Cancer Immunotherapy: A promising platform for predicting toxicity
Killing cancer cells can be done directly or by harnessing the immune system through an approach called cancer immunotherapy (Robert, 2020). Indeed, over the past decade, most of the advances in oncology have involved boosting immune cells to destroy tumor cells. Some of these treatments have resulted in impressive improvements in survival, bringing the possibility of a cure closer for some patients (Robert et al., 2018). This is a big milestone in oncology.
Despite these advances, immunotherapies can be associated with toxicity, which forces the treatment to be stopped: if immune cells become too activated, they can mistakenly recognize and destroy healthy tissues. This can lead, for example, to rashes, hepatitis or colitis depending on whether the skin, liver or gut are attacked (Marin-Acevedo et al., 2019).
A big challenge in oncology is therefore to predict which new immunotherapy drugs are going to be too harmful for patients. This is usually examined in animal models, but since their immune systems differ from the human immune system, it can be difficult to reliably predict toxicity (Zschaler et al., 2014). Now, in eLife, Nikolce Gjorevski (Roche), Lauriane Cabon (Roche) and colleagues – including Jordan Kerns and Chaitra Belgur of Emulate Inc in Boston as joint first authors – report how an in vitro model can help bypass this problem for T cell bispecific antibodies immunotherapy (Kerns et al., 2021).
T cell bispecific antibodies (or TCBs) can recognise and bind to ‘antigen’ proteins present on the surface of tumors, as well as receptors displayed by immune ‘T cells’: by bringing the two types of cells closer, this process helps to activate T cells and allows them to kill their targets. However, the antigens that TCBs bind to are not always exclusive to cancer cells. Recognition of non-cancer cells which share antigens with tumors – known as the on-target, off-tumor effect – can lead to normal cells being damaged (Labrijn et al., 2019; Figure 1). Predicting which TCBs under clinical development will cause such undesired toxicity represents an important challenge in oncology.
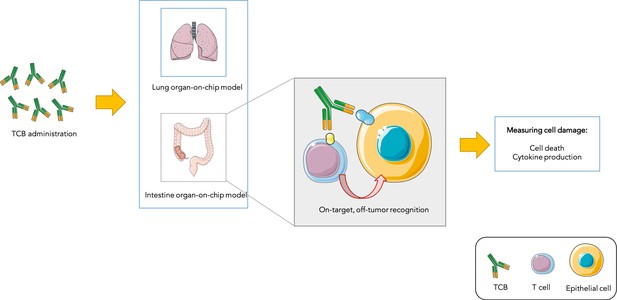
Testing drug toxicity using an organ-on-chip platform.
Administering TCBs to an organ-on-chip platform (such as a model of the lung or intestine) allows the antibodies (Y-shaped structures) to recognize antigens (shown in yellow and blue) present in both healthy epithelial and cancerous cells. This brings T cells close to their targets and also activates them, which can lead to healthy cells being damaged (as assessed by measuring cell death and inflammation levels). This model is able to partially reproduce toxicity observed in animals and humans.
Image credit: Image was prepared with Servier Medical Art (CC BY 3.0).
To address this issue, Kerns et al. first took advantage of a lung-on-chip model (Huh et al., 2010) – a system grown under conditions mimicking those found in the body – to predict toxicity to TCBs. This ‘mini-organ’ was exposed to a TCB that recognises an antigen present in ovarian, lung and breast cancer cells, but which can also be expressed, at lower levels, in healthy lung cells. This manipulation led to healthy cells being damaged in the lung-on-chip model, which was then used to determine which TCB dose could kill tumors while sparing normal lung cells. This dose was then administered to mouse models, whose lungs remained undamaged. This demonstrates that the lung-on-chip was able to efficiently predict toxicity in these animals.
Kerns et al. then used an intestine-on-chip model to test a TCB which targets an antigen present on both colon cancer and normal intestine cells. Healthy cells survived the treatment but signs of toxicity emerged that could mimick side-effects that commonly occur in patients, such as increased inflammation.
Using an organ-on-chip model to test TCBs therefore has two main advantages. First, it reproduces toxicity observed in vivo, and can help to determine which doses are effective while remaining safe for normal cells. Second, it serves to predict toxicity before patients are administered with newly-developed TCBs, filling the gap between animal and human studies for drugs that are yet to be clinically tested. Organ-on-chip models could therefore make drug development more efficient by helping to screen out toxic TCBs before they reach cancer patients.
References
-
Bispecific antibodies: a mechanistic review of the pipelineNature Reviews Drug Discovery 18:585–608.https://doi.org/10.1038/s41573-019-0028-1
-
Immune checkpoint inhibitor toxicitiesMayo Clinic Proceedings 94:1321–1329.https://doi.org/10.1016/j.mayocp.2019.03.012
-
Durable complete response after discontinuation of pembrolizumab in patients with metastatic melanomaJournal of Clinical Oncology 36:1668–1674.https://doi.org/10.1200/JCO.2017.75.6270
-
A decade of immune-checkpoint inhibitors in cancer therapyNature Communications 11:3801.https://doi.org/10.1038/s41467-020-17670-y
-
Differences in innate immune response between man and mouseCritical Reviews in Immunology 34:433–454.https://doi.org/10.1615/CritRevImmunol.2014011600
Article and author information
Author details
Publication history
Copyright
© 2021, Ochoa de Olza
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Cancer Biology
- Cell Biology
Testicular microcalcifications consist of hydroxyapatite and have been associated with an increased risk of testicular germ cell tumors (TGCTs) but are also found in benign cases such as loss-of-function variants in the phosphate transporter SLC34A2. Here, we show that fibroblast growth factor 23 (FGF23), a regulator of phosphate homeostasis, is expressed in testicular germ cell neoplasia in situ (GCNIS), embryonal carcinoma (EC), and human embryonic stem cells. FGF23 is not glycosylated in TGCTs and therefore cleaved into a C-terminal fragment which competitively antagonizes full-length FGF23. Here, Fgf23 knockout mice presented with marked calcifications in the epididymis, spermatogenic arrest, and focally germ cells expressing the osteoblast marker Osteocalcin (gene name: Bglap, protein name). Moreover, the frequent testicular microcalcifications in mice with no functional androgen receptor and lack of circulating gonadotropins are associated with lower Slc34a2 and higher Bglap/Slc34a1 (protein name: NPT2a) expression compared with wild-type mice. In accordance, human testicular specimens with microcalcifications also have lower SLC34A2 and a subpopulation of germ cells express phosphate transporter NPT2a, Osteocalcin, and RUNX2 highlighting aberrant local phosphate handling and expression of bone-specific proteins. Mineral disturbance in vitro using calcium or phosphate treatment induced deposition of calcium phosphate in a spermatogonial cell line and this effect was fully rescued by the mineralization inhibitor pyrophosphate. In conclusion, testicular microcalcifications arise secondary to local alterations in mineral homeostasis, which in combination with impaired Sertoli cell function and reduced levels of mineralization inhibitors due to high alkaline phosphatase activity in GCNIS and TGCTs facilitate osteogenic-like differentiation of testicular cells and deposition of hydroxyapatite.
-
- Cancer Biology
TAK1 is a serine/threonine protein kinase that is a key regulator in a wide variety of cellular processes. However, the functions and mechanisms involved in cancer metastasis are still not well understood. Here, we found that TAK1 knockdown promoted esophageal squamous cancer carcinoma (ESCC) migration and invasion, whereas TAK1 overexpression resulted in the opposite outcome. These in vitro findings were recapitulated in vivo in a xenograft metastatic mouse model. Mechanistically, co-immunoprecipitation and mass spectrometry demonstrated that TAK1 interacted with phospholipase C epsilon 1 (PLCE1) and phosphorylated PLCE1 at serine 1060 (S1060). Functional studies revealed that phosphorylation at S1060 in PLCE1 resulted in decreased enzyme activity, leading to the repression of phosphatidylinositol 4,5-bisphosphate (PIP2) hydrolysis. As a result, the degradation products of PIP2 including diacylglycerol (DAG) and inositol IP3 were reduced, which thereby suppressed signal transduction in the axis of PKC/GSK-3β/β-Catenin. Consequently, expression of cancer metastasis-related genes was impeded by TAK1. Overall, our data indicate that TAK1 plays a negative role in ESCC metastasis, which depends on the TAK1-induced phosphorylation of PLCE1 at S1060.






