Vibrio cholerae’s ToxRS bile sensing system
Figures
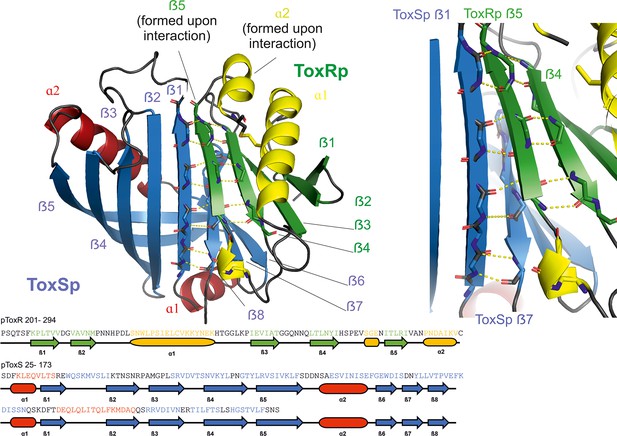
Heterodimer formation of ToxRSp (pdb: 8ALO).
ToxRp ß strand 5 completes main chain hydrogen bond network of ToxSp ß-barrel formation, by interaction with ToxSp ß1. ToxSp ß8 position is stabilized by main chain hydrogen bonds with ToxSp ß1, as well as side chain interactions with ToxRp ß5 and ß4. ToxRp ß5 and α2 are formed upon ToxSp interaction. ToxRSp crystal structure was determined by molecular replacement using an AlphaFold (Evans et al., 2021; Jumper et al., 2021) model as template.

Inner membrane spanning domains of Vibrio cholerae regulators ToxR and ToxS.
ToxRp, ToxSp stand for the periplasmic domains of ToxR and ToxS, respectively. ToxRSp is the complex formation of periplasmic domains of ToxR and ToxS.

ToxRSp structure highlighting L33ToxSp core stabilizing hydrophobic network.
According to previous studies ToxSp L33S mutant results in a loss of ToxSp protective function against ToxRp proteolysis (Pfau and Taylor, 1998). ToxRSp crystal structure reveals a central function of ToxSp L33 in stabilizing the N-terminal membrane oriented core visible in the zoomed region.

Structural alignment of bile interacting protein complexes ToxRS V. cholerae and VtrAC V. parahaemolyticus.
VtrAC (gold; pdb: 5KEV) and ToxRSp (dark blue; pdb: 8ALO) reveal a similar fold (RMSD: 3.8 Å). VtrC and ToxSp form a ß-barrel fold, which is stabilized by ß-strand 5 from VtrA and ToxRp, resulting in an obligate heterodimer formation including a binding pocket in the barrel cavity.
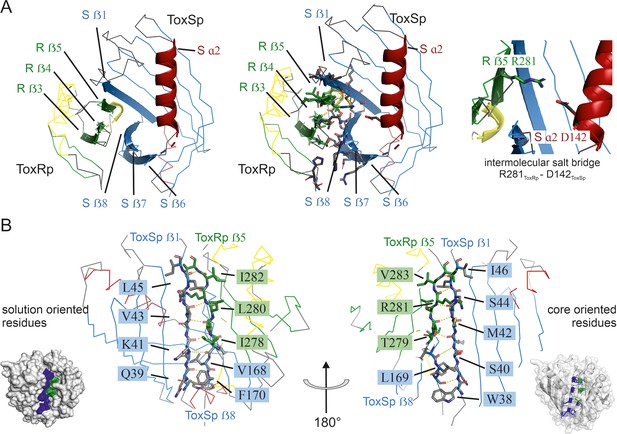
Detailed description of ToxRSp interface.
(A) Overview of ToxRSp interface. The interaction is mainly established via ToxRp ß strands 3, 4 and 5, and ToxSp ß strands 6, 7 and 8. Additionally, an intermolecular salt bridge is formed between ToxRp R281 and ToxSp D142 located at the opening of ToxSp barrel. (B) The intermolecular main chain H-bond pattern of ToxRSp. The interprotein H-bond network is established between ToxRp ß5 and ToxSp ß1 thereby filling the gap between ToxSp ß1 and ß8, which interact only in the C-terminal region of the strands.
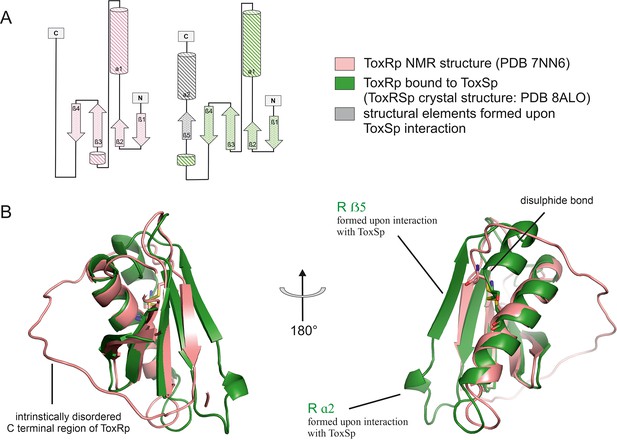
Comparison of ToxRp NMR structure (salmon) to ToxRp bound to ToxSp (green).
Upon binding to ToxSp, ToxRp intrinsically disordered C-terminal region forms ß5 and α2. (A) Topology diagram of ToxRp and ToxRp bound to ToxSp. Newly formed structural elements are highlighted in grey. (B) Structural alignment between ToxRp (PDB 7NN6) and ToxRp in complex with ToxSp (PDB 8ALO). The C-terminal disulphide bond is shown in sticks.
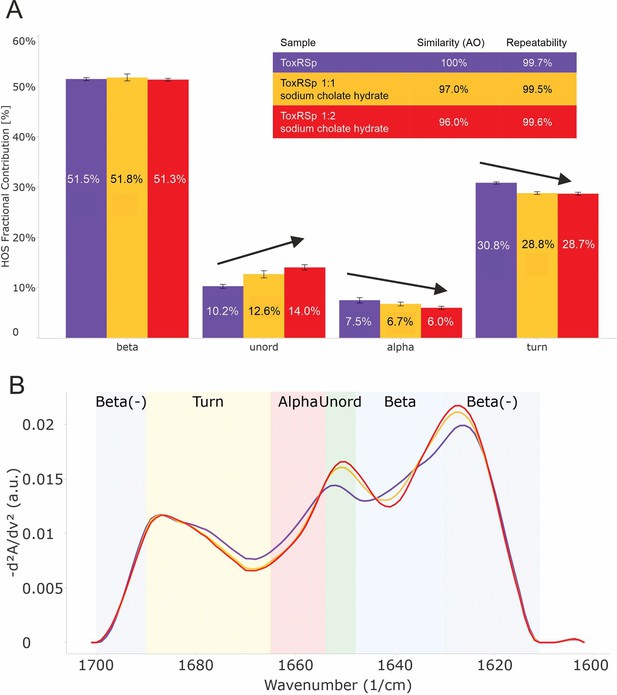
Analysis of microfluidic modulation spectroscopy MMS experiments.
MMS experiments of ToxRSp with different S-CH concentrations show conformational changes upon binding of bile to ToxRSp. Additional structural rearrangements could be detected upon higher additions of bile. (A) Bar chart of the relative abundance of secondary structural motifs based on Gaussian deconvolution of the corresponding spectra. (B) MMS spectra (baselined, inverted 2nd Derivative) of ToxRSp with and without bile, showing spectral changes upon higher bile additions.
-
Figure 4—source data 1
MMS data.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig4-data1-v2.zip

MMS titration experiment with ToxRSp and S-CH.
Six samples were measured with increasing molar ratios of S-CH to ToxRSp: 0, 1, 2, 3, 5, 20. Significant conformational changes occur within 1:1 and 1:2 samples, whereas higher bile to protein ratios does not seem to induce additional significant spectral changes.
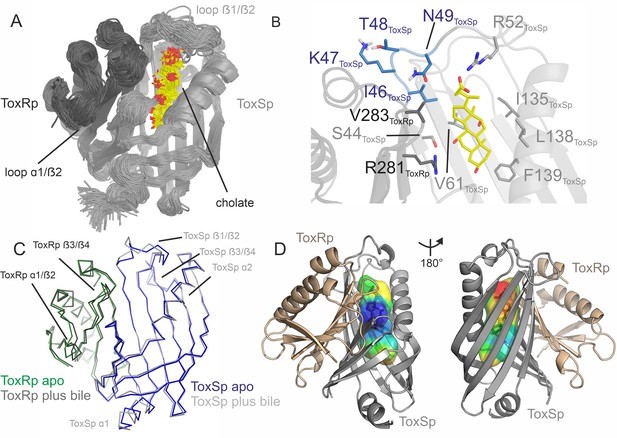
ToxRSp binding pocket for bile acid.
(A) Superimposition of 100 structures of the ToxRSp complex (dark grey/light grey cartoons) with cholate (sticks, C-atoms in yellow) along one representative classical MD simulation. (B) Detailed view of the binding mode of cholate to ToxRSp. The residues with the larger contribution to the binding of the ligand are highlighted as well as residues 46–49 of ToxSp located on loop β1/β2. (C) Overlay of ToxRSp apo and bile-bound structures. Conformational changes mainly occur in ToxSp loop β1/β2 and ToxRp loop α1/β2. ToxSp helices α1 and α2 and ToxRp loop β3/β4, both close to the ToxRSp interface, experience minor conformational rearrangements upon bile interaction. (D) Hydrophobic properties of the bile binding cavity calculated with the CavMan (v. 0.1, 2019, Innophore GmbH plugin in PyMOL). The entrance of the cavity (blue to green) faces the solvent exhibiting a hydrophilic environment, in contrast to the buried hydrophobic areas (yellow to red). For the analysis of the hydrophobicity of the cavities the program VASCo (Steinkellner et al., 2009) was used; cavities were calculated using a LIGSITE algorithm (Hendlich et al., 1997).
-
Figure 5—source data 1
ToxRSp MD data.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig5-data1-v2.zip

Hydrophobic cleft of ToxSp binding pocket in complex with ToxRp. Hydrophobic (ochre) and hydrophilic (teal) surface representation of ToxS calculated with ChimeraX mlp. The bile binding cavity is highly hydrophobic (arrow).
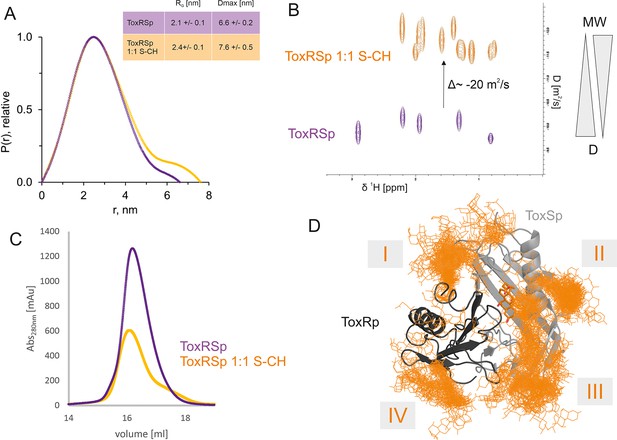
ToxRSp bile interaction experiments propose additional bile interaction areas I-IV.
(A) SEC-SAXS experiments with ToxRSp and S-CH reveal an increase of the radius of gyration RG and the effective maximum particle dimension Dmax upon bile addition. (B) Superimposed NMR DOSY spectra of ToxRSp with and without bile acid S-CH. Addition of bile acid causes a decrease of the diffusion coefficient due to an increase of molecular weight. (C) SEC experiments with ToxRSp and S-CH reveal a slight decrease of retention volume upon bile acid presence from 16.19 ml to to 16.08 ml and a broadening of the peak. (D) MD determined surface exposed bile interacting regions of ToxRSp. Besides the bile binding cavity of ToxSp, four regions (marked as I-IV) on the ToxRSp surface could be mapped as additional bile binding areas. Region I and III involves both proteins, whereas region II is located on ToxSp and region IV on ToxRp. The representative structure of the ToxRSp complex (ToxRp: dark grey, ToxSp: light grey) is shown with cholate bound at the cavity (orange) and additional cholate molecules attach to surface exposed regions of ToxRSp.
-
Figure 6—source data 1
SEC data.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig6-data1-v2.zip

Analysis of size-exclusion chromatography coupled with solution small-angle X-ray scattering SEC-SAXS experiments.
ToxRp (green), ToxSp (blue), ToxRSp (purple) and ToxRSp:bile (orange). (A) The SEC-SAXS profiles are compared to the theoretical scattering curves (Crysol). Chi² values are indicated. Profiles are shifted along the y-axis for clarity. The following models were used ToxRp – NMR structure pdb: 7NN6; ToxSp – the dimeric model (AlphaFold-Multimer); ToxRSp – crystal structure pdb (8ALO); ToxRSp plus bile acid (1:1)- MD complex (Figure 2). (B) p(r) functions. (C) Overlay of the models listed in (A) with the respective bead models calculated from the SAXS profiles. Note, for ToxRp and ToxRSp:bile an excess of volume indicates overall larger size. (D) Dimensionless Kratkyplot ((qRG)2I(q)/I(0) vs qRG). Note, dotted lines are drawn at y=1.104 and x = √ 3. Folded proteins have a local maximum where the two lines intersect. Here, the maxima of ToxRp and ToxRSp:bile are shifted, indicating a flexible/elongated structure.

SAXS-derived chromatogram showing Integrated intensities (0.1 nm−1<s < 0.8 nm−1) vs. elution of indicated samples.
Note, the bile complex was separated on a Superdex 75 increase 5/150 column; the other samples on the larger Superdex 75 10/300 column.

Saturation transfer difference STD NMR spectra of ToxRSp with S-CH.
STD experiments reveal a binding of ToxRSp to bile acid S-CH. Saturation of the amide region of ToxRSp resulted in signals of the ligand in the STD spectra. (A) Overlay of ToxRSp spectra with and without S-CH. (B–D) STD spectra after saturation of different regions: (B) amide region 5000 Hz, (C) amide region 6000 Hz, (D) negative control –5000 Hz. (E) Reference spectrum for S-CH 2 mM.
-
Figure 6—figure supplement 3—source data 1
STD ToxRSp plus S-CH.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig6-figsupp3-data1-v2.zip

Saturation transfer difference NMR spectra of ToxSp with sodium cholate hydrate.
Spectra does not show an interaction between ToxSp and bile. No ligand signals could be observed upon saturation of amide signals of ToxSp. (A) Overlay of ToxSp spectra with and without S-CH. (B–D) STD spectra after saturation of different regions: (B) amide region 5000 Hz, (C) amide region 6000 Hz, (D) negative control –5000 Hz.
-
Figure 6—figure supplement 4—source data 1
STD ToxSp plus S-CH.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig6-figsupp4-data1-v2.zip

1 H NMR spectrum of ToxRSp upon S-CH addition.
Addition of increasing amounts of bile acid S-CH causes broadening of peaks and chemical shift changes indicating a binding event and increase of molecular weight. Reference spectrum of ToxRSp is shown in purple, spectrum recorded after addition of 0.5 and equimolar molar ratios of S-CH to ToxRSp is shown in orange and green, respectively.

Superimposed NMR DOSY spectra of ToxRSp with and without bile acid sodium cholate hydrate.
X-axis represents 1H chemical shift, y-axis the determined diffusion coefficient D [m²/s]. Three samples were measured: ToxRSp (purple), ToxRSp 1:1 bile acid (orange), ToxRSp 1:20 bile acid (green). Addition of bile acid causes a decrease of the diffusion coefficient due to an increase of molecular weight. Higher bile acid concentrations result in higher diffusion coefficients via averaging due to exchange of bound and free bile molecules.

SEC runs of ToxRSp with and without bile acid S-CH.
x-axis represents the retention volume in ml, y-axis shows the absorption at 280 nm [mAU]. Maximum absorption of ToxRSp (purple) and ToxRSp with bile acid (1:1) (orange) is observed at 16.19 ml and 16.08 ml, respectively. Bile addition leads to a slight decrease of the retention volume and peak broadening.

NMR interaction experiment of ToxRSp with cyclo-PhePro (cFP).
Previous experiments revealed a ToxR-dependent downregulation of virulence factor production in response to cFP (Bina et al., 2013; Bina and Bina, 2010). NMR interaction experiments with ToxRSp do not indicate a binding event between ToxRSp and cFP since upon cFP addition no significant changes of chemical shift or intensity of ToxRSp signals could be observed.
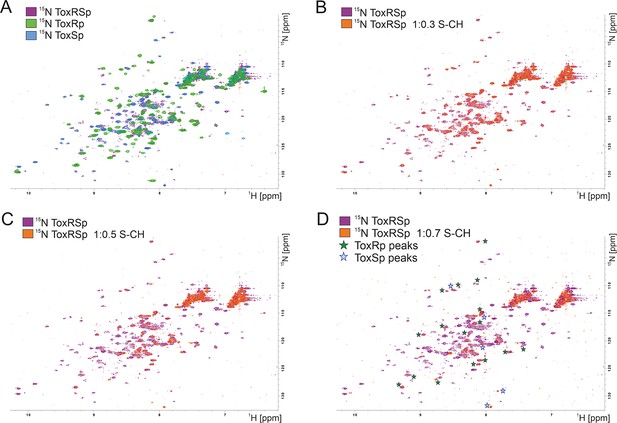
NMR titration of S-CH to 15 N labelled ToxRSp.
(A) Overlay of 15N-HSQC spectra of 15N-labelled ToxRSp (purple) with 15N-labelled ToxRp (green) and ToxSp (blue) for ascription of peaks to each of the proteins. (B–D) Overlay of 15N-HSQC spectra of 15N-labelled ToxRSp before (purple) and after the addition of increasing ratios of S-CH (orange): 0.3 (B), 0.5 (C), 0.7 (D). Upon bile acid addition signal intensities significantly decrease. Upon higher additions of bile acid mainly ToxRp peaks are visible indicated by green asterisk in (D), whereas most ToxSp signals disappear indicating direct interaction with S-CH. Remaining ToxSp signals are marked with blue asterisk.
-
Figure 7—source data 1
NMR titration 1 ToxRSp plus S-CH.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig7-data1-v2.zip
-
Figure 7—source data 2
NMR titration 2 ToxRSp plus S-CH.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig7-data2-v2.zip
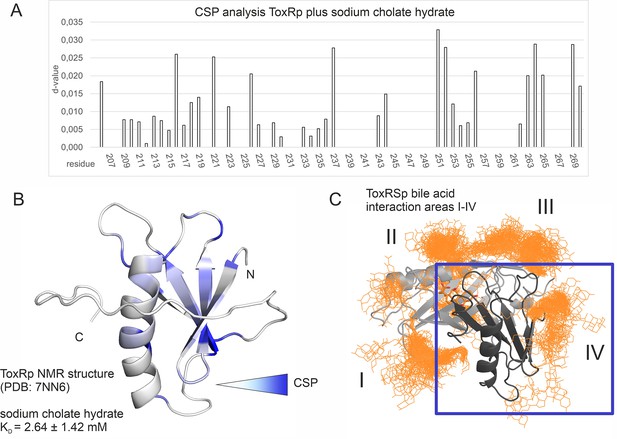
Binding studies with ToxRp and bile.
(A) Chemical shift perturbation experiments expose residues mostly affected upon binding of S-CH. (B) Residues which experience a change of their chemical shift upon bile addition are coloured according to a gradient from white to blue. Residues highlighted in dark blue are mostly affected upon bile addition and are therefore most likely located at the interaction area. The calculated dissociation constant of 2.6 mM suggests a low affinity binding of bile to ToxRp. (C) Binding area IV determined by MD simulations (Figure 6) is located at the ß-sheet of ToxRp. Taken together, MD simulation and CSP experiments reveal a bile interacting area located at the ß-sheet of ToxRp proposing a bile sensing function of ToxRp independent on ToxRSp complex formation relevant only at high bile salt concentrations.
-
Figure 8—source data 1
NMR titration ToxRp plus S-CH.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig8-data1-v2.zip

NMR titration experiment with ToxRp and S-CH.
Overlay of 15N-HSQC spectra of ToxRp with increasing amounts of bile acid S-CH proposes a weak interaction event. Significant chemical shift changes of ToxRp could be observed with high ligand concentrations of 1:50-1:200.
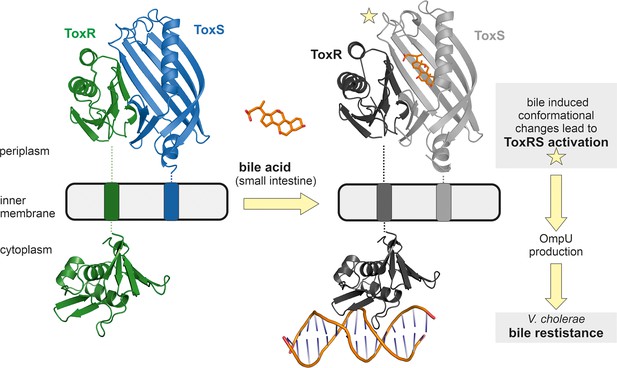
Model of bile induced activation of V. cholerae ToxRS.
When entering the human gut V. cholerae senses the presence of bile acids by binding of bile to the periplasmic domains of inner membrane proteins ToxR and ToxS. The bile binding pocket is formed by ToxS and stabilized by ToxR. Subsequently, the interaction with bile induces ToxR activation which leads to a change of outer-membrane proteins from OmpT to OmpU. OmpU then provides bile resistance and thus enables the bacterium’s survival in the human body. PDB accession codes: ToxRSp crystal structure: 8ALO, cytoplasmic domain of ToxR: 7NMB. The cytoplasmic domain of ToxR bound to ompU operon is a NMR guided HADDOCK model (Gubensäk et al., 2021a).
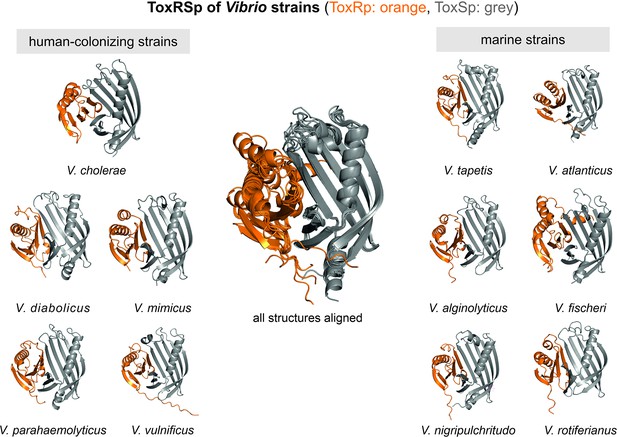
Heterodimer structures of periplasmic sensory domains of ToxRS from Vibrio organisms calculated using AlphaFold-Multimer (Evans et al., 2021; Jumper et al., 2021).
ToxRSp dimer formation occurs in different Vibrio strains. The structural alignment of all models reveal that the ToxSp fold is highly similar in whereas ToxRp seems to be more diverse.
-
Figure 10—source data 1
coordinate files of Vibrio AlphaFold models.
- https://cdn.elifesciences.org/articles/88721/elife-88721-fig10-data1-v2.zip

Overlay of ToxRSp from different Vibrio species.
Structures are calculated using AlphaFold-Multimer (Evans et al., 2021; Jumper et al., 2021). The structural alignment reveals a conserved fold of ToxSp (grey), whereas ToxRp (orange) varies among different Vibrio strains. The intermolecular salt bridge between ToxRp and ToxSp is highlighted, and residues types involved are mentioned in the table beside the aligned structures. ToxSp salt bridge forming aspartate residue is strictly conserved.
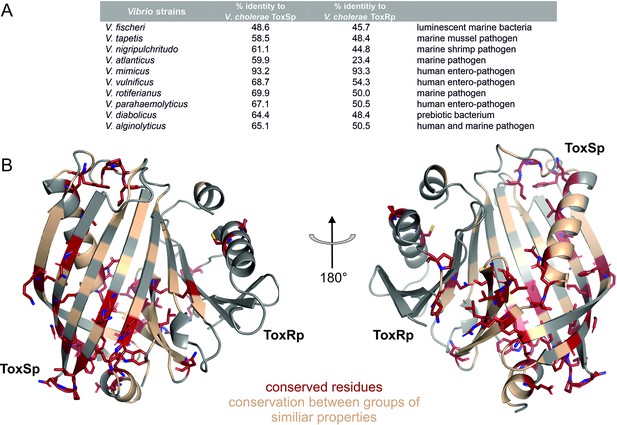
Sequential and structural analysis of sensory domains of ToxRS from different Vibrio strains.
ToxRSp from different Vibrio species exhibit similar heterodimer formations. Especially ToxSp ß-barrel structure, which forms the binding cavity, seems to be conserved. (A) Sequential analysis of conserved residues of ToxRSp. Sequential alignment was done using ToxRS amino acid sequences from Vibrio species (Supplementary file 1e). The sequence identity (%) of ToxR and ToxS proteins from different strains to V. cholerae is shown in the table above the structures. (B) Conserved residues are highlighted on the ToxRSp structure. Conserved residues are coloured in red and shown in sticks, residues which are conserved between groups of similar properties are coloured in gold. Most conserved regions could be found in ToxSp, especially at the openings of the barrel. ToxRp seems to have a higher sequential variability.

Sequential analysis of ToxSp proteins from different Vibrio species.
Analysis was performed with Clustal2.1 (Madeira et al., 2022). Strongly conserved residues are marked with an asterisk (*), conserved residues with colon (:) and period (.). Below the sequence identities are shown in percent.

Sequential analysis of ToxRp proteins from different Vibrio species.
Analysis was performed with Clustal2.1 (Madeira et al., 2022). Strongly conserved residues are marked with an asterisk (*), conserved residues with colon (:) and period (.). Below the sequence identities are shown in percent. Due to the higher variability of ToxRp in Vibrio’s an additional analysis was performed for closely related strains, which are mostly human colonizing. Source files.
Additional files
-
Supplementary file 1
Supplementary tables for 'Vibrio cholerae's ToxRS Bile Sensing System'.
(a) Description of expression constructs. List of plasmids and amino acid sequences used for the expression of ToxRp and ToxSp. (b) Crystal data and structure refinement table of ToxRSp. (c) Analysis of size-exclusion chromatography coupled with solution small-angle X-ray scattering SEC-SAXS. According to SAXS curves, ToxRp is monomeric in solution, whereas ToxSp seems to form dimers. Upon bile addition, the radius of gyration RG, the maximum particle size Dmax and the molecular weight of the ToxRSp complex increases. (d) NMR derived dissociation constants for ToxRp and sodium cholate hydrate. NMR titration experiments reveal a weak binding of bile to ToxRp with a dissociation constant of 2.6 mM. Table S5 includes ToxRp residue number, chemical shift distance, dissociation constant Kd [M], error of Kd. The calculated average Kd is 2.7±1.4 mM. (e) List of amino acid sequences of sensory domains of ToxRS proteins from different Vibrio species. Table S5 includes UniProt ID of the used sequences as well as sequence identity values to ToxRSp from V. cholerae. (f) Structural homology search of ToxSp bound to ToxRp using Dali server (Holm, 2022).
- https://cdn.elifesciences.org/articles/88721/elife-88721-supp1-v2.docx
-
Supplementary file 2
ToxRSp atom-atom interactions.
List of all atom interactions of ToxRSp (pdb: 8ALO). The list was created using PDBsum online server (Laskowski et al., 2018).
- https://cdn.elifesciences.org/articles/88721/elife-88721-supp2-v2.docx
-
MDAR checklist
- https://cdn.elifesciences.org/articles/88721/elife-88721-mdarchecklist1-v2.pdf





