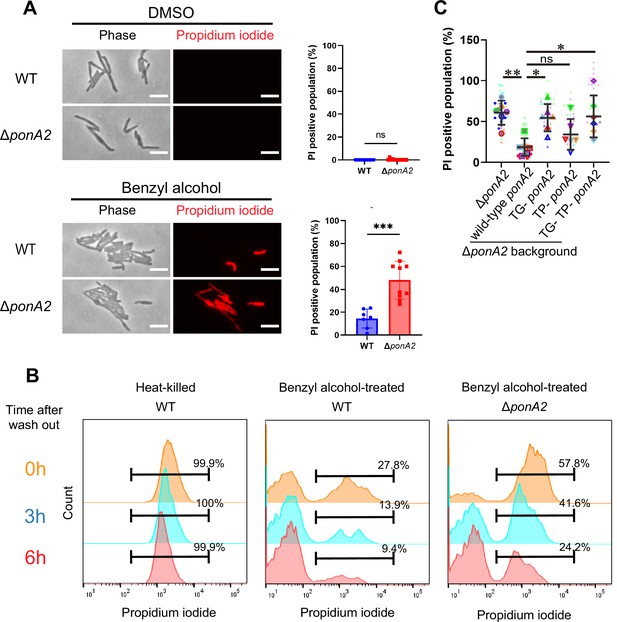
PonA2 promotes cell impermeability upon benzyl alcohol treatment.
(A, left) Benzyl alcohol- or DMSO-treated wild-type or ΔponA2 strains were stained with propidium iodide to detect membrane-compromised cells. Scale bars, 5 µm. Images are representative of three independent experiments. (A, right) Propidium iodide-positive cells in individual fields of wild-type (n = 160) or ΔponA2 (n = 327) were enumerated. Mann–Whitney U-test. ns, not statistically significant (p=0.4805); ***p=0.0002. Data were combined from two independent experiments. (B) Flow cytometry analysis of propidium iodide-positive cells in time course after benzyl alcohol washout. Cells were incubated at 65℃ for 1 hr for heat-killed control. Data are representative of three independent experiments. (C) ∆ponA2 M. smegmatis complemented with wild-type ponA2 or alleles with mutated transpeptidase (TP-) and/or transglycosylase (TG- and TP- TG-) domains were treated with benzyl alcohol and stained with propidium iodide. Propidium iodide staining quantitated in the same way as the right panel of (A). Data were combined from six individual experiments. The line show the average and standard deviation. Statistical significance was determined by the Kruskal–Wallis test, followed by Dunn’s multiple-comparison test. ns, not statistically significant (p=0.9001); **p=0.0033; *p=0.0235 or 0.0287. In total, 577 < n < 1410 cells were examined across five independent experiments.
-
Figure 7—figure supplement 1—source data 1
Dataset for propidium-iodide-positive population.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig7-figsupp1-data1-v2.zip
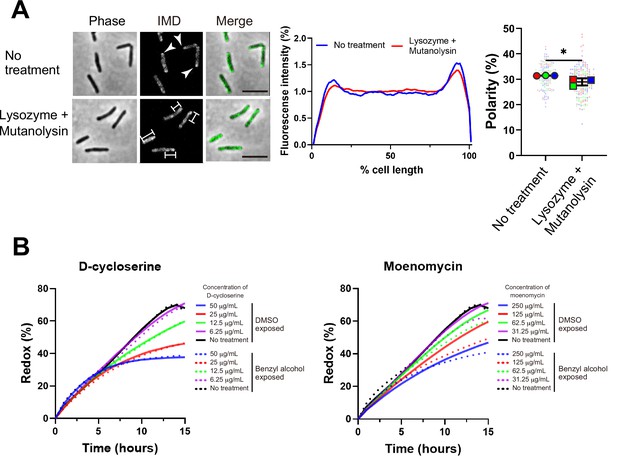
Maintenance and establishment of membrane partitioning are respectively supported by the cell wall glycan backbone and active cell wall polymerization.
(A) Left, images of M. smegmatis expressing inner membrane domain (IMD) marker Ppm1-mNeonGreen ± 60 min treatment with cell wall hydrolases lysozyme and mutanolysin. Arrowheads indicate subpolar foci of Ppm1-mNeonGreen. Bars highlight dispersed fluorescent signal of Ppm1-mNeonGreen. Scale bars, 5 µm. Middle, quantitation of Ppm1-mNeonGreen polarity for cells with no treatment (n = 134) or lysozyme/mutanolysin treatment (n = 187). Lines show the average of all cells. Right, the percentage of Ppm1-mNeonGreen signal associated with the distal 15% of rod-shaped cells is quantified to indicate polarity of fluorescence distribution. Each color in the super plots (Lord et al., 2020) represents an independent biological replicate. Smaller symbols are the polarities of each cell and larger symbols are the means of each replicate. The line shows the average and standard deviation. Data were obtained from three independent experiments. Mann–Whitney U p-value, *p=0.04. (B) Bacteria treated with benzyl alcohol (dashed line) or DMSO vehicle (solid line) were washed then grown in Middlebrook 7H9 with d-cycloserine or moenomycin at the indicated concentrations. The lines are the average of two or three biological replicates.
-
Figure 8—source data 1
Raw values of plot profiles and resazurin growth curves.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig8-data1-v2.zip

Top, images of M. smegmatis expressing inner membrane domain (IMD) marker mCherry-GlfT2 ±60 min treatment with cell wall hydrolases lysozyme and mutanolysin.
Arrowheads indicate subpolar foci of mCherry-GlfT2. Scale bars, 5 µm. Bottom, quantitation of mCherry-GlfT2 signals with no treatment (n = 475) or lysozyme/mutanolysin treatment (n = 370). Bottom left, lines show the average of total cells. Bottom right, the percentage of signal associated with the distal 15% of rod-shaped cells is quantified to indicate polarity of fluorescence distribution (details in ‘Materials and methods’). The line shows the average and standard deviation. Each color in the super plots represents an independent biological replicate. Smaller symbols are the polarities of each cell and larger symbols are the means of the polarity in each of three biological replicates. Mann–Whitney U p-value, p=0.4685.
-
Figure 8—figure supplement 1—source data 1
Dataset for plot profiles of inner membrane domain (IMD) protein and super plot of polarity.
- https://cdn.elifesciences.org/articles/81924/elife-81924-fig8-figsupp1-data1-v2.zip
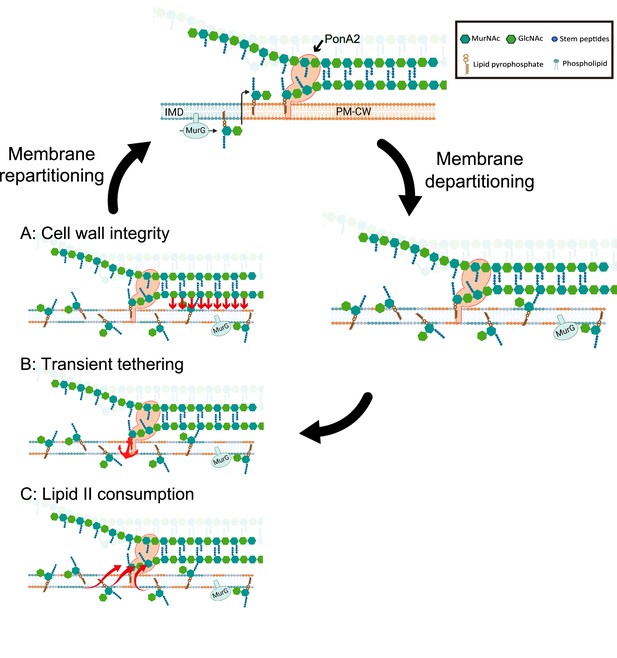
Working models for the role PonA2 in de novo plasma membrane partitioning.
Inner membrane domain (IMD) (blue) and PM-CW (orange) have distinct proteomes and lipidomes (Hayashi et al., 2016). PonA2, likely enriched in the PM-CW (García-Heredia et al., 2021; Hayashi et al., 2016), polymerizes peptidoglycan using lipid II as a donor substrate, which in turn is produced by MurG in the IMD. PonA2 also cross-links nascent glycan strands into the existing cell wall. Upon membrane fluidization, lipid II and IMD-enriched proteins are no longer confined to the IMD (García-Heredia et al., 2021). PonA2 may promote membrane repartitioning post-fluidization via: (A) a mature peptidoglycan structure that directly or indirectly interacts with the membrane (red arrows), (B) nascent peptidoglycan polymers that transiently tether the membrane to the cell wall, and/or (C) consumption of lipid II, in turn regulating membrane fluidity.
Tables
Genes identified by Tn-seq.
| Underrepresented in benzyl alcohol-treated M. smegmatis (candidates for promoting survival) | ||
|---|---|---|
| Gene locus | Gene name | Gene description |
| MSMEG_0846c | – | Putative monovalent cation/H+ antiporter subunit D |
| MSMEG_2775 | nhaA | Na+/H+ antiporter NhaA |
| MSMEG_4533c | – | Sulfate-binding protein |
| MSMEG_5488c | – | DNA-binding response regulator |
| MSMEG_5781c | pstC | Phosphate ABC transporter, permease protein PstC |
| MSMEG_6201 | ponA2 | Bifunctional transglycosylase/transpeptidase |
| Overrepresented in benzyl alcohol-treated M. smegmatis (candidates for impairing survival) | ||
| MSMEG_2768 | – | OB-fold nucleic acid binding domain-containing protein |
| MSMEG_2772 | – | Amino acid permease |
| MSMEG_5694 | – | Hypothetical protein MSMEG_5694 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Mycobacterium smegmatis mc2155) | M. smegmatis | NC_008596 in GenBank | Wild-type M. smegmatis | |
| Strain, strain background (M. smegmatis) | mCherry-GlfT2 and Ppm1-mNeonGreen | Hayashi et al., 2016 | See reference for details | |
| Strain, strain background (M. smegmatis) | mCherry-GlfT2 and MurG-Dendra2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 wild-type background | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 wild-type background complementing wild-type ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 wild-type background complementing TG- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 wild-type background complementing TP- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 wild-type background complementing TG- and TP- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 mC-GlfT2 and Ppm1-mNG background | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 mC-GlfT2 and Ppm1-mNG background complementing wild-type ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 mC-GlfT2 and Ppm1-mNG background complementing TG- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 mC-GlfT2 and Ppm1-mNG background complementing TP- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | DponA2 mC-GlfT2 and Ppm1-mNG background complementing TG- and TP- ponA2 | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | mCherry-GlfT2/DivIVA-eGFP-ID | García-Heredia et al., 2021 | See reference for details | |
| Strain, strain background (M. smegmatis) | GFP expressing wild-type | This study | The mutant was generated as described in Supplementary material and methods | |
| Strain, strain background (M. smegmatis) | GFP expressing DponA2 background of wild-type | This study | The mutant was generated as described in Supplementary material and methods | |
| Software, algorithm | MATLAB codes | García-Heredia et al., 2018; Paintdakhi et al., 2016 | RRID:SCR_001622 | Scripts designed for MATLAB to analyze the fluorescence profiles along a cell body from data collected in Oufti. |
| Other | Python script | This study | See ‘Materials and methods’ and Source code 1 for details | |
| Software, algorithm | ImageJ | Schindelin et al., 2012 | RRID:SCR_003070 | See reference for details |
| Software, algorithm | GraphPad Prism 9 | Commercially available | RRID:SCR_002798 |
Additional files
-
Supplementary file 1
Primers used in this study.
- https://cdn.elifesciences.org/articles/81924/elife-81924-supp1-v2.docx
-
Supplementary file 2
The Tn-seq result of Mann–Whitney U-test.
- https://cdn.elifesciences.org/articles/81924/elife-81924-supp2-v2.xlsx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/81924/elife-81924-mdarchecklist1-v2.pdf
-
Source code 1
Code to analyze cell width profile.
- https://cdn.elifesciences.org/articles/81924/elife-81924-code1-v2.zip





