Cylicins are a structural component of the sperm calyx being indispensable for male fertility in mice and human
Figures
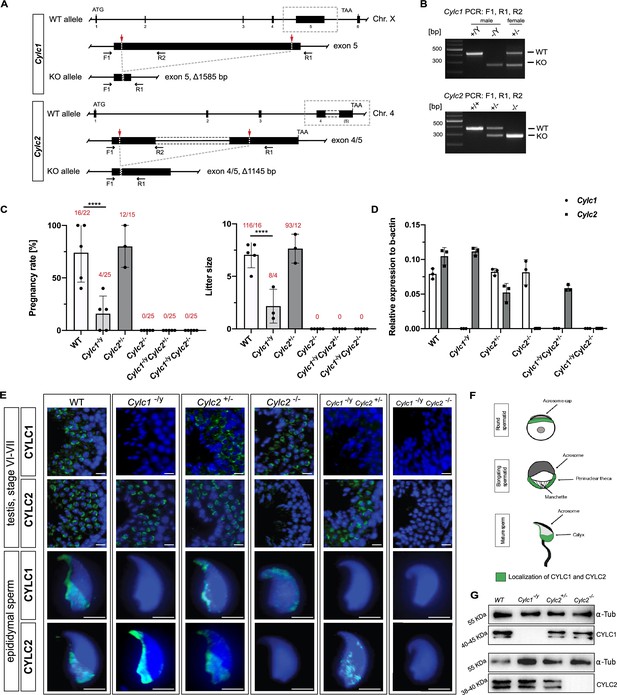
Loss of Cylc1 or Cylc2 results in impaired male fertility.
(A) Schematic representation of the Cylc1 and Cylc2 gene structure and targeting strategy for CRISPR/Cas9-mediated generation of Cylc1- and Cylc2-deficient alleles. Targeting sites of guide RNAs are depicted by red arrows. Genotyping primer binding sites are depicted by black arrows. (B) Representative genotyping PCR of Cylc1- and Cylc2-deficient mice. N=3. (C) Fertility analysis of Cylicin-deficient mice visualized by mean litter size and pregnancy rate (%) in comparison to wildtype (WT) matings. Black dots represent mean values obtained for each male included in fertility testing. Columns represent mean values ± standard deviation (SD). Total number of offspring per total number of pregnancies as well as total number of pregnancies per total number of plugs are depicted above each bar. (D) Expression of Cylc1 and Cylc2 in testicular tissue of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice analyzed by quantitative reverse transcription-polymerase chain reaction (qRT-PCR). Biological replicate of 3 was used. (E) Immunofluorescent staining of testicular tissue and cauda epididymal sperm from WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- males against CYLC1 and CYLC2. Cell nuclei were counterstained with DAPI. Staining was performed on three animals from each genotype. Scale bar: 5 µm. (F) Schematic illustration of CYLC localization during spermiogenesis. CYLC localization (green) is shown for round and elongating spermatids as well as mature sperm. (G) Representative immunoblot against CYLC1 and CYLC2 on cytoskeletal protein fractions from WT, Cylc1-/y, Cylc2+/-, and Cylc2-/- testes. α-Tubulin was used as load control.
-
Figure 1—source data 1
PCR-genotyping of Cylicin-deficient mice.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig1-data1-v2.zip
-
Figure 1—source data 2
Pregnancy rates and litter sizes of WT female mice mated to Cylicin-deficient male mice.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig1-data2-v2.zip
-
Figure 1—source data 3
Cylicin1 and Cylicin2 staining of mature sperm in testis tissues.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig1-data3-v2.zip
-
Figure 1—source data 4
Western-blot validation of the knockout.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig1-data4-v2.zip
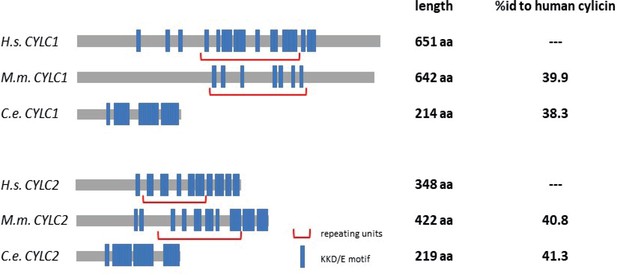
Amino acid sequence comparison of CYLC1 and CYLC2 in Caenorhabditis elegans and Mus musculus to Homo sapiens.
KKD/E motifs are highlighted in blue and repeating units are marked by red brackets.

Immunohistochemical staining against CYLC1 and CYLC2 in tissue sections of testis, brain, thymus, and spleen.
Scale bar: 100 µm.
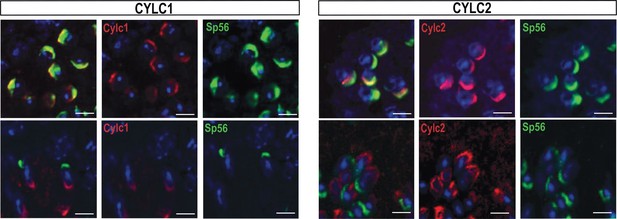
Immunofluorescence staining against the acrosomal matrix marker protein SP56 (green) and CYLC1 or CYLC2 (red) in round and elongating spermatids.
Nuclei were stained with DAPI. Scale bar: 5 µm.
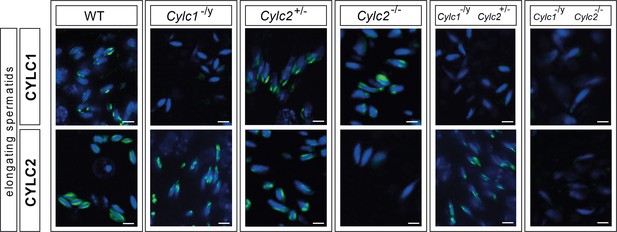
Immunofluorescence staining of CYLC1 and CYLC2 in elongating spermatids of wildtype (WT), Cylc1-/y, Cylc2 +/-, Cylc2 / - , Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice.
Scale bar: 5μm.
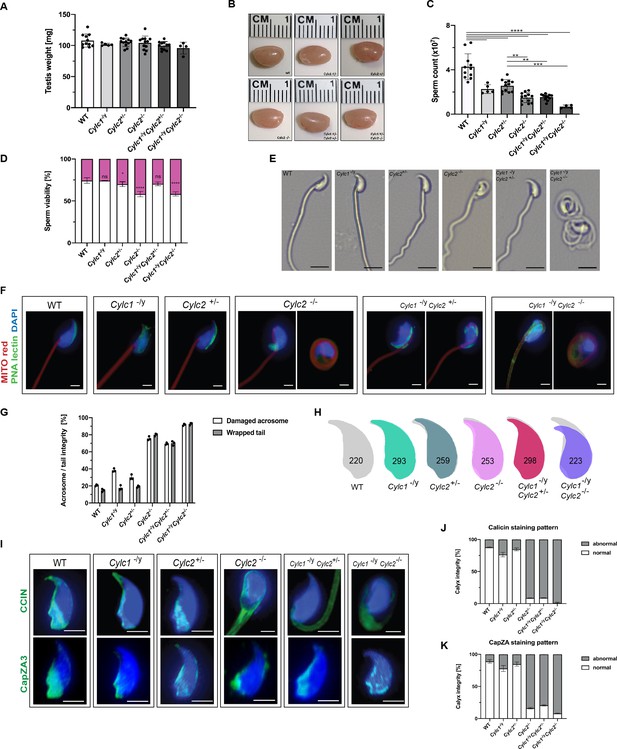
Sperm morphology is severely altered in Cylicin-deficient mice.
(A) Testis weight (mg) and sperm count (×107) of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- males. Mean values ± SD are shown; black dots represent data points for individual males. (B) Comparable photographs of the testes of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice. (C) Epididymal sperm count (×107) of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- males. Mean values ± SD are shown; black dots represent data points for individual males. (D) Viability of the epididymal sperm stained with Eosin-Nigrosin. Percentage of Eosin negative (viable) and Eosin positive (inviable) sperm is shown. Data represented as mean ± SD. Staining was performed on three animals from each genotype. (E) Bright-field microscopy pictures of epididymal sperm from WT, Cylc1-/y, Cylc2+/-, and Cylc2-/- mice. Scale bar: 10 μm. (F) Immunofluorescence staining of epididymal sperm acrosomes with peanut agglutinin (PNA) lectin (green) and tails with MITOred (red). Nuclei were counterstained with DAPI. Scale bar: 5 µm. (G) Quantification of abnormal sperm of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice is shown. Acrosome aberrations and tail coiling were counted separately. Staining was performed on three animals from each genotype. (H) Nuclear morphology analysis of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- sperm. Number of cells analyzed for each genotype is shown. (I) Representative pictures of immunofluorescent staining against perinuclear theca (PT) proteins CCIN (upper panel) and CAPZa3 (lower panel) in WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- sperm. Nuclei were counterstained with DAPI. Staining was performed on three animals from each genotype. Scale bar: 5 µm. (J–K) Quantification of sperm with abnormal calyx integrity in WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice based on CCIN and CapZA staining patterns.
-
Figure 2—source data 1
Testis weights and sperm counts.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data1-v2.zip
-
Figure 2—source data 2
Sperm viability.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data2-v2.zip
-
Figure 2—source data 3
Brightfield micrographs of mature sperm.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data3-v2.zip
-
Figure 2—source data 4
Staining and quantification of acrosomal and flagellar defects.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data4-v2.zip
-
Figure 2—source data 5
Sperm head morphology.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data5-v2.zip
-
Figure 2—source data 6
Staining of calyx proteins.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig2-data6-v2.zip
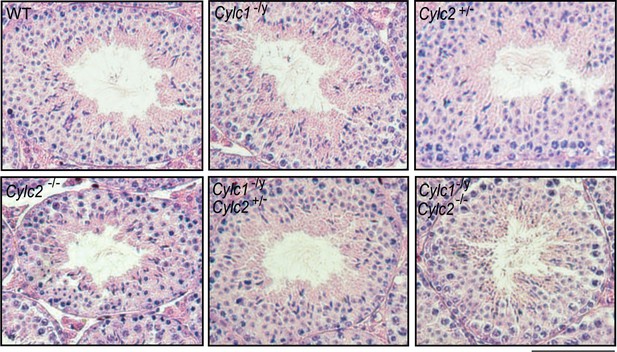
Hematoxylin and eosin (HE)-stained testicular tissue sections of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice.
Scale bar: 100 µm.
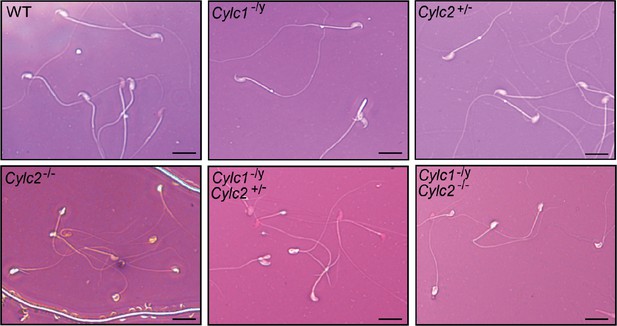
Eosin-Nigrosin staining of epididymal sperm samples from wildtype (WT), Cylc1- /y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice.
Scale bar: 10 μm.
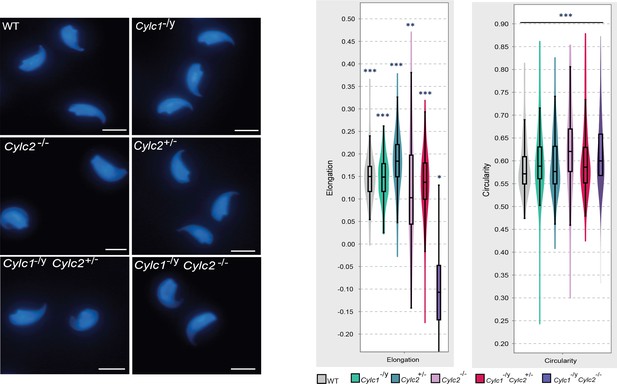
Nuclei of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- sperm stained with DAPI.
Scale bar: 5 μm. Elongation and circularity of nuclei from WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- sperm. The minimum detection area was set to 1.000 pixels, while the maximum detection area was 7.000 pixels.
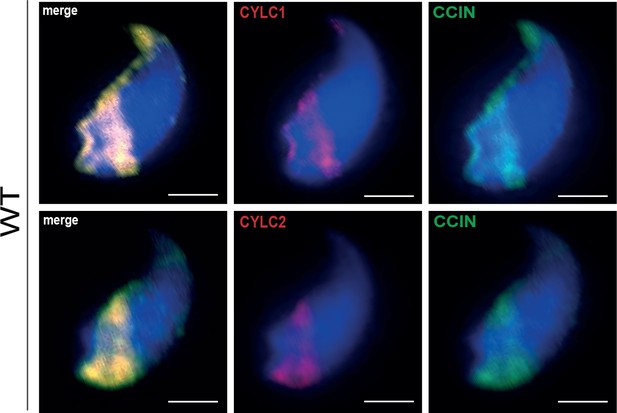
Co-staining against CYLC1/CYLC2 (red) and CCIN (green) in epididymal sperm cells of wildtype (WT) mouse.
Nuclei were counterstained with DAPI. Scale bar: 2 μm.
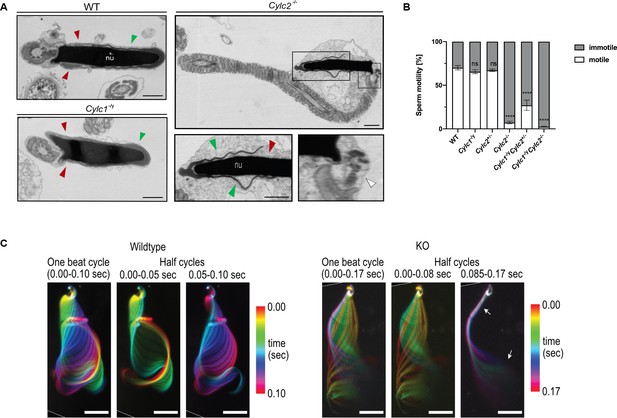
Cylc2-/- sperm cells have altered flagellar beat.
(A) Transmission electron microscopy (TEM) micrographs of wildtype (WT), Cylc1-/y and Cylc2-/- epididymal sperm. Acrosome appears detached from the nucleus in Cylc2-/- sperm (green arrowheads), while the calyx is missing entirely (red arrowheads). The head-tail connecting piece shifted from the basal plate is shown by white arrowheads causing the looping of the flagellum and formation of a cytoplasmatic sac. Cylc1-/y sperm appears comparable to WT. Scale bar: 1 µm. (B) Motility of the epididymal sperm of WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- males activated in TYH medium. (C) Full and half-beat cycle plots of the flagellar beat are shown for WT and Cylc2-/- spermatozoa. Half-beat cycle shows the stiffness of the midpiece (upper arrow) and high oscillations (lower arrow) in Cylc2-/- sperm in one direction of the beat.
-
Figure 3—source data 1
Uncropped TEM-micrographs of mature sperm.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig3-data1-v2.zip
-
Figure 3—source data 2
Sperm motility.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig3-data2-v2.zip
-
Figure 3—source data 3
Program codes for flagellar beat analysis.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig3-data3-v2.zip
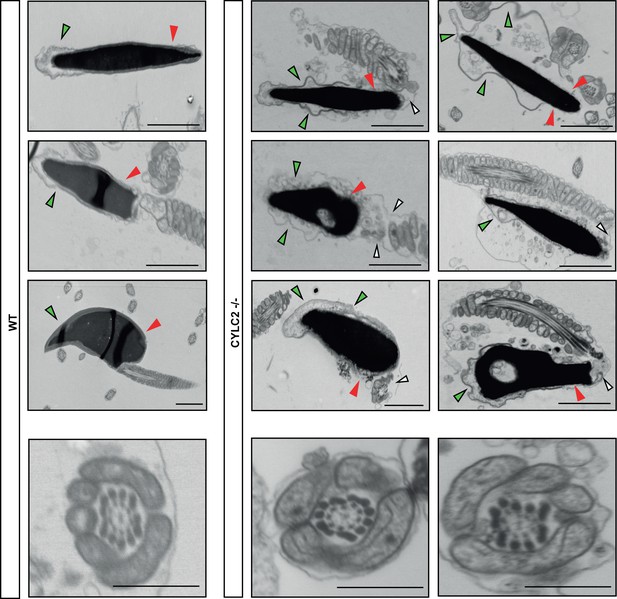
Transmission electron microscopy (TEM) micrographs of wildtype (WT) and Cylc2-/- sperm and axonemes.

SpermQ analysis of the flagellar beat of wildtype (WT) (green) and Cylc2-/- (red) sperm.
Average curvature of the flagellum and the arc length are shown.
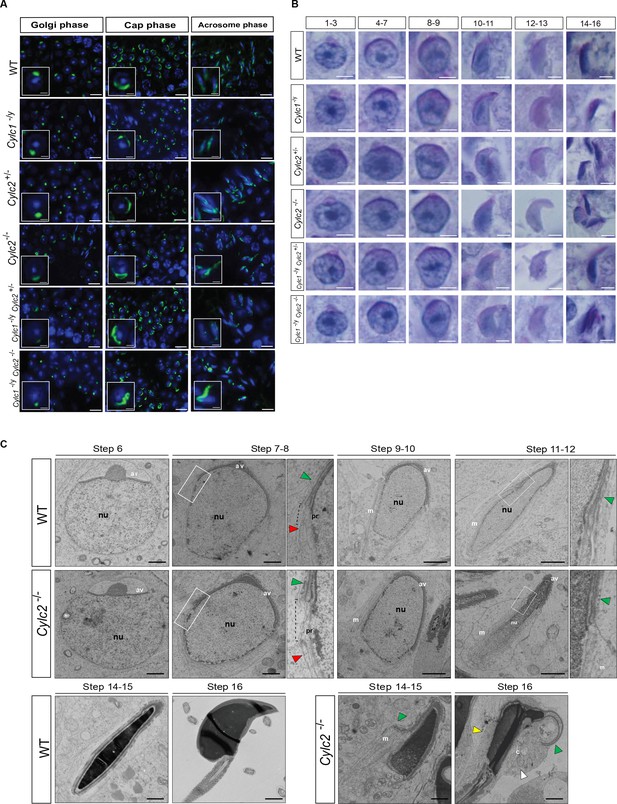
Cylicins are required for acrosome attachment to the nuclear envelope.
(A) Peanut agglutinin (PNA)-fluorescein isothiocyanite (FITC) lectin immunofluorescence staining of the acrosome in testicular tissue of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice (green). Golgi phase of acrosome biogenesis at round spermatid stage (I–IV) is visible in the left panel. Middle panel shows cap phases on round spermatids (stage V–VIII). In the right panel acrosomal phase is shown (stage IX–XI). Nuclei were counterstained with DAPI. Staining was performed on three animals from each genotype. Scale bar: 10 µm. Insets show representative single spermatids at higher magnification (scale bar: 2 µm). (B) Periodic acid Schiff (PAS) staining of testicular sections from WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice. Representative spermatids at different steps of spermiogenesis are shown. Scale bar: 3 µm. (C) Transmission electron microscopy (TEM) micrographs of testicular tissues of WT and Cylc2-/- mice. Single spermatids from step 6 to step 16 are shown. nu: nucleus; av: acrosomal vesicle; pr: perinuclear ring; m: manchette microtubules; cy: cytoplasm; green arrowheads: developing acrosome; red arrowheads: manchette; white arrowhead: cytoplasm; yellow arrowhead: remaining microtubules in mature sperm. Scale bar: 1 µm.
-
Figure 4—source data 1
PNA-lectin stained testes tissues.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig4-data1-v2.zip
-
Figure 4—source data 2
Uncropped TEM-micrographs of testes tissues.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig4-data2-v2.zip

Peanut agglutinin (PNA)-lectin immunofluorescence staining of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- testicular tissue.
Spermatids at cap phase and acrosome phase of acrosome biogenesis are shown individually. Scale bar: 5 μm.
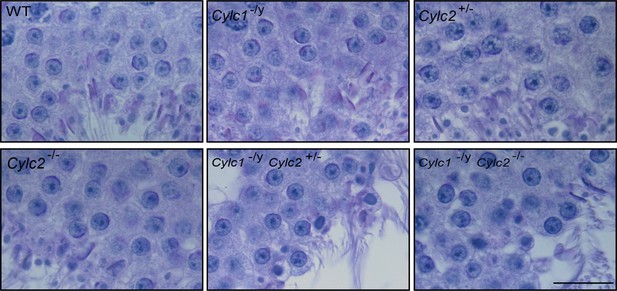
Periodic acid Schiff (PAS)-stained testicular tissue sections of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice.
Scale bar: 20 µm.
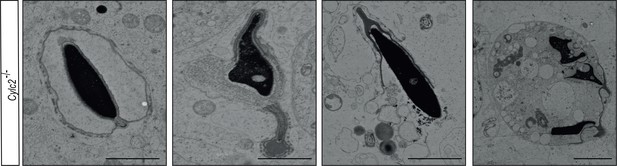
Transmission electron microscopy (TEM) micrographs of degrading damaged spermatids in testicular sections of Cylc2-/- mice.
Scale bar: 5 µm.
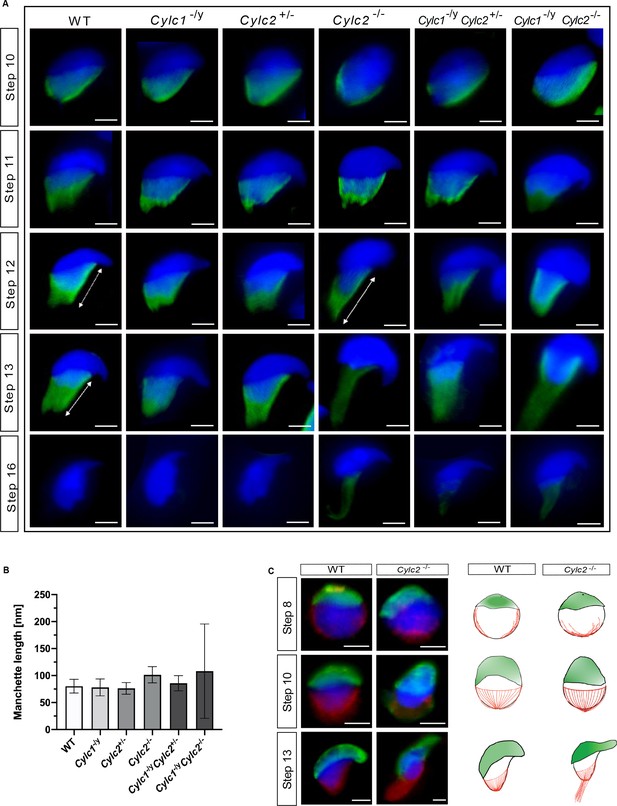
Cylc2 deficiency causes delay in manchette removal.
(A) Immunofluorescence staining of α-tubulin to visualize manchette structure in squash testis samples of wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- mice. Spermatids in different steps of spermiogenesis were shown, for step-to-step comparison. Scale bar: 5 µm. (B) Quantification of manchette length in WT, Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y, Cylc2+/-, and Cylc1-/y Cylc2-/- α-tubulin-stained spermatids at steps 10–13. (C) Co-staining of the manchette with HOOK1 (red) and acrosome with peanut agglutinin (PNA)-lectin (green) is shown in round, elongating and elongated spermatids of WT (upper panel) and Cylc2-/- mice (lower panel). Scale bar: 3 µm. Schematic representation shows acrosomal structure (green) and manchette filaments (red).
-
Figure 5—source data 1
Manchette length.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig5-data1-v2.zip
-
Figure 5—source data 2
Uncropped PNA-Hook1 staining.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig5-data2-v2.zip
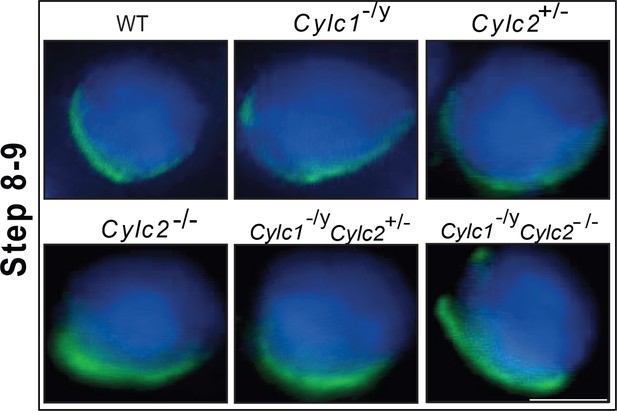
Immunofluorescence staining of α-tubulin in wildtype (WT), Cylc1-/y, Cylc2+/-, Cylc2-/-, Cylc1-/y Cylc2+/-, and Cylc1-/y Cylc2-/- squash testis samples.
Spermatids at steps 8–9 are shown. Scale bar: 10 μm.
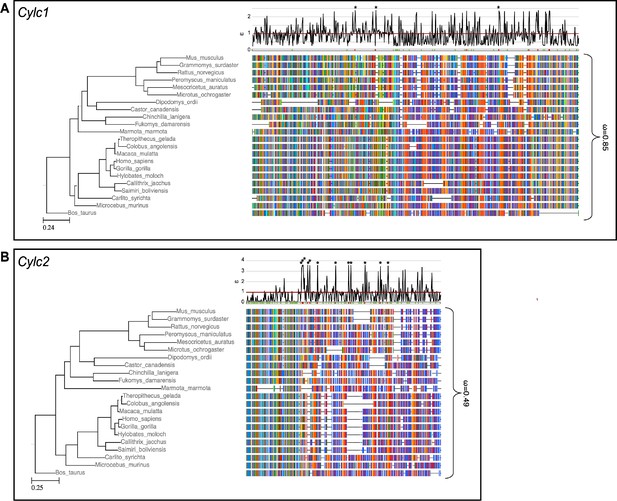
Species phylogeny with branch length representing number of nucleotide substitutions per codon with schematic representation of (A) CYLC1 and (B) CYLC2 amino acid alignment used in the PAML CodeML analysis.
Alignments were stripped of columns with gaps in more than 80% of species. Evolutionary rate (ω) obtained by CodeML models M0 is shown for the whole alignment. The graph on top shows the evolutionary rate (ω) per codon sites across the whole tree (CodeML model M2a). Significantly positively selected sites are marked by asterisks. Sites with a probability of higher than 0.95 to belonging to the conserved or positively selected site class are marked in green and red respectively below the graph.
-
Figure 6—source data 1
Code ML results table.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig6-data1-v2.zip
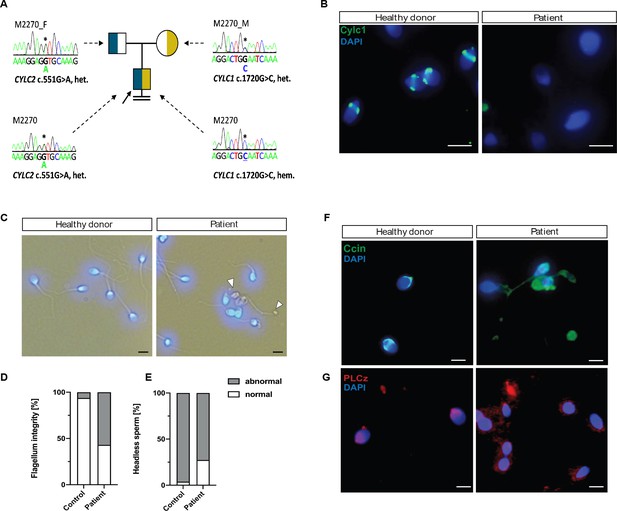
Cylicins are required for human male fertility.
(A) Pedigree of patient M2270. His father (M2270_F) is carrier of the heterozygous CYLC2 variant c.551G>A and his mother (M2270_M) carries the X-linked CYLC1 variant c.1720G>C in a heterozygous state. Asterisks (*) indicate the location of the variants in CYLC1 and CYLC2 within the electropherograms. (B) Immunofluorescence staining of CYLC1 in spermatozoa from healthy donor and patient M2270. In donor’s sperm cells CYLC1 localizes in the calyx, while patient’s sperm cells are completely missing the signal. Scale bar: 5 µm. (C) Bright-field images of the spermatozoa from healthy donor and M2270 show visible head and tail anomalies, coiling of the flagellum, as well as headless spermatozoa, which carry cytoplasmatic residues without nuclei (white arrowhead). Heads were counterstained with DAPI. Scale bar: 5 µm. (D–E) Quantification of flagellum integrity (D) and headless sperm (E) in the semen of patient M2270 and a healthy donor. (F–G) Immunofluorescence staining of CCIN (F) and PLCz (G) in sperm cells of patient M2270 and a healthy donor. Nuclei were counterstained with DAPI. Scale bar: 3 µm.
-
Figure 7—source data 1
Uncropped images of stainings on human sperm.
- https://cdn.elifesciences.org/articles/86100/elife-86100-fig7-data1-v2.zip

Variants in CYLC1 and CYLC2 identified in subject M2270 and their localization on the DNA and protein level.
(A) Localization of the CYLC1 variant found in M2270. The variant affects exon 4 and an intolerant part of the C-terminal region of CYLC1 according to metadome 41. (B) CYLC2 variant localization. The missense variant in CYLC2 detected in M2270 affects exon 5 and a tolerant part of CYLC2 according to metadome 41.
Videos
Full beat cycle of sperm from WT male.
Full beat cycle of sperm from Cylc2-/- male.
Tables
Semen analysis of the patient M2770 carrying variants in the CYLC1 and CYLC2 genes.
| First visit | Second visit | WHO reference rang | |
|---|---|---|---|
| Abstinence time (day) | 4.0 | 5.0 | |
| Volume (ml) | 4.2 | 5.8 | >1.4 |
| Concentration (Mill./ml) | 10.5 | 16.3 | >16 |
| Total sperm count (Mill.) | 44.1 | 94.5 | >39 |
| Vitality (%) | 53 | 27 | >54 |
| Motility | |||
| a (%) | 7 | 9 | a+b > 30 |
| b (%) | 5 | 4 | |
| c (%) | 19 | 8 | |
| d (%) | 69 | 79 | |
| Morphology | |||
| Normal (%) | 2 | 2 | >4 |
| Head defects (%) | 99 | 99 | |
| Midpiece defects (%) | 63 | 59 | |
| Flagella defects (%) | 18 | 47 |
Protospacer sequences.
| Name | Protospacer sequence (5′–3′) |
|---|---|
| Mm.Cas9.CYLC1.sg1 | GGTTTATCCATACGTGAGT |
| Mm.Cas9.CYLC1.sg2 | GGCTTAGGTGATGCTCTAAA |
| Mm.Cas9.CYLC2.1.AB | AAGGGAGAGTCGAAAAGCGT |
| Mm.Cas9.CYLC2.1.AF | GGATCCAAGGATAAAGTGTC |
PCR primer sequences.
| Cylc1 | 5′–3′ | Expected band size |
|---|---|---|
| Cylc1_F1 | TATACACACAATCCACAATCTTGAAAT | WT: 427 bp |
| Cylc1_R1 | TCACTTCAAAATCCAACTTGTCCT | KO: 264 bp |
| Cylc1_R2 | TGCCTAGTATTTCAGGTTCCCC | |
| Cylc2 | ||
| Cylc2_F1 | ACCACCATTATGGATGCACCG | WT: 376 bp |
| Cylc2_R1 | AGTGTTTCTTGTGAGCTCGTTG | KO: 286 bp |
| Cylc2_R2 | GGCTGAATCTTTACCCTTAGGT |
qRT primer sequences.
| Name | Forward (5′-3′) | Reverse (5′-3′) |
|---|---|---|
| Cylc1 | GGGGAAAAATAAGCTCATGTGTAG | TTCAGGTTCCCCATTGGTTA |
| Cylc2 | GCATTTCCCAAACCACCAAGG | AACGGATGGTCTCTCGGATATT |
| Beta-actin | TGTTACCAACTGGGACGACA | GGGTGTTGAAGGTCTCAAA |
Antibodies.
| Antibody | Company | Catalogue number | Antigen | Dilution IF | Dilution WB |
|---|---|---|---|---|---|
| α-Tubulin | Abcam | ab7291 | 1:10,000 in 3% BSA | ||
| α-Tubulin | Merck Millipore (Billerica, MA, USA) | 16-232 | 1:1000 | ||
| CapZa3 | Progen | GP-SH4 | 1:500 | ||
| Ccin | Progen | GP-SH3 | 1:500 | ||
| Cylc1 (used for mouse) | Davids Biotechnology (Regensburg, Germany) | Custom-made polyclonal antibody | AESRKSKNDERRKTLKIKFRGK and KDAKKEGKKKGKRESRKKR | 1:1000 (sperm cells) 1:500 (testis tissue) 1:1000 western blot | 1:1000 in 5% milk in TBST |
| Cylc1 (used for human samples) | Santa Cruz | sc-166400 | 1:500 | ||
| Cylc2 | Davids Biotechnology (Regensburg, Germany) | Custom-made polyclonal antibody | KSVGTHKSLASEKTKKEVK and ESGGEKAGSKKEAKDDKKDA | 1:1000 (sperm cells) 1:500 (testis tissue) 1:1000 western blot | 1:1000 in 5% milk in TBST |
| Hook1 | Proteintech | 10871-1-AP | 1:500 | ||
| PLCζ | Invitrogen | PA5-50476 | 1:100 | ||
| Sp56 | Invitrogen | MA1-10866 | 1:500 |







